Accepted Manuscript
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Department of Planning and Zoning
Department of Planning and Zoning Subject: Howard County Landscape Manual Updates: Recommended Street Tree List (Appendix B) and Recommended Plant List (Appendix C) - Effective July 1, 2010 To: DLD Review Staff Homebuilders Committee From: Kent Sheubrooks, Acting Chief Division of Land Development Date: July 1, 2010 Purpose: The purpose of this policy memorandum is to update the Recommended Plant Lists presently contained in the Landscape Manual. The plant lists were created for the first edition of the Manual in 1993 before information was available about invasive qualities of certain recommended plants contained in those lists (Norway Maple, Bradford Pear, etc.). Additionally, diseases and pests have made some other plants undesirable (Ash, Austrian Pine, etc.). The Howard County General Plan 2000 and subsequent environmental and community planning publications such as the Route 1 and Route 40 Manuals and the Green Neighborhood Design Guidelines have promoted the desirability of using native plants in landscape plantings. Therefore, this policy seeks to update the Recommended Plant Lists by identifying invasive plant species and disease or pest ridden plants for their removal and prohibition from further planting in Howard County and to add other available native plants which have desirable characteristics for street tree or general landscape use for inclusion on the Recommended Plant Lists. Please note that a comprehensive review of the street tree and landscape tree lists were conducted for the purpose of this update, however, only -

Likely to Have Habitat Within Iras That ALLOW Road
Item 3a - Sensitive Species National Master List By Region and Species Group Not likely to have habitat within IRAs Not likely to have Federal Likely to have habitat that DO NOT ALLOW habitat within IRAs Candidate within IRAs that DO Likely to have habitat road (re)construction that ALLOW road Forest Service Species Under NOT ALLOW road within IRAs that ALLOW but could be (re)construction but Species Scientific Name Common Name Species Group Region ESA (re)construction? road (re)construction? affected? could be affected? Bufo boreas boreas Boreal Western Toad Amphibian 1 No Yes Yes No No Plethodon vandykei idahoensis Coeur D'Alene Salamander Amphibian 1 No Yes Yes No No Rana pipiens Northern Leopard Frog Amphibian 1 No Yes Yes No No Accipiter gentilis Northern Goshawk Bird 1 No Yes Yes No No Ammodramus bairdii Baird's Sparrow Bird 1 No No Yes No No Anthus spragueii Sprague's Pipit Bird 1 No No Yes No No Centrocercus urophasianus Sage Grouse Bird 1 No Yes Yes No No Cygnus buccinator Trumpeter Swan Bird 1 No Yes Yes No No Falco peregrinus anatum American Peregrine Falcon Bird 1 No Yes Yes No No Gavia immer Common Loon Bird 1 No Yes Yes No No Histrionicus histrionicus Harlequin Duck Bird 1 No Yes Yes No No Lanius ludovicianus Loggerhead Shrike Bird 1 No Yes Yes No No Oreortyx pictus Mountain Quail Bird 1 No Yes Yes No No Otus flammeolus Flammulated Owl Bird 1 No Yes Yes No No Picoides albolarvatus White-Headed Woodpecker Bird 1 No Yes Yes No No Picoides arcticus Black-Backed Woodpecker Bird 1 No Yes Yes No No Speotyto cunicularia Burrowing -

Indiana's Native Magnolias
FNR-238 Purdue University Forestry and Natural Resources Know your Trees Series Indiana’s Native Magnolias Sally S. Weeks, Dendrologist Department of Forestry and Natural Resources Purdue University, West Lafayette, IN 47907 This publication is available in color at http://www.ces.purdue.edu/extmedia/fnr.htm Introduction When most Midwesterners think of a magnolia, images of the grand, evergreen southern magnolia (Magnolia grandiflora) (Figure 1) usually come to mind. Even those familiar with magnolias tend to think of them as occurring only in the South, where a more moderate climate prevails. Seven species do indeed thrive, especially in the southern Appalachian Mountains. But how many Hoosiers know that there are two native species Figure 2. Cucumber magnolia when planted will grow well throughout Indiana. In Charles Deam’s Trees of Indiana, the author reports “it doubtless occurred in all or nearly all of the counties in southern Indiana south of a line drawn from Franklin to Knox counties.” It was mainly found as a scattered, woodland tree and considered very local. Today, it is known to occur in only three small native populations and is listed as State Endangered Figure 1. Southern magnolia by the Division of Nature Preserves within Indiana’s Department of Natural Resources. found in Indiana? Very few, I suspect. No native As the common name suggests, the immature magnolias occur further west than eastern Texas, fruits are green and resemble a cucumber so we “easterners” are uniquely blessed with the (Figure 3). Pioneers added the seeds to whisky presence of these beautiful flowering trees. to make bitters, a supposed remedy for many Indiana’s most “abundant” species, cucumber ailments. -

