(12) Patent Application Publication (10) Pub. No.: US 2011/0086407 A1 Berka Et Al
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

The Structure of an Authentic Spore Photoproduct Lesion in DNA Suggests a Basis for Recognition
addenda and errata Acta Crystallographica Section D Biological addenda and errata Crystallography ISSN 1399-0047 The structure of an authentic spore photoproduct lesion in DNA suggests a basis for recognition. Corrigendum Isha Singh,a Yajun Jian,b Lei Lia,b and Millie M. Georgiadisa,b* aDepartment of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, IN 46202, USA, and bDepartment of Chemistry and Chemical Biology, Indiana University–Purdue University at Indianapolis, Indianapolis, IN 46202, USA Correspondence e-mail: [email protected] The article by Singh et al. [ (2014). Acta Cryst. D70, 752–759] is corrected. In the article by Singh et al. (2014) the name of one of the authors was given incorrectly. The correct name is Yajun Jian as given above. References Singh, I., Lian, Y., Li, L. & Georgiadis, M. M. (2014). Acta Cryst. D70, 752–759. Acta Cryst. (2014). D70, 1173 doi:10.1107/S1399004714006130 # 2014 International Union of Crystallography 1173 research papers Acta Crystallographica Section D Biological The structure of an authentic spore photoproduct Crystallography lesion in DNA suggests a basis for recognition ISSN 1399-0047 Isha Singh,a Yajun Lian,b Lei Lia,b The spore photoproduct lesion (SP; 5-thymine-5,6-dihydro- Received 12 September 2013 and Millie M. Georgiadisa,b* thymine) is the dominant photoproduct found in UV- Accepted 5 December 2013 irradiated spores of some bacteria such as Bacillus subtilis. Upon spore germination, this lesion is repaired in a light- PDB references: N-terminal aDepartment of Biochemistry and Molecular independent manner by a specific repair enzyme: the spore fragment of MMLV RT, SP Biology, Indiana University School of Medicine, DNA complex, 4m94; non-SP Indianapolis, IN 46202, USA, and bDepartment photoproduct lyase (SP lyase). -

Letters to Nature
letters to nature Received 7 July; accepted 21 September 1998. 26. Tronrud, D. E. Conjugate-direction minimization: an improved method for the re®nement of macromolecules. Acta Crystallogr. A 48, 912±916 (1992). 1. Dalbey, R. E., Lively, M. O., Bron, S. & van Dijl, J. M. The chemistry and enzymology of the type 1 27. Wolfe, P. B., Wickner, W. & Goodman, J. M. Sequence of the leader peptidase gene of Escherichia coli signal peptidases. Protein Sci. 6, 1129±1138 (1997). and the orientation of leader peptidase in the bacterial envelope. J. Biol. Chem. 258, 12073±12080 2. Kuo, D. W. et al. Escherichia coli leader peptidase: production of an active form lacking a requirement (1983). for detergent and development of peptide substrates. Arch. Biochem. Biophys. 303, 274±280 (1993). 28. Kraulis, P.G. Molscript: a program to produce both detailed and schematic plots of protein structures. 3. Tschantz, W. R. et al. Characterization of a soluble, catalytically active form of Escherichia coli leader J. Appl. Crystallogr. 24, 946±950 (1991). peptidase: requirement of detergent or phospholipid for optimal activity. Biochemistry 34, 3935±3941 29. Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and (1995). the thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281±296 (1991). 4. Allsop, A. E. et al.inAnti-Infectives, Recent Advances in Chemistry and Structure-Activity Relationships 30. Meritt, E. A. & Bacon, D. J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505± (eds Bently, P. H. & O'Hanlon, P. J.) 61±72 (R. Soc. Chem., Cambridge, 1997). -

