Expansion During Axolotl Spinal Cord Regeneration
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

ATRX Induction by Mutant Huntingtin Via Cdx2 Modulates Heterochromatin Condensation and Pathology in Huntington’S Disease
Cell Death and Differentiation (2012) 19, 1109–1116 & 2012 Macmillan Publishers Limited All rights reserved 1350-9047/12 www.nature.com/cdd ATRX induction by mutant huntingtin via Cdx2 modulates heterochromatin condensation and pathology in Huntington’s disease J Lee1,2, YK Hong3, GS Jeon4, YJ Hwang4, KY Kim4, KH Seong4, M-K Jung4, DJ Picketts5, NW Kowall1,2, KS Cho3 and H Ryu*,1,2,4 Aberrant chromatin remodeling is involved in the pathogenesis of Huntington’s disease (HD) but the mechanism is not known. Herein, we report that mutant huntingtin (mtHtt) induces the transcription of alpha thalassemia/mental retardation X linked (ATRX), an ATPase/helicase and SWI/SNF-like chromatin remodeling protein via Cdx-2 activation. ATRX expression was elevated in both a cell line model and transgenic model of HD, and Cdx-2 occupancy of the ATRX promoter was increased in HD. Induction of ATRX expanded the size of promyelocytic leukemia nuclear body (PML-NB) and increased trimethylation of H3K9 (H3K9me3) and condensation of pericentromeric heterochromatin, while knockdown of ATRX decreased PML-NB and H3K9me3 levels. Knockdown of ATRX/dXNP improved the hatch rate of fly embryos expressing mtHtt (Q127). ATRX/dXNP overexpression exacerbated eye degeneration of eye-specific mtHtt (Q127) expressing flies. Our findings suggest that transcriptional alteration of ATRX by mtHtt is involved in pericentromeric heterochromatin condensation and contributes to the pathogenesis of HD. Cell Death and Differentiation (2012) 19, 1109–1116; doi:10.1038/cdd.2011.196; published -

ASCL2 Expression Contributes to Gastric Tumor Migration and Invasion by Downregulating Mir223 and Inducing EMT
MOLECULAR MEDICINE REPORTS 18: 3751-3759, 2018 ASCL2 expression contributes to gastric tumor migration and invasion by downregulating miR223 and inducing EMT QINGSONG ZUO*, JIE WANG*, CHAO CHEN, YONG ZHANG, DIAN-XU FENG, RONGHUA ZHAO and TENG CHEN Department of General Surgery, Putuo Hospital, Shanghai University of Traditional Chinese Medicine, Shanghai 200062, P.R. China Received May 15, 2017; Accepted November 23, 2017 DOI: 10.3892/mmr.2018.9363 Abstract. Achaete-scute homolog 2 (ASCL2), a basic results of the present study demonstrated that ASCL2 was able helix-loop-helix transcription factor, serves an essential role in to downregulate the expression level of miR223, contribute to the maintenance of adult intestinal stem cells and the growth EMT and promote gastric tumor metastasis, which indicated of gastric cancer (GC). However, the function of ASCL2 in that ASCL2 may serve as a therapeutic target in the treatment the metastasis of GC is poorly understood. The present study of GC. aimed to evaluate the effect of ASCL2 expression on gastric tumor metastasis. ASCL2 protein expression was detected in Introduction 32 cases of gastric metastasis and its relevant primary tumors using western blotting and immunohistochemistry. The Gastric cancer (GC) is the second most common cause of data suggested that the expression of ASCL2 was highest in cancer-associated mortality in the world, and the incidence metastatic tumors, among adjacent normal tissues, primary GC is highest in East Asia, Eastern Europe and parts of Latin gastric tumors and gastric metastatic tumors. Furthermore, America (1,2). However, the precise mechanisms underlying ASCL2-overexpressing GC cell lines MKN1-ASCL2 and gastric carcinogenesis are not yet completely understood (3). -

