The Cyclin K/Cdk12 Complex Maintains Genomic Stability Via Regulation of Expression of DNA Damage Response Genes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Analysis of Trans Esnps Infers Regulatory Network Architecture
Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2014 © 2014 Anat Kreimer All rights reserved ABSTRACT Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer eSNPs are genetic variants associated with transcript expression levels. The characteristics of such variants highlight their importance and present a unique opportunity for studying gene regulation. eSNPs affect most genes and their cell type specificity can shed light on different processes that are activated in each cell. They can identify functional variants by connecting SNPs that are implicated in disease to a molecular mechanism. Examining eSNPs that are associated with distal genes can provide insights regarding the inference of regulatory networks but also presents challenges due to the high statistical burden of multiple testing. Such association studies allow: simultaneous investigation of many gene expression phenotypes without assuming any prior knowledge and identification of unknown regulators of gene expression while uncovering directionality. This thesis will focus on such distal eSNPs to map regulatory interactions between different loci and expose the architecture of the regulatory network defined by such interactions. We develop novel computational approaches and apply them to genetics-genomics data in human. We go beyond pairwise interactions to define network motifs, including regulatory modules and bi-fan structures, showing them to be prevalent in real data and exposing distinct attributes of such arrangements. We project eSNP associations onto a protein-protein interaction network to expose topological properties of eSNPs and their targets and highlight different modes of distal regulation. -

Cyclin K Interacts with Β-Catenin to Induce Cyclin D1 Expression And
Theranostics 2020, Vol. 10, Issue 24 11144 Ivyspring International Publisher Theranostics 2020; 10(24): 11144-11158. doi: 10.7150/thno.42578 Research Paper Cyclin K interacts with β-catenin to induce Cyclin D1 expression and facilitates tumorigenesis and radioresistance in lung cancer Guojun Yao*, Jing Tang*, Xijie Yang, Ye Zhao, Rui Zhou, Rui Meng, Sheng Zhang, Xiaorong Dong, Tao Zhang, Kunyu Yang, Gang Wu and Shuangbing Xu Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. *These authors contributed equally to this work. Corresponding author: Shuangbing Xu or Gang Wu, Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. E-mail: [email protected] or [email protected]. © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2019.11.29; Accepted: 2020.08.24; Published: 2020.09.11 Abstract Rationale: Radioresistance remains the major cause of local relapse and distant metastasis in lung cancer. However, the underlying molecular mechanisms remain poorly defined. This study aimed to investigate the role and regulatory mechanism of Cyclin K in lung cancer radioresistance. Methods: Expression levels of Cyclin K were measured by immunohistochemistry in human lung cancer tissues and adjacent normal lung tissues. Cell growth and proliferation, neutral comet and foci formation assays, G2/M checkpoint and a xenograft mouse model were used for functional analyses. Gene expression was examined by RNA sequencing and quantitative real-time PCR. -
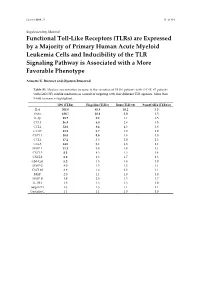
Are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway Is Associated with a More Favorable Phenotype
Cancers 2019, 11 S1 of S19 Supplementary Material Functional Toll-Like Receptors (TLRs) are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway is Associated with a More Favorable Phenotype Annette K. Brenner and Øystein Bruserud Table S1. Median concentration increase in the secretion of 19 (16 patients with G-CSF, 67 patients with GM-CSF) soluble mediators as a result of targeting with four different TLR agonists. More than 5-fold increase is highlighted. LPS (TLR4) Flagellin (TLR5) R848 (TLR7/8) Pam3CSK4 (TLR1/2) IL-6 301.0 45.9 10.2 5.3 TNFα 188.7 10.6 5.0 1.5 IL-1β 89.7 9.2 1.1 1.5 CCL3 56.4 6.0 2.4 1.5 CCL2 52.6 9.4 4.3 2.8 G-CSF 51.9 2.7 1.0 1.0 CXCL1 38.0 5.8 1.4 1.5 CCL4 17.2 3.3 2.0 1.3 CCL5 14.5 2.1 1.3 1.1 MMP-1 11.1 3.4 1.4 1.1 CXCL5 8.3 4.3 1.3 1.4 CXCL8 6.0 4.3 1.7 1.3 GM-CSF 5.2 1.5 1.4 1.0 MMP-2 4.0 1.5 1.2 1.1 CXCL10 3.9 1.6 2.2 1.1 HGF 2.0 1.1 1.0 1.0 MMP-9 1.9 2.5 1.3 1.7 IL-1RA 1.8 1.3 1.3 1.0 Serpin E1 1.3 1.3 1.1 1.1 Cystatin C 1.1 1.1 1.0 1.0 Cancers 2019, 11 S2 of S19 Table S2. -
Transcriptional Signature of Accessory Cells in the Lateral Line, Using the Tnk1bp1:EGFP Transgenic Zebrafish Line Behra Et Al
Transcriptional signature of accessory cells in the lateral line, using the Tnk1bp1:EGFP transgenic zebrafish line Behra et al. Behra et al. BMC Developmental Biology 2012, 12:6 http://www.biomedcentral.com/1471-213X/12/6 (24 January 2012) Behra et al. BMC Developmental Biology 2012, 12:6 http://www.biomedcentral.com/1471-213X/12/6 RESEARCHARTICLE Open Access Transcriptional signature of accessory cells in the lateral line, using the Tnk1bp1:EGFP transgenic zebrafish line Martine Behra1,3*, Viviana E Gallardo1, John Bradsher3, Aranza Torrado3, Abdel Elkahloun1, Jennifer Idol1, Jessica Sheehy1, Seth Zonies1, Lisha Xu1, Kenna M Shaw2,5, Chie Satou4, Shin-ichi Higashijima4, Brant M Weinstein2 and Shawn M Burgess1 Abstract Background: Because of the structural and molecular similarities between the two systems, the lateral line, a fish and amphibian specific sensory organ, has been widely used in zebrafish as a model to study the development/ biology of neuroepithelia of the inner ear. Both organs have hair cells, which are the mechanoreceptor cells, and supporting cells providing other functions to the epithelium. In most vertebrates (excluding mammals), supporting cells comprise a pool of progenitors that replace damaged or dead hair cells. However, the lack of regenerative capacity in mammals is the single leading cause for acquired hearing disorders in humans. Results: In an effort to understand the regenerative process of hair cells in fish, we characterized and cloned an egfp transgenic stable fish line that trapped tnks1bp1, a highly conserved gene that has been implicated in the maintenance of telomeres’ length. We then used this Tg(tnks1bp1:EGFP) line in a FACsorting strategy combined with microarrays to identify new molecular markers for supporting cells. -