Download PCN Magnolia Multisite
Institution name plant NAMES for inventory::print name Accession # Provenanc Quantity Plant source The Scott Arboretum atMagnolia Swarthmore acuminata College 2005-355UN*A G 1 Unknown The Scott Arboretum atMagnolia Swarthmore acuminata College 2001-188UN*A U 1 Unknown The Scott Arboretum atMagnolia Swarthmore acuminata College 96-129*A G 1 Princeton Nurseries The Scott Arboretum atMagnolia Swarthmore acuminata College var. subcordata 99-203*B G 1 Longwood Gardens The Scott Arboretum atMagnolia Swarthmore acuminata College var. subcordata 93-206*A G 1 Woodlanders Nursery The Scott Arboretum atMagnolia Swarthmore acuminata College var. subcordata 'Brenda'2004-239*A G 1 Pat McCracken The Scott Arboretum atMagnolia Swarthmore 'Anilou' College 2008-202*A G 1 Pleasant Run Nursery The Scott Arboretum atMagnolia Swarthmore 'Anilou' College 2008-202*B G 1 Pleasant Run Nursery The Scott Arboretum atMagnolia Swarthmore 'Ann' College 68-165*A G 1 U. S. National Arboretum, Washington, DC The Scott Arboretum atMagnolia Swarthmore 'Banana College Split' 2004-237*A G 1 Pat McCracken The Scott Arboretum atMagnolia Swarthmore 'Betty' College 68-166*A G 1 U. S. National Arboretum, Washington, DC The Scott Arboretum atMagnolia Swarthmore 'Big Dude' College 2008-203*A G 1 Pleasant Run Nursery The Scott Arboretum atMagnolia Swarthmore ×brooklynensis College 'Black Beauty' 2008-204*A G 1 Pleasant Run Nursery The Scott Arboretum atMagnolia Swarthmore ×soulangeana College 'Jurmag1' 2010-069*A G 1 Pleasant Run Nursery The Scott Arboretum atMagnolia Swarthmore -

IAWA Journal, Vol 14 (4), 1993: 391-412 WOOD ANATOMY OF
IAWA Journal, VoL 14 (4), 1993: 391-412 WOOD ANATOMY OF TREES AND SIffiUBS FROM CHINA. VI. MAGNOLIACEAE by Chen Bao Liangt 1, Pieter Baas2, Elisabeth A. Wheeler3 and Wu Shuming4 Summary The wood anatomy offive genera of Mag ces; oil cells in rays mostly occur in the taxa noliaceae (59 native species, 2 introduced from tropical and subtropical provenances. species) of China is described. Although the Simple perforation plates are mostly present wood anatomy of this family is rather homo in the temperate taxa. Counter to trends for geneous, it is possible to identify most speci the dicotyledons at large, helical thickenings mens to genus. Magnoliaceae wood from are more common in tropical species than in China is characterised by diffuse-porosity, temperate species, and, when present, are scalariform to opposite vessel wall pitting, usually not distinct in deciduous species. scalariform perforations with few bars or in Key words: Magnoliaceae, Kmeria, Lirio some Magnolia species simple perforations, dendron, Magnolia, Mangiietia, Michelia, ground tissue fibres with distinctly to minute China, systematic wood anatomy, ecologi ly bordered pits, marginal parenchyma and cal wood anatomy, wood identification. heterocellular rays mostly with one marginal row of square/upright cells. Intervessel and vessel-parenchyma pits are almost exclusive Introduction ly opposite in the Liriodendroideae; they are Magnoliaceae are shrubs, small trees, or almost exclusively scalariform in the Magno trees, and range from tropical and subtropical lioideae, except for Magnolia section Rhyti to temperate areas. In China, most Magnolia dospermum in which pits are predominantly ceae occur in tropical to subtropical forests, a opposite. Although the wood anatomical char few species occur in temperate areas. -