Isolation and Characterization of BEN, a Member of the TFII-I Family of DNA-Binding Proteins Containing Distinct Helix–Loop–Helix Domains
Isolation and characterization of BEN, a member of the TFII-I family of DNA-binding proteins containing distinct helix–loop–helix domains Dashzeveg Bayarsaihan and Frank H. Ruddle* Department of Molecular, Cellular, and Developmental Biology, Kline Biology Tower, Yale University, 266 Whitney Avenue, New Haven, CT 06520 Contributed by Frank H. Ruddle, May 4, 2000 The transcriptional regulation of the Hoxc8 gene is controlled during paraxial mesoderm at the 14th somite, and in the lateral plate early mouse embryogenesis by an enhanceosome-like control region, mesoderm at the 12th somite. Later in development, posterior termed the early enhancer (EE), located 3 kb upstream from the Hoxc8 expression of Hoxc8 decreases, whereas intense expression is translation start site. The EE is involved in establishing the posterior maintained at previously determined anterior limits within the expression pattern of Hoxc8 at embryonic day (E) 8.5–9.0. Genetic and thorax (somites and lateral plate mesoderm) and in the brachial biochemical data have shown that nuclear factors interact with this region (neural tube) (7). region in a sequence-specific manner. We have used a yeast one- Transgenic reporter analysis was used to identify cis-regulatory hybrid screen in a search for transcription factors that bind to EE domains critical for the normal expression of Hoxc8. Two distinct motifs and have isolated a novel murine DNA-binding protein, termed genomic regions were identified that regulate the early and late BEN (binding factor for early enhancer). The ORF of BEN encodes a phases of Hoxc8 expression (5, 6). The early phase is regulated by protein of 1072 amino acids and contains six helix–loop–helix do- DNA elements located 3 kb upstream from the translation start site mains, a hydrophobic leucine zipper-like motif, and a serine-rich of the gene, whereas the late expression is regulated by elements repeat. -

The Human Gene Map for Performance and Health-Related Fitness Phenotypes: the 2006–2007 Update
BASIC SCIENCES The Human Gene Map for Performance and Health-Related Fitness Phenotypes: The 2006–2007 Update MOLLY S. BRAY1, JAMES M. HAGBERG2, LOUIS PE´ RUSSE3, TUOMO RANKINEN4, STEPHEN M. ROTH2, BERND WOLFARTH5, and CLAUDE BOUCHARD4 1USDA/ARS Children’s Nutrition Research Center, Baylor College of Medicine, Houston, TX; 2Department of Kinesiology, School of Public Health, University of Maryland, College Park, MD; 3Division of Kinesiology, Department of Preventive Medicine, Laval University, Ste-Foy, Que´bec, CANADA; 4Human Genomics Laboratory, Pennington Biomedical Research Center, Baton Rouge, LA; and 5Preventive and Rehabilitative Sports Medicine, Technical University Munich, Munich, GERMANY ABSTRACT BRAY, M. S., J. M. HAGBERG, L. PE´ RUSSE, T. RANKINEN, S. M. ROTH, B. WOLFARTH, and C. BOUCHARD. The Human Gene Map for Performance and Health-Related Fitness Phenotypes: The 2006–2007 Update. Med. Sci. Sports Exerc., Vol. 41, No. 1, pp. 34–72, 2009. This update of the human gene map for physical performance and health-related fitness phenotypes covers the research advances reported in 2006 and 2007. The genes and markers with evidence of association or linkage with a performance or a fitness phenotype in sedentary or active people, in responses to acute exercise, or for training-induced adaptations are positioned on the map of all autosomes and sex chromosomes. Negative studies are reviewed, but a gene or a locus must be supported by at least one positive study before being inserted on the map. A brief discussion on the nature of the evidence and on what to look for in assessing human genetic studies of relevance to fitness and performance is offered in the introduction, followed by a review of all studies published in 2006 and 2007. -