Cytotoxic Effects and Changes in Gene Expression Profile
Toxicology in Vitro 34 (2016) 309–320 Contents lists available at ScienceDirect Toxicology in Vitro journal homepage: www.elsevier.com/locate/toxinvit Fusarium mycotoxin enniatin B: Cytotoxic effects and changes in gene expression profile Martina Jonsson a,⁎,MarikaJestoib, Minna Anthoni a, Annikki Welling a, Iida Loivamaa a, Ville Hallikainen c, Matti Kankainen d, Erik Lysøe e, Pertti Koivisto a, Kimmo Peltonen a,f a Chemistry and Toxicology Research Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, Finland b Product Safety Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, c The Finnish Forest Research Institute, Rovaniemi Unit, P.O. Box 16, FI-96301 Rovaniemi, Finland d Institute for Molecular Medicine Finland (FIMM), University of Helsinki, P.O. Box 20, FI-00014, Finland e Plant Health and Biotechnology, Norwegian Institute of Bioeconomy, Høyskoleveien 7, NO -1430 Ås, Norway f Finnish Safety and Chemicals Agency (Tukes), Opastinsilta 12 B, FI-00521 Helsinki, Finland article info abstract Article history: The mycotoxin enniatin B, a cyclic hexadepsipeptide produced by the plant pathogen Fusarium,isprevalentin Received 3 December 2015 grains and grain-based products in different geographical areas. Although enniatins have not been associated Received in revised form 5 April 2016 with toxic outbreaks, they have caused toxicity in vitro in several cell lines. In this study, the cytotoxic effects Accepted 28 April 2016 of enniatin B were assessed in relation to cellular energy metabolism, cell proliferation, and the induction of ap- Available online 6 May 2016 optosis in Balb 3T3 and HepG2 cells. The mechanism of toxicity was examined by means of whole genome ex- fi Keywords: pression pro ling of exposed rat primary hepatocytes. -

Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain a Target Enabling Package (TEP)
Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain A Target Enabling Package (TEP) Gene ID / UniProt ID / EC CDK12, 51755 / Q9NYV4/ 2.7.11.22, 2.7.11.23 CCNK, 8812 / O75909/ - Target Nominator Gregg Morin (UBC, Canada), Nathanael Gray (Harvard) SGC Authors Sarah E. Dixon-Clarke, Jonathan M. Elkins, Nathanael S. Gray, and Alex N. Bullock Collaborating Authors S.-W. Grace Cheng1, Tinghu Zhang2, Nicholas Kwiatkowski3, Calla M. Olson2, Brian J. Abraham3, Ann K. Greifenberg4, Scott B. Ficarro2, Yanke Liang2, Nancy M. Hannett3, Theresa Manz5, Mingfeng Hao2, Bartlomiej Bartkowiak6, Arno L. Greenleaf6, Jarrod A. Marto2, Matthias Geyer4, Richard A. Young3, and Gregg B. Morin1 Target PI Alex Bullock (SGC Oxford) Therapeutic Area(s) Oncology Disease Relevance CDK12 loss sensitises cancer cells to DNA damage Date approved by TEP 17th June 2016 Evaluation Group Document version Version 3 Document version date April 2018 DOI https://doi.org/10.5281/zenodo.1219680 Affiliations 1. Canada's Michael Smith Genome Sciences Centre, BC Cancer Agency 2. Department of Cancer Biology, Dana-Farber Cancer Institute 3. Whitehead Institute for Biomedical Research 4. Department of Structural Immunology, Institute of Innate Immunity 5. Pharmaceutical and Medicinal Chemistry, Saarland University 6. Department of Biochemistry, Duke University Medical Center USEFUL LINKS (Please note that the inclusion of links to external sites should not be taken as an endorsement of that site by the SGC in any way) SUMMARY OF PROJECT Cyclin-dependent kinase 12 (CDK12) phosphorylates RNA Pol II C-terminal domain (CTD) to promote transcriptional elongation of large DNA damage response genes. CDK12 is frequently mutated or amplified in cancer and its loss sensitises cells to DNA damage. -

WO 2012/174282 A2 20 December 2012 (20.12.2012) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2012/174282 A2 20 December 2012 (20.12.2012) P O P C T (51) International Patent Classification: David [US/US]; 13539 N . 95th Way, Scottsdale, AZ C12Q 1/68 (2006.01) 85260 (US). (21) International Application Number: (74) Agent: AKHAVAN, Ramin; Caris Science, Inc., 6655 N . PCT/US20 12/0425 19 Macarthur Blvd., Irving, TX 75039 (US). (22) International Filing Date: (81) Designated States (unless otherwise indicated, for every 14 June 2012 (14.06.2012) kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, English (25) Filing Language: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, Publication Language: English DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, KR, (30) Priority Data: KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, ME, 61/497,895 16 June 201 1 (16.06.201 1) US MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, 61/499,138 20 June 201 1 (20.06.201 1) US OM, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SC, SD, 61/501,680 27 June 201 1 (27.06.201 1) u s SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, 61/506,019 8 July 201 1(08.07.201 1) u s TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. -

Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain a Target Enabling Package (TEP)
Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain A Target Enabling Package (TEP) Gene ID / UniProt ID / EC CDK12, 51755 / Q9NYV4/ 2.7.11.22, 2.7.11.23 CCNK, 8812 / O75909/ - Target Nominator Gregg Morin (UBC, Canada), Nathanael Gray (Harvard) SGC Authors Sarah E. Dixon-Clarke, Jonathan M. Elkins, and Alex N. Bullock Collaborating Authors S.-W. Grace Cheng1, Tinghu Zhang2, Nicholas Kwiatkowski3, Calla M. Olson2, Brian J. Abraham3, Ann K. Greifenberg4, Scott B. Ficarro2, Yanke Liang2, Nancy M. Hannett3, Theresa Manz5, Mingfeng Hao2, Bartlomiej Bartkowiak6, Arno L. Greenleaf6, Jarrod A. Marto2, Matthias Geyer4, Richard A. Young3, Gregg B. Morin1 and Nathanael S. Gray2 Target PI Alex Bullock (SGC Oxford) Therapeutic Area(s) Oncology Disease Relevance CDK12 loss sensitises cancer cells to DNA damage Date approved by TEP 17th June 2016 Evaluation Group Document version Version 9 Document version date March 3rd 2021 Citation Sarah E. Dixon-Clarke, Jonathan M. Elkins, S.-W. Grace Cheng, Tinghu Zhang, Nicholas Kwiatkowski, Calla M. Olson, Brian J. Abraham, Ann K. Greifenberg, Scott B. Ficarro, Yanke Liang, Nancy M. Hannett, Theresa Manz, Mingfeng Hao, Bartlomiej Bartkowiak, Arno L. Greenleaf, Jarrod A. Marto, Matthias Geyer, Richard A. Young, Gregg B. Morin, Nathanael S. Gray and Alex N. Bullock. (2016) Human Cyclin-dependent kinase 12 (CDK12), kinase domain; A Target Enabling Package. 10.5281/zenodo.1219680. Affiliations 1. Canada's Michael Smith Genome Sciences Centre, BC Cancer Agency; 2. Department of Cancer Biology, Dana-Farber Cancer Institute; 3. Whitehead Institute for Biomedical Research; 4. Department of Structural Immunology, Institute of Innate Immunity; 5. Pharmaceutical and Medicinal Chemistry, Saarland University; 6. -