Vascular Flora of the Possum Walk Trail at the Infinity Science Center, Hancock County, Mississippi
The University of Southern Mississippi The Aquila Digital Community Honors Theses Honors College Spring 5-2016 Vascular Flora of the Possum Walk Trail at the Infinity Science Center, Hancock County, Mississippi Hanna M. Miller University of Southern Mississippi Follow this and additional works at: https://aquila.usm.edu/honors_theses Part of the Biodiversity Commons, and the Botany Commons Recommended Citation Miller, Hanna M., "Vascular Flora of the Possum Walk Trail at the Infinity Science Center, Hancock County, Mississippi" (2016). Honors Theses. 389. https://aquila.usm.edu/honors_theses/389 This Honors College Thesis is brought to you for free and open access by the Honors College at The Aquila Digital Community. It has been accepted for inclusion in Honors Theses by an authorized administrator of The Aquila Digital Community. For more information, please contact [email protected]. The University of Southern Mississippi Vascular Flora of the Possum Walk Trail at the Infinity Science Center, Hancock County, Mississippi by Hanna Miller A Thesis Submitted to the Honors College of The University of Southern Mississippi in Partial Fulfillment of the Requirement for the Degree of Bachelor of Science in the Department of Biological Sciences May 2016 ii Approved by _________________________________ Mac H. Alford, Ph.D., Thesis Adviser Professor of Biological Sciences _________________________________ Shiao Y. Wang, Ph.D., Chair Department of Biological Sciences _________________________________ Ellen Weinauer, Ph.D., Dean Honors College iii Abstract The North American Coastal Plain contains some of the highest plant diversity in the temperate world. However, most of the region has remained unstudied, resulting in a lack of knowledge about the unique plant communities present there. -

A Critically Endangered Plant Species Endemic to South-West China
Integrated conservation for Parakmeria omeiensis (Magnoliaceae), a Critically Endangered plant species endemic to south-west China D AOPING Y U ,XIANGYING W EN,CEHONG L I ,TIEYI X IONG,QIXIN P ENG X IAOJIE L I ,KONGPING X IE,HONG L IU and H AI R EN Abstract Parakmeria omeiensis is a Critically Endangered tree attractive and large, and their seed arils are orange, making species in the family Magnoliaceae, endemic to south-west the tree an attractive ornamental plant. However, the species China. The tree is functionally dioecious, but little is known has a restricted range. It has been considered a Grade-I about the species’ status in the wild. We investigated the Key-Protected Wild Plant Species in China since and range, population size, age structure, habitat characteristics has been categorized as Critically Endangered on the and threats to P. omeiensis. We located a total of individuals IUCN Red List since (China Expert Workshop, ), in two populations on the steep slopes of Mount Emei, Sichuan the Chinese Higher Plants Red List since (Yin, ), and province, growing under the canopy of evergreen broadleaved the Red List of Magnoliaceae since (Malin et al., ). forest in well-drained gravel soil. A male-biased sex ratio, lack The tree has also been identified as a plant species with an of effective pollinating insects, and habitat destruction result extremely small population (Ren et al., ; State Forestry in low seed set and poor seedling survival in the wild. We Administration of China, ). have adopted an integrated conservation approach, including Parakmeria includes five species (P. -

THE Magnoliaceae Liriodendron L. Magnolia L
THE Magnoliaceae Liriodendron L. Magnolia L. VEGETATIVE KEY TO SPECIES IN CULTIVATION Jan De Langhe (1 October 2014 - 28 May 2015) Vegetative identification key. Introduction: This key is based on vegetative characteristics, and therefore also of use when flowers and fruits are absent. - Use a 10× hand lens to evaluate stipular scars, buds and pubescence in general. - Look at the entire plant. Young specimens, shade, and strong shoots give an atypical view. - Beware of hybridisation, especially with plants raised from seed other than wild origin. Taxa treated in this key: see page 10. Questionable/frequently misapplied names: see page 10. Names referred to synonymy: see page 11. References: - JDL herbarium - living specimens, in various arboreta, botanic gardens and collections - literature: De Meyere, D. - (2001) - Enkele notities omtrent Liriodendron tulipifera, L. chinense en hun hybriden in BDB, p.23-40. Hunt, D. - (1998) - Magnolias and their allies, 304p. Bean, W.J. - (1981) - Magnolia in Trees and Shrubs hardy in the British Isles VOL.2, p.641-675. - or online edition Clarke, D.L. - (1988) - Magnolia in Trees and Shrubs hardy in the British Isles supplement, p.318-332. Grimshaw, J. & Bayton, R. - (2009) - Magnolia in New Trees, p.473-506. RHS - (2014) - Magnolia in The Hillier Manual of Trees & Shrubs, p.206-215. Liu, Y.-H., Zeng, Q.-W., Zhou, R.-Z. & Xing, F.-W. - (2004) - Magnolias of China, 391p. Krüssmann, G. - (1977) - Magnolia in Handbuch der Laubgehölze, VOL.3, p.275-288. Meyer, F.G. - (1977) - Magnoliaceae in Flora of North America, VOL.3: online edition Rehder, A. - (1940) - Magnoliaceae in Manual of cultivated trees and shrubs hardy in North America, p.246-253. -