Genome-Wide DNA Methylation Analysis Reveals Molecular Subtypes of Pancreatic Cancer
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 17), pp: 28990-29012 Research Paper Genome-wide DNA methylation analysis reveals molecular subtypes of pancreatic cancer Nitish Kumar Mishra1 and Chittibabu Guda1,2,3,4 1Department of Genetics, Cell Biology and Anatomy, University of Nebraska Medical Center, Omaha, NE, 68198, USA 2Bioinformatics and Systems Biology Core, University of Nebraska Medical Center, Omaha, NE, 68198, USA 3Department of Biochemistry and Molecular Biology, University of Nebraska Medical Center, Omaha, NE, 68198, USA 4Fred and Pamela Buffet Cancer Center, University of Nebraska Medical Center, Omaha, NE, 68198, USA Correspondence to: Chittibabu Guda, email: [email protected] Keywords: TCGA, pancreatic cancer, differential methylation, integrative analysis, molecular subtypes Received: October 20, 2016 Accepted: February 12, 2017 Published: March 07, 2017 Copyright: Mishra et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Pancreatic cancer (PC) is the fourth leading cause of cancer deaths in the United States with a five-year patient survival rate of only 6%. Early detection and treatment of this disease is hampered due to lack of reliable diagnostic and prognostic markers. Recent studies have shown that dynamic changes in the global DNA methylation and gene expression patterns play key roles in the PC development; hence, provide valuable insights for better understanding the initiation and progression of PC. In the current study, we used DNA methylation, gene expression, copy number, mutational and clinical data from pancreatic patients. -

Adjusting the Molecular Clock: the Importance of Circadian Rhythms in the Development of Glioblastomas and Its Intervention As a Therapeutic Strategy
International Journal of Molecular Sciences Review Adjusting the Molecular Clock: The Importance of Circadian Rhythms in the Development of Glioblastomas and Its Intervention as a Therapeutic Strategy Paula M. Wagner 1,2,†,César G. Prucca 1,2,† , Beatriz L. Caputto 1,2 and Mario E. Guido 1,2,* 1 CIQUIBIC-CONICET, Facultad de Ciencias Químicas, Universidad Nacional de Córdoba, Córdoba 5000, Argentina; [email protected] (P.M.W.); [email protected] (C.G.P.); [email protected] (B.L.C.) 2 Departamento de Química Biológica Ranwel Caputto, Facultad de Ciencias Químicas, Universidad Nacional de Córdoba, Córdoba 5000, Argentina * Correspondence: [email protected] † Equal contribution. Abstract: Gliomas are solid tumors of the central nervous system (CNS) that originated from different glial cells. The World Health Organization (WHO) classifies these tumors into four groups (I-IV) with increasing malignancy. Glioblastoma (GBM) is the most common and aggressive type of brain tumor classified as grade IV. GBMs are resistant to conventional therapies with poor prognosis after diagnosis even when the Stupp protocol that combines surgery and radiochemotherapy is applied. Nowadays, few novel therapeutic strategies have been used to improve GBM treatment, looking for higher efficiency and lower side effects, but with relatively modest results. The circadian timing system temporally organizes the physiology and behavior of most organisms and daily regulates several cellular processes in organs, tissues, and even in individual cells, including tumor cells. The Citation: Wagner, P.M.; Prucca, C.G.; Caputto, B.L.; Guido, M.E. Adjusting potentiality of the function of the circadian clock on cancer cells modulation as a new target for novel the Molecular Clock: The Importance treatments with a chronobiological basis offers a different challenge that needs to be considered in of Circadian Rhythms in the further detail. -
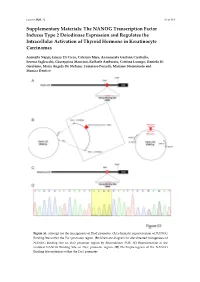
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