A Phosphopantetheinylating Polyketide Synthase Producing a Linear Polyene to Initiate Enediyne Antitumor Antibiotic Biosynthesis
A phosphopantetheinylating polyketide synthase producing a linear polyene to initiate enediyne antitumor antibiotic biosynthesis Jian Zhang†, Steven G. Van Lanen‡, Jianhua Ju‡, Wen Liu‡, Pieter C. Dorrestein§, Wenli Li‡, Neil L. Kelleher§, and Ben Shen†‡¶ʈ †Department of Chemistry, ‡Division of Pharmaceutical Sciences, and ¶University of Wisconsin National Cooperative Drug Discovery Group, University of Wisconsin, Madison, WI 53705; and §Department of Chemistry, University of Illinois at Urbana–Champaign, Urbana, IL 61801 Communicated by Arnold L. Demain, Drew University, Madison, NJ, December 11, 2007 (received for review November 29, 2007) The enediynes, unified by their unique molecular architecture and can have anywhere from all to none of these activities, resulting mode of action, represent some of the most potent anticancer drugs in a product with various levels of oxidation. ever discovered. The biosynthesis of the enediyne core has been Bacterial PKSs are often categorized by domain architecture predicted to be initiated by a polyketide synthase (PKS) that is distinct and the iterative or noniterative nature during polyketide for- from all known PKSs. Characterization of the enediyne PKS involved mation, and this classification has led to three groupings (17). in C-1027 (SgcE) and neocarzinostatin (NcsE) biosynthesis has now Noniterative type I PKS are large proteins with multiple domains revealed that (i) the PKSs contain a central acyl carrier protein domain forming modules, a minimal module consisting of a KS, AT, and and C-terminal phosphopantetheinyl transferase domain; (ii) the PKSs ACP domain, wherein each domain catalyzes a single event are functional in heterologous hosts, and coexpression with an during polyketide biosynthesis (18). -

The Microbiota-Produced N-Formyl Peptide Fmlf Promotes Obesity-Induced Glucose
Page 1 of 230 Diabetes Title: The microbiota-produced N-formyl peptide fMLF promotes obesity-induced glucose intolerance Joshua Wollam1, Matthew Riopel1, Yong-Jiang Xu1,2, Andrew M. F. Johnson1, Jachelle M. Ofrecio1, Wei Ying1, Dalila El Ouarrat1, Luisa S. Chan3, Andrew W. Han3, Nadir A. Mahmood3, Caitlin N. Ryan3, Yun Sok Lee1, Jeramie D. Watrous1,2, Mahendra D. Chordia4, Dongfeng Pan4, Mohit Jain1,2, Jerrold M. Olefsky1 * Affiliations: 1 Division of Endocrinology & Metabolism, Department of Medicine, University of California, San Diego, La Jolla, California, USA. 2 Department of Pharmacology, University of California, San Diego, La Jolla, California, USA. 3 Second Genome, Inc., South San Francisco, California, USA. 4 Department of Radiology and Medical Imaging, University of Virginia, Charlottesville, VA, USA. * Correspondence to: 858-534-2230, [email protected] Word Count: 4749 Figures: 6 Supplemental Figures: 11 Supplemental Tables: 5 1 Diabetes Publish Ahead of Print, published online April 22, 2019 Diabetes Page 2 of 230 ABSTRACT The composition of the gastrointestinal (GI) microbiota and associated metabolites changes dramatically with diet and the development of obesity. Although many correlations have been described, specific mechanistic links between these changes and glucose homeostasis remain to be defined. Here we show that blood and intestinal levels of the microbiota-produced N-formyl peptide, formyl-methionyl-leucyl-phenylalanine (fMLF), are elevated in high fat diet (HFD)- induced obese mice. Genetic or pharmacological inhibition of the N-formyl peptide receptor Fpr1 leads to increased insulin levels and improved glucose tolerance, dependent upon glucagon- like peptide-1 (GLP-1). Obese Fpr1-knockout (Fpr1-KO) mice also display an altered microbiome, exemplifying the dynamic relationship between host metabolism and microbiota. -