A Note on Magnolia, Mainly of Sections Manglietia and Michelia Subgenus
A note on Magnolia, mainly of sections Manglietia and Michelia subgenus Magnolia (Magnoliaceae) A note of caution concerning the ultimate heights that may be achieved in cultivation by numerous of the newer evergreen magnolias from Asia, is the theme of this article by CHRIS CALLAGHAN and S. K. PNG of the Australian Bicentennial Arboretum (ABA). Following Thomas Methuen-Campbell’s interesting report in the 2011 IDS Yearbook concerning the study weekend held in June of that year at RHS Wisley, Surrey, to discuss “summer” flowering magnolias (see Endnote), the authors thought they should write to mention an important consideration before contemplating planting of these trees in gardens, or indeed any tree in a garden, particularly the average small garden. We are not sure if the ultimate size of many of these magnolias was discussed with those attending the study weekend, since most of their maximum known heights were not mentioned in the article. However, we believe any readers tempted by the article to purchase and plant out most of the evergreen magnolias featured (previously in the genera Manglietia, Michelia or Parakmeria) in a normal suburban front or backyard in relatively 46 warm, sheltered, near frost-free areas, will be ultimately dismayed by the sizes they reach (see Figlar 2009 for reasons behind the reduction of genera). Even allowing that these predominantly warm-temperate to sub-tropical forest trees may not achieve their maximum potential sizes in the milder regions of temperate Europe, most are still likely to overtop (and overshadow!) two or three storey homes or apartments, especially with a warming climate. -

Approved Plants For
Perennials, Ground Covers, Annuals & Bulbs Scientific name Common name Achillea millefolium Common Yarrow Alchemilla mollis Lady's Mantle Aster novae-angliae New England Aster Astilbe spp. Astilbe Carex glauca Blue Sedge Carex grayi Morningstar Sedge Carex stricta Tussock Sedge Ceratostigma plumbaginoides Leadwort/Plumbago Chelone glabra White Turtlehead Chrysanthemum spp. Chrysanthemum Convallaria majalis Lily-of-the-Valley Coreopsis lanceolata Lanceleaf Tickseed Coreopsis rosea Rosy Coreopsis Coreopsis tinctoria Golden Tickseed Coreopsis verticillata Threadleaf Coreopsis Dryopteris erythrosora Autumn Fern Dryopteris marginalis Leatherleaf Wood Fern Echinacea purpurea 'Magnus' Magnus Coneflower Epigaea repens Trailing Arbutus Eupatorium coelestinum Hardy Ageratum Eupatorium hyssopifolium Hyssopleaf Thoroughwort Eupatorium maculatum Joe-Pye Weed Eupatorium perfoliatum Boneset Eupatorium purpureum Sweet Joe-Pye Weed Geranium maculatum Wild Geranium Hedera helix English Ivy Hemerocallis spp. Daylily Hibiscus moscheutos Rose Mallow Hosta spp. Plantain Lily Hydrangea quercifolia Oakleaf Hydrangea Iris sibirica Siberian Iris Iris versicolor Blue Flag Iris Lantana camara Yellow Sage Liatris spicata Gay-feather Liriope muscari Blue Lily-turf Liriope variegata Variegated Liriope Lobelia cardinalis Cardinal Flower Lobelia siphilitica Blue Cardinal Flower Lonicera sempervirens Coral Honeysuckle Narcissus spp. Daffodil Nepeta x faassenii Catmint Onoclea sensibilis Sensitive Fern Osmunda cinnamomea Cinnamon Fern Pelargonium x domesticum Martha Washington -