Structural and Functional Analysis of the Cdk13/Cyclin K Complex
MASARYK UNIVERSITY FACULTY OF SCIENCE The role of the C-terminal domain of RNA polymerase II in transcriptionally regulated process of genomic instability Ph.D. Dissertation Květa Pilařová Supervisor: Mgr. Dalibor Blažek, Ph.D. Department of biochemistry Brno 2020 Bibliografický záznam Autor: Mgr. Květa Pilařová Přírodovědecká fakulta, Masarykova univerzita Ústav biochemie Název práce: Úloha C-terminální domény RNA polymerasy II v transkripčně regulovaném procesu genomové nestability Studijní program: Biochemie Vedoucí práce: Mgr. Dalibor Blažek, Ph.D. Akademický rok: 2019/2020 Počet stran: 132 Klíčová slova: CDK12, CDK13, kinázová aktivita, analog-senzitivní kináza, C- terminální doména RNAPII, transkripce, genová exprese, přechod mezi G1/S, genomová nestabilita, nádorový biomarker Bibliographic entry Author: Mgr. Květa Pilařová Faculty of Science, Masaryk University Department of biochemistry Title of thesis: The role of the C-terminal domain of RNA polymerase II in transcriptionally regulated process of genomic instability Degree programme: Biochemistry Supervisor: Mgr. Dalibor Blažek, Ph.D. Academic year: 2019/2020 Number of pages: 132 Keywords: CDK12, CDK13, kinase activity, analogue-sensitive kinase, C- terminal domain of RNAPII, transcription, gene expression, G1/S progression, genome instability, tumour biomarker Abstrakt Transkripce protein-kódujících genů je řízena v eukaryotických buňkách RNA polymerázou II (RNAPII). Cyklin-dependentní kinázy 12 a 13 (CDK12 a CDK13) se řadí do skupiny transkripčních CDKs, které asociují s RNAPII při elongaci a fosforylují její C-terminální doménu (CTD). Obě kinázy působí také na expresi genů. Abnormální funkce těchto proteinů jsou u lidských buněk spojovány s různými typy onemocnění a v posledním desetiletí se proto staly předmětem studia výzkumů v oblasti medicíny. Jakým způsobem se CDK12 a CDK13 konkrétně podílí na regulaci transkripce a fosforylačním statusu CTD však není příliš známo. -

Structural Basis and Sequence Rules for Substrate Recognition by Tankyrase Explain the Basis for Cherubism Disease
Structural Basis and Sequence Rules for Substrate Recognition by Tankyrase Explain the Basis for Cherubism Disease Sebastian Guettler,1,2 Jose LaRose,3 Evangelia Petsalaki,1,2 Gerald Gish,1 Andy Scotter,3 Tony Pawson,1,2,* Robert Rottapel,3,4,* and Frank Sicheri1,2,* 1Centre for Systems Biology, Samuel Lunenfeld Research Institute, Mount Sinai Hospital, 600 University Avenue, Toronto, Ontario M5G 1X5, Canada 2Department of Molecular Genetics, University of Toronto, 1 Kings College Circle, Toronto, Ontario M5S 1A8, Canada 3Ontario Cancer Institute and the Campbell Family Cancer Research Institute, 101 College Street, Room 8-703, Toronto Medical Discovery Tower, University of Toronto, Toronto, Ontario M5G 1L7, Canada 4Division of Rheumatology, Department of Medicine, Saint Michael’s Hospital, 30 Bond Street, Toronto, Ontario M5B 1W8, Canada *Correspondence: [email protected] (T.P.), [email protected] (R.R.), [email protected] (F.S.) DOI 10.1016/j.cell.2011.10.046 SUMMARY asparagine, arginine, lysine, cysteine, phosphoserine, and diph- thamide residues (reviewed in Hottiger et al., 2010). As a large The poly(ADP-ribose)polymerases Tankyrase 1/2 posttranslational modification of substantial negative charge, (TNKS/TNKS2) catalyze the covalent linkage of protein poly(ADP-ribosyl)ation (PARsylation) can influence pro- ADP-ribose polymer chains onto target proteins, tein fate through several mechanisms, including a direct effect regulating their ubiquitylation, stability, and function. on protein activity, recruitment of binding partners -