Proteome Analysis of Peroxisomes from Dark-Treated Senescent Arabidopsis Leavesfa
Journal of Integrative JIPB Plant Biology Proteome analysis of peroxisomes from dark-treated senescent Arabidopsis leavesFA † † Ronghui Pan1 , Sigrun Reumann1,2,3,4 , Piotr Lisik3, Stefanie Tietz1, Laura J. Olsen2 and Jianping Hu1,5* 1. MSU-Department of Energy Plant Research Laboratory, Michigan State University, East Lansing, MI 48824, USA 2. Department of Molecular, Cellular, and Developmental Biology, University of Michigan, Ann Arbor, MI, USA 3. Center of Organelle Research, University of Stavanger, N-4021 Stavanger, Norway 4. Department of Plant Biochemistry and Infection Biology, Institute of Plant Science and Microbiology, University of Hamburg, D-22609 Hamburg, Germany 5. Plant Biology Department, Michigan State University, East Lansing, MI 48824, USA † These authors contributed equally to this work *Correspondence: Jianping Hu ([email protected]) doi: 10.1111/jipb.12670 High-Impact Article Abstract Peroxisomes compartmentalize a dynamic suite of a total of 111 peroxisomal proteins and verified the peroxisomal biochemical reactions and play a central role in plant localization for six new proteins with potential roles in fatty metabolism, such as the degradation of hydrogen peroxide, acid metabolism and stress response by in vivo targeting metabolism of fatty acids, photorespiration, and the biosyn- analysis. Metabolic pathways compartmentalized in the three thesis of plant hormones. Plant peroxisomes have been major subtypes of peroxisomes were also compared, which traditionally classified into three major subtypes, and in-depth revealed a higher number of proteins involved in the mass spectrometry (MS)-based proteomics has been per- detoxification of reactive oxygen species in peroxisomes formed to explore the proteome of the two major subtypes from senescent leaves. -

Radical SAM Enzymes in the Biosynthesis of Ribosomally Synthesized and Post-Translationally Modified Peptides (Ripps) Alhosna Benjdia, Clémence Balty, Olivier Berteau
Radical SAM enzymes in the biosynthesis of ribosomally synthesized and post-translationally modified peptides (RiPPs) Alhosna Benjdia, Clémence Balty, Olivier Berteau To cite this version: Alhosna Benjdia, Clémence Balty, Olivier Berteau. Radical SAM enzymes in the biosynthesis of ribosomally synthesized and post-translationally modified peptides (RiPPs). Frontiers in Chemistry, Frontiers Media, 2017, 5, 10.3389/fchem.2017.00087. hal-02627786 HAL Id: hal-02627786 https://hal.inrae.fr/hal-02627786 Submitted on 26 May 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License REVIEW published: 08 November 2017 doi: 10.3389/fchem.2017.00087 Radical SAM Enzymes in the Biosynthesis of Ribosomally Synthesized and Post-translationally Modified Peptides (RiPPs) Alhosna Benjdia*, Clémence Balty and Olivier Berteau* Micalis Institute, ChemSyBio, INRA, AgroParisTech, Université Paris-Saclay, Jouy-en-Josas, France Ribosomally-synthesized and post-translationally modified peptides (RiPPs) are a large and diverse family of natural products. They possess interesting biological properties such as antibiotic or anticancer activities, making them attractive for therapeutic applications. In contrast to polyketides and non-ribosomal peptides, RiPPs derive from ribosomal peptides and are post-translationally modified by diverse enzyme families. -