VDR) Gene in Preeclampsia Patients in the Chinese Han Population
Hypertension Research (2015) 38, 513–517 & 2015 The Japanese Society of Hypertension All rights reserved 0916-9636/15 www.nature.com/hr ORIGINAL ARTICLE Genetic variations in the vitamin-D receptor (VDR) gene in preeclampsia patients in the Chinese Han population Ying Zhan1,5, Mengchun Liu2,5, Yuelan You3, Yan Zhang1, Jingli Wang4, Xunfeng Wang4, Shiguo Liu1,4 and Xuemei Liu2 Previous studies have indicated that vitamin D deficiency is linked to a risk of preeclampsia (PE). The aim of our study was to investigate the association between genetic variations in the vitamin-D receptor (VDR) gene and the susceptibility to PE in the Chinese Han population. We examined the genotypes VDR rs2228570, rs11568820 and rs1544410 in 402 PE patients and 554 normal pregnant women in the third trimester by TaqMan allelic discrimination real-time polymerase chain reaction. The clinical data of the individuals were collected to enable genotype–phenotype analysis. A significant statistical difference in the genotypic frequencies of rs2228570 between cases and controls was found (χ2 = 13.750, P = 0.001). The G allele was the risk factor for the risk of PE (χ2 = 9.456, P = 0.002, OR = 1.137, 95% CI 1.111–1.610). There was no difference in the genotypic and allelic distributions of rs11568820 and rs1544410 between the two groups (P> 0.05). Our results provide evidence for a possible link between VDR and the development of PE in the Chinese Han population. Hypertension Research (2015) 38, 513–517; doi:10.1038/hr.2015.29; published online 19 March 2015 Keywords: Chinese Han population; polymorphism; preeclampsia; vitamin-D receptor 12 INTRODUCTION ligand-binding domain. -

A Crucial Role for the Ubiquitously Expressed Transcription Factor Sp1 at Early Stages of Hematopoietic Specification
This is a repository copy of A crucial role for the ubiquitously expressed transcription factor Sp1 at early stages of hematopoietic specification.. White Rose Research Online URL for this paper: http://eprints.whiterose.ac.uk/83683/ Version: Published Version Article: Gilmour, J, Assi, SA, Jaegle, U et al. (6 more authors) (2014) A crucial role for the ubiquitously expressed transcription factor Sp1 at early stages of hematopoietic specification. Development, 141 (12). 2391 - 2401. ISSN 0950-1991 https://doi.org/10.1242/dev.106054 Reuse Unless indicated otherwise, fulltext items are protected by copyright with all rights reserved. The copyright exception in section 29 of the Copyright, Designs and Patents Act 1988 allows the making of a single copy solely for the purpose of non-commercial research or private study within the limits of fair dealing. The publisher or other rights-holder may allow further reproduction and re-use of this version - refer to the White Rose Research Online record for this item. Where records identify the publisher as the copyright holder, users can verify any specific terms of use on the publisher’s website. Takedown If you consider content in White Rose Research Online to be in breach of UK law, please notify us by emailing [email protected] including the URL of the record and the reason for the withdrawal request. [email protected] https://eprints.whiterose.ac.uk/ © 2014. Published by The Company of Biologists Ltd | Development (2014) 141, 2391-2401 doi:10.1242/dev.106054 RESEARCH ARTICLE STEM CELLS AND REGENERATION A crucial role for the ubiquitously expressed transcription factor Sp1 at early stages of hematopoietic specification Jane Gilmour1, Salam A. -