Phylogenomic Approach
Toward the ultimate phylogeny of Magnoliaceae: phylogenomic approach Sangtae Kim*1, Suhyeon Park1, and Jongsun Park2 1 Sungshin University, Korea 2 InfoBoss Co., Korea Mr. Carl Ferris Miller Founder of Chollipo Arboretum in Korea Chollipo Arboretum Famous for its magnolia collection 2020. Annual Meeting of Magnolia Society International Cholliop Arboretum in Korea. April 13th~22th, 2020 http://WWW.Chollipo.org Sungshin University, Seoul, Korea Dr. Hans Nooteboom Dr. Liu Yu-Hu Twenty-one years ago... in 1998 The 1st International Symposium on the Family Magnoliaceae, Gwangzhow Dr. Hiroshi Azuma Mr. Richard Figlar Dr. Hans Nooteboom Dr. Qing-wen Zeng Dr. Weibang Sun Handsome young boy Dr. Yong-kang Sima Dr. Yu-wu Law Presented ITS study on Magnoliaceae - never published Ten years ago... in 2009 Presented nine cp genome region study (9.2 kbp) on Magnoliaceae – published in 2013 2015 1st International Sympodium on Neotropical Magnoliaceae Gadalajara, 2019 3rd International Sympodium and Workshop on Neotropical Magnoliaceae Asterales Dipsacales Apiales Why magnolia study is Aquifoliales Campanulids (Euasterids II) Garryales Gentianales Laminales Solanales Lamiids important in botany? Ericales Asterids (Euasterids I) Cornales Sapindales Malvales Brassicales Malvids Fagales (Eurosids II) • As a member of early-diverging Cucurbitales Rosales Fabales Zygophyllales Celestrales Fabids (Eurosid I) angiosperms, reconstruction of the Oxalidales Malpighiales Vitales Geraniales Myrtales Rosids phylogeny of Magnoliaceae will Saxifragales Caryphyllales -
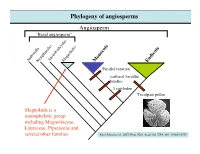
Phylogeny of Angiosperms Angiosperm “Basal Angiosperm”
Phylogeny of angiosperms Angiosperm “Basal angiosperm” AmborellaNymphaealesAustrobaileyalesMagnoliidss Monocots Eudicots Parallel venation scattered vascular bundles 1 cotyledon Tricolpate pollen Magnoliids is a monophyletic group including Magnoliaceae, Lauraceae, Piperaceae and several other families After Jansen et al., 2007, Proc. Natl. Acad. Sci. USA 104: 19369-19374 Magnoliaceae (Magnolia family) Textbook DVD KRR Magnolia X soulangeana; Magnoliaceae (Magnolia family) Textbook DVD WSJ Textbook DVD KRR Magnolia grandiflora; Magnolia macrophylla; Note leaf simple, entire, pinnate venation, numerous tepals, numerous stamens and carples. Textbook DVD KRR Magnolia sieboldii; Magnoliaceae (Magnolia family) Textbook DVD KMN Textbook DVD SMK-KRR Magnolia figo; Magnolia grandiflora; Note the elongated receptacle, Note the aggregate of follicles, and laminar stamens and red fleshy seed coat Magnoliaceae (Magnolia family) Photo: Yaowu Yuan Photo: Yaowu Yuan Liriodendron tulipifera; Note the elongated receptacle, and laminar stamens Magnoliaceae (Magnolia family) Note the lobed, T-shirt-like leaf, and pinnate venation Note the aggregate of samara Magnoliaceae (Magnolia family) Magnoliaceae - 2 genera/220 species. Trees or shrubs; Ethereal oils (aromatic terpenoids) - (remember the smell of bay leaves?); Leaves alternate, simple (Magnolia) or lobed (Liriodendron), entire; Flowers large and showy, actinomorphic, bisexual Tepals 6-numerous, stamens and carpels numerous, Spirally arranged on an elongated receptacle, Laminar stamens poorly differentiated into anther and filament. Fruit usually an aggregate of follicle (Magnolia) or samara (Liriodendron); follicle: 1-carpellate fruit that dehisces on the side samara: 1-carpellate winged, indehiscent fruit Phylogeny of Eudicots (or Tricolpates) Eudicots (or Tricolpates) “Basal eudicots” Asterids Buxales Rosids Caryophyllales RanunculalesProteales Ranunculales is a monophyletic group including Ranunculaceae, Berberidaceae, Papaveraceae, and 4 other families. After Jansen et al., 2007, Proc.