Neocarzinostatin Naphthoate Synthase: an Unique Iterative Type I PKS from Neocarzinostatin Producer Streptomyces Carzinostaticus
FEBS 28398 FEBS Letters 566 (2004) 201–206 Neocarzinostatin naphthoate synthase: an unique iterative type I PKS from neocarzinostatin producer Streptomyces carzinostaticus Basundhara Sthapita, Tae-Jin Oha, Rajan Lamichhanea, Kwangkyoung Lioua, Hei Chan Leea, Chun-Gyu Kimb, Jae Kyung Sohnga,* aInstitute of Biomolecule Reconstruction (iBR), Department of Chemistry, Sun Moon University, #100, Kalsan-ri, Tangjeong-myeon, Asansi, Chung-Nam 336-708, Republic of Korea bDepartment of Pharmaceutical Engineering, Inje University, 607 Obang-dong, Kimhae City, Kyungnam 621-749, Republic of Korea Received 13 March 2004; revised 7 April 2004; accepted 7 April 2004 Available online 28 April 2004 Edited by Stuart Ferguson ketide synthase (PKS) catalyzes a decarboxylative condensa- Abstract Enediyne antibiotics are known for their potent antitumor activities. One such enediyne, neocarzinostatin tion of these compounds into the growing polyketide chain. (NCS), consists of a 1:1 complex of non-peptide chromophore Microbial PKSs are classified into three major types. A type I (1a), and peptide apoprotein. The structurally diverse non- PKS system consists of one or more multifunctional proteins peptide chromophore is responsible for its biological activity. that contain a different active site for each enzyme-catalyzed One of its structural components, the naphthoic acid moiety (2,7- reaction in polyketide carbon chain assembly and modification dihydroxy-5-methyl-1-naphthoic acid, 1d) is synthesized by a [1]. The macrolides of erythromycin and avermectin are ex- polyketide synthase (PKS) pathway through condensing six amples of type I PKS products. Type II PKSs, on the other intact acetate units. The 5.45 kb iterative type I PKS, hand, comprise sets of iteratively used individual proteins for neocarzinostatin naphthoate synthase (NNS), responsible for the production of multicyclic aromatic compounds (e.g., ac- naphthoic acid moiety biosynthesis, shares sequence homology tinorhodin and tetrocenomycin) [2]. -