Genome-Wide Screening Identifies Genes and Biological Processes
Louisiana State University LSU Digital Commons LSU Doctoral Dissertations Graduate School 10-12-2018 Genome-Wide Screening Identifies Genes and Biological Processes Implicated in Chemoresistance and Oncogene-Induced Apoptosis Tengyu Ko Louisiana State University and Agricultural and Mechanical College, [email protected] Follow this and additional works at: https://digitalcommons.lsu.edu/gradschool_dissertations Part of the Cancer Biology Commons, Cell Biology Commons, and the Genomics Commons Recommended Citation Ko, Tengyu, "Genome-Wide Screening Identifies Genes and Biological Processes Implicated in Chemoresistance and Oncogene- Induced Apoptosis" (2018). LSU Doctoral Dissertations. 4715. https://digitalcommons.lsu.edu/gradschool_dissertations/4715 This Dissertation is brought to you for free and open access by the Graduate School at LSU Digital Commons. It has been accepted for inclusion in LSU Doctoral Dissertations by an authorized graduate school editor of LSU Digital Commons. For more information, please [email protected]. GENOME-WIDE SCREENING IDENTIFIES GENES AND BIOLOGICAL PROCESSES IMPLICATED IN CHEMORESISTANCE AND ONCOGENE- INDUCED APOPTOSIS A Dissertation Submitted to the Graduate Faculty of the Louisiana State University and Agricultural and Mechanical College in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical and Veterinary Medical Sciences through the Department of Comparative Biomedical Sciences by Tengyu Ko B.S., University of California, Santa Barbara 2010 December 2018 ACKNOWLEDGEMENTS I would like to express my sincerest gratitude to my major supervisor Dr. Shisheng Li for giving me the opportunity to join his team and the freedom to pursue projects. I appreciate all of his thoughts and efforts. Truly, none of these findings would be possible without his supervisions, supports, insightful discussions, and patience. -

Using Network Inference to Discover Molecular Pathways Underlying Cytokine Synergism and Age-Related Neurodegeneration by Bryce Hwang S.B., M.I.T
Using network inference to discover molecular pathways underlying cytokine synergism and age-related neurodegeneration by Bryce Hwang S.B., M.I.T. (2018) Submitted to the Department of Electrical Engineering and Computer Science in partial fulfillment of the requirements for the degree of Master of Engineering in Computer Science and Molecular Biology at the MASSACHUSETTS INSTITUTE OF TECHNOLOGY June 2018 ○c Bryce Hwang. All rights reserved. The author hereby grants to M.I.T. permission to reproduce and to distribute publicly paper and electronic copies of this thesis document in whole or in part in any medium now known or hereafter created. Author.............................................................. Department of Electrical Engineering and Computer Science May 25, 2018 Certified by. Ernest Fraenkel Professor Thesis Supervisor Accepted by . Katrina LaCurts Chair, Master of Engineering Thesis Committee Using network inference to discover molecular pathways underlying cytokine synergism and age-related neurodegeneration by Bryce Hwang Submitted to the Department of Electrical Engineering and Computer Science on May 25, 2018, in partial fulfillment of the requirements for the degree of Master of Engineering in Computer Science and Molecular Biology Abstract New high-throughput “omic” methods can help shed light on molecular pathways underpinning diseases ranging from cancers to neurodegenerative disorders. However, effectively integrating information across these diverse data types is challenging. Network modeling approaches can help bridge this gap. In particular, the Prize- Collecting Steiner Forest approach (PCSF) is a network modeling method that provides high-confidence subnetworks of physically interacting molecules by integrating diverse “omics” data with prior knowledge from protein-protein interaction networks (PPIs). However, PCSF is sensitive to initial parameterization and generating biological hypotheses from the resulting subnetworks can often be difficult.