The Vitamin D Receptor: Contemporary Genomic Approaches Reveal New Basic and Translational Insights
REVIEW SERIES: NUCLEAR RECEPTORS The Journal of Clinical Investigation Series Editor: Mitchell A. Lazar The vitamin D receptor: contemporary genomic approaches reveal new basic and translational insights J. Wesley Pike, Mark B. Meyer, Seong-Min Lee, Melda Onal, and Nancy A. Benkusky Department of Biochemistry, University of Wisconsin — Madison, Madison, Wisconsin, USA. The vitamin D receptor (VDR) is the single known regulatory mediator of hormonal 1,25-dihydroxyvitamin D3 [1,25(OH)2D3] in higher vertebrates. It acts in the nucleus of vitamin D target cells to regulate the expression of genes whose products control diverse, cell type–specific biological functions that include mineral homeostasis. In this Review we describe progress that has been made in defining new cellular sites of action of this receptor, the mechanisms through which this mediator controls the expression of genes, the biology that ensues, and the translational impact of this receptor on human health and disease. We conclude with a brief discussion of what comes next in understanding vitamin D biology and the mechanisms that underlie its actions. Introduction involved (14, 15). With this background, we then comment briefly on In the early 1970s, 1,25-dihydroxyvitamin D3 [1,25(OH)2D3] was the current translational impact of several features of VDR action identified as both the exclusive, metabolically active form of vita- and function on human health and disease. min D and a key component of what proved to be an exquisite endo- crine system that regulates numerous biologic processes in higher The VDR vertebrates (1, 2). This pioneering discovery ended a decades-long VDR tissue distribution. -

WO 2012/054896 Al
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number ι (43) International Publication Date ¾ ί t 2 6 April 2012 (26.04.2012) WO 2012/054896 Al (51) International Patent Classification: AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, C12N 5/00 (2006.01) C12N 15/00 (2006.01) CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, C12N 5/02 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, (21) International Application Number: KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, PCT/US201 1/057387 ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, (22) International Filing Date: NO, NZ, OM, PE, PG, PH, PL, PT, QA, RO, RS, RU, 2 1 October 201 1 (21 .10.201 1) RW, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, (25) Filing Language: English ZM, ZW. (26) Publication Language: English (84) Designated States (unless otherwise indicated, for every (30) Priority Data: kind of regional protection available): ARIPO (BW, GH, 61/406,064 22 October 2010 (22.10.2010) US GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, SZ, TZ, 61/415,244 18 November 2010 (18.1 1.2010) US UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, MD, RU, TJ, TM), European (AL, AT, BE, BG, CH, CY, CZ, (71) Applicant (for all designated States except US): BIO- DE, DK, EE, ES, FI, FR, GB, GR, HR, HU, IE, IS, IT, TIME INC. -

DNA Methylation in Placentas of Interspecies Mouse Hybrids
Copyright 2003 by the Genetics Society of America DNA Methylation in Placentas of Interspecies Mouse Hybrids Sabine Schu¨tt,*,1 Andrea R. Florl,† Wei Shi,*,‡ Myriam Hemberger,*,2 Annie Orth,§ Sabine Otto,* Wolfgang A. Schulz† and Reinald H. Fundele*,‡,3 *Max-Planck-Institute for Molecular Genetics, 14195 Berlin, Germany, †Department of Oncology, Heinrich-Heine-University, 40225 Du¨sseldorf, Germany, ‡Department of Development and Genetics, University of Uppsala, Norbyva¨gen 18A, S-75236, Sweden and §Laboratory of Genomes and Populations, University of Montpellier II, 34095 Montpellier Cedex 5, France Manuscript received October 14, 2002 Accepted for publication April 2, 2003 ABSTRACT Interspecific hybridization in the genus Mus results in several hybrid dysgenesis effects, such as male sterility and X-linked placental dysplasia (IHPD). The genetic or molecular basis for the placental pheno- types is at present not clear. However, an extremely complex genetic system that has been hypothesized to be caused by major epigenetic changes on the X chromosome has been shown to be active. We have investigated DNA methylation of several single genes, Atrx, Esx1, Mecp2, Pem, Psx1, Vbp1, Pou3f4, and Cdx2, and, in addition, of LINE-1 and IAP repeat sequences, in placentas and tissues of fetal day 18 mouse interspecific hybrids. Our results show some tendency toward hypomethylation in the late gestation mouse placenta. However, no differential methylation was observed in hyper- and hypoplastic hybrid placentas when compared with normal-sized littermate placentas or intraspecific Mus musculus placentas of the same developmental stage. Thus, our results strongly suggest that generalized changes in methylation patterns do not occur in trophoblast cells of such hybrids.