(MNQ) Biosynthesis in Impatiens Balsamina L. Lian Chee Foong1,2, Jian Yi Chai1, Anthony Siong Hock Ho1, Brandon Pei Hui Yeo3, Yang Mooi Lim4 & Sheh May Tam1*
www.nature.com/scientificreports OPEN Comparative transcriptome analysis to identify candidate genes involved in 2‑methoxy‑1,4‑naphthoquinone (MNQ) biosynthesis in Impatiens balsamina L. Lian Chee Foong1,2, Jian Yi Chai1, Anthony Siong Hock Ho1, Brandon Pei Hui Yeo3, Yang Mooi Lim4 & Sheh May Tam1* Impatiens balsamina L. is a tropical ornamental and traditional medicinal herb rich in natural compounds, especially 2‑methoxy‑1,4‑naphthoquinone (MNQ) which is a bioactive compound with tested anticancer activities. Characterization of key genes involved in the shikimate and 1,4‑dihydroxy‑2‑naphthoate (DHNA) pathways responsible for MNQ biosynthesis and their expression profles in I. balsamina will facilitate adoption of genetic/metabolic engineering or synthetic biology approaches to further increase production for pre‑commercialization. In this study, HPLC analysis showed that MNQ was present in signifcantly higher quantities in the capsule pericarps throughout three developmental stages (early‑, mature‑ and postbreaker stages) whilst its immediate precursor, 2‑hydroxy‑1,4‑naphthoquinone (lawsone) was mainly detected in mature leaves. Transcriptomes of I. balsamina derived from leaf, fower, and three capsule developmental stages were generated, totalling 59.643 Gb of raw reads that were assembled into 94,659 unigenes (595,828 transcripts). A total of 73.96% of unigenes were functionally annotated against seven public databases and 50,786 diferentially expressed genes (DEGs) were identifed. Expression profles of 20 selected genes from four major secondary metabolism pathways were studied and validated using qRT‑PCR method. Majority of the DHNA pathway genes were found to be signifcantly upregulated in early stage capsule compared to fower and leaf, suggesting tissue‑specifc synthesis of MNQ. -
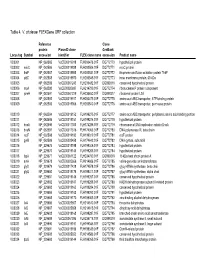
Table 4. V. Cholerae Flexgene ORF Collection
Table 4. V. cholerae FLEXGene ORF collection Reference Clone protein PlasmID clone GenBank Locus tag Symbol accession identifier FLEX clone name accession Product name VC0001 NP_062585 VcCD00019918 FLH200476.01F DQ772770 hypothetical protein VC0002 mioC NP_062586 VcCD00019938 FLH200506.01F DQ772771 mioC protein VC0003 thdF NP_062587 VcCD00019958 FLH200531.01F DQ772772 thiophene and furan oxidation protein ThdF VC0004 yidC NP_062588 VcCD00019970 FLH200545.01F DQ772773 inner membrane protein, 60 kDa VC0005 NP_062589 VcCD00061243 FLH236482.01F DQ899316 conserved hypothetical protein VC0006 rnpA NP_062590 VcCD00025697 FLH214799.01F DQ772774 ribonuclease P protein component VC0007 rpmH NP_062591 VcCD00061229 FLH236450.01F DQ899317 ribosomal protein L34 VC0008 NP_062592 VcCD00019917 FLH200475.01F DQ772775 amino acid ABC transporter, ATP-binding protein VC0009 NP_062593 VcCD00019966 FLH200540.01F DQ772776 amino acid ABC transproter, permease protein VC0010 NP_062594 VcCD00019152 FLH199275.01F DQ772777 amino acid ABC transporter, periplasmic amino acid-binding portion VC0011 NP_062595 VcCD00019151 FLH199274.01F DQ772778 hypothetical protein VC0012 dnaA NP_062596 VcCD00017363 FLH174286.01F DQ772779 chromosomal DNA replication initiator DnaA VC0013 dnaN NP_062597 VcCD00017316 FLH174063.01F DQ772780 DNA polymerase III, beta chain VC0014 recF NP_062598 VcCD00019182 FLH199319.01F DQ772781 recF protein VC0015 gyrB NP_062599 VcCD00025458 FLH174642.01F DQ772782 DNA gyrase, subunit B VC0016 NP_229675 VcCD00019198 FLH199346.01F DQ772783 hypothetical protein -

PURDUE UNIVERSITY GRADUATE SCHOOL Thesis/Dissertation Acceptance
Graduate School ETD Form 9 (Revised 12/07) PURDUE UNIVERSITY GRADUATE SCHOOL Thesis/Dissertation Acceptance This is to certify that the thesis/dissertation prepared By Renae Nelson Entitled Exploring the Mechanism of Action of Spore Photoproduct Lyase Master of Science For the degree of Is approved by the final examining committee: Dr. Lei Li Chair Dr. Eric Long Dr. Michael McLeish To the best of my knowledge and as understood by the student in the Research Integrity and Copyright Disclaimer (Graduate School Form 20), this thesis/dissertation adheres to the provisions of Purdue University’s “Policy on Integrity in Research” and the use of copyrighted material. Approved by Major Professor(s): ____________________________________Dr. Lei Li ____________________________________ Approved by: Dr. Eric Long 11/20/2013 Head of the Graduate Program Date i EXPLORING THE MECHANISM OF ACTION OF SPORE PHOTOPRODUCT LYASE A Thesis Submitted to the Faculty of Purdue University by Renae Nelson In Partial Fulfillment of the Requirements for the Degree of Master of Science i December 2013 Purdue University Indianapolis, Indiana ii To Steven, thank you for holding my hand as we’ve grown up together. To my children, Adelene and Calvin, thank you for motivating me to be the best person I can be. To my Momma, for making me think that a master’s degree made you the smartest person in the world. ii iii ACKNOWLEDGEMENTS I would like to thank Dr. Lei Li for providing me with the opportunity to challenge and develop my skills as a research scientist, and his constant patience with my failures as well as my successes. -

INVASIVE PLANKTON Implications of and for Ballast Water Management
INVASIVE PLANKTON Implications of and for ballast water management Dissertation Zur Erlangung der Würde des Doktors der Naturwissenschaften des Fachbereichs Biologie, der Fakultät für Mathematik, Informatik und Naturwissenschaften, der Universität Hamburg vorgelegt von Viola Liebich aus Berlin Hamburg, Dezember 2012 Title: INVASIVE PLANKTON - implications of and for ballast water management Contents: Chapter 1. Introduction: Invasive species and ballast water 4 Chapter 2. Understanding (marine) invasions through the application of a comprehensive stage-transition framework – review of invasion theory and terminology 15 Chapter 3. Re-growth of potential invasive phytoplankton following UV-based ballast water treatment 32 Chapter 4. Incubation experiments after ballast water treatment: focus on the forgotten fraction of organisms smaller than 10 µm 47 Chapter 5. Overall Discussion, Conclusion and Perspective 66 List of publications 78 Acknowledgement 79 Summary in English 80 Summary in German 82 Declaration on oath 85 Annex: Stehouwer PP, Liebich V, Peperzak L (2012) Flow cytometry, microscopy, and DNA analysis as complementary phytoplankton screening methods in ballast water treatment studies. Journal of Applied Phycology 86 2 Chapter 1. Introduction: Invasive species and ballast water Invasive species Invasive species are considered one of the biggest threats to our world’s oceans (in addition to marine pollution and overexploitation). The man-aided introduction of non- native organisms via a vector (transport element) into new areas and their successful establishment as invasive species pose risks to native biodiversity and habitats (Zaiko et al. 2011). Commonly recognized is the phenomenon of competitive exclusion of native populations by invasive species (Huxel 1999), with increasing probability due to climate change (Philippart et al. -

Diphthamide Biosynthesis: Characterization and Mechanistic Studies of an Unconventional Radical Sam Enzyme Phdph2
DIPHTHAMIDE BIOSYNTHESIS: CHARACTERIZATION AND MECHANISTIC STUDIES OF AN UNCONVENTIONAL RADICAL SAM ENZYME PHDPH2 A Dissertation Presented to the Faculty of the Graduate School of Cornell University In Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy by Xuling Zhu January 2011 © 2011 Xuling Zhu DIPHTHAMIDE BIOSYNTHESIS: CHARACTERIZATION AND MECHANISTIC STUDIES OF AN UNCONVENTIONAL RADICAL SAM ENZYME PHDPH2 Xuling Zhu, Ph. D. Cornell University 2011 Diphthamide, the target of diphtheria toxin, is a unique posttranslational modification on eukaryotic and archaeal translation elongation factor 2 (EF2). The proposed biosynthesis of diphthamide involves three steps. The first step is the formation of a C-C bond between the histidine residue and the 3-amino-3- carboxylpropyl group of S-adenosylmethionine (SAM), which is catalyzed by four enzymes Dph1-Dph4 in eukaryotic or only one enzyme Dph2 in archaea; the second step is the trimethylation of the amino group by Dph5; and the last step is an ATP depended amidation of the carboxyl group by an unknown enzyme. We have recently found that in an archaeal species Pyrococcus horikoshii (P. horikoshii), the first step uses an S-adenosyl-L-methionine (SAM)-dependent [4Fe– 4S] enzyme, PhDph2, to catalyze the formation of a C–C bond. Crystal structure shows that PhDph2 is a homodimer and each monomer contains three conserved cysteine residues that can bind a [4Fe–4S] cluster. In the reduced state, the [4Fe–4S] cluster can provide one electron to reductively cleave the bound SAM molecule. However, different from classical radical SAM enzymes, biochemical evidence suggests that a 3-amino-3-carboxypropyl radical is generated in PhDph2.