Supplementary Figure 1. HEK293 Were Transiently Transfected with The
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic Variation Across the Human Olfactory Receptor Repertoire Alters Odor Perception
bioRxiv preprint doi: https://doi.org/10.1101/212431; this version posted November 1, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Genetic variation across the human olfactory receptor repertoire alters odor perception Casey Trimmer1,*, Andreas Keller2, Nicolle R. Murphy1, Lindsey L. Snyder1, Jason R. Willer3, Maira Nagai4,5, Nicholas Katsanis3, Leslie B. Vosshall2,6,7, Hiroaki Matsunami4,8, and Joel D. Mainland1,9 1Monell Chemical Senses Center, Philadelphia, Pennsylvania, USA 2Laboratory of Neurogenetics and Behavior, The Rockefeller University, New York, New York, USA 3Center for Human Disease Modeling, Duke University Medical Center, Durham, North Carolina, USA 4Department of Molecular Genetics and Microbiology, Duke University Medical Center, Durham, North Carolina, USA 5Department of Biochemistry, University of Sao Paulo, Sao Paulo, Brazil 6Howard Hughes Medical Institute, New York, New York, USA 7Kavli Neural Systems Institute, New York, New York, USA 8Department of Neurobiology and Duke Institute for Brain Sciences, Duke University Medical Center, Durham, North Carolina, USA 9Department of Neuroscience, University of Pennsylvania School of Medicine, Philadelphia, Pennsylvania, USA *[email protected] ABSTRACT The human olfactory receptor repertoire is characterized by an abundance of genetic variation that affects receptor response, but the perceptual effects of this variation are unclear. To address this issue, we sequenced the OR repertoire in 332 individuals and examined the relationship between genetic variation and 276 olfactory phenotypes, including the perceived intensity and pleasantness of 68 odorants at two concentrations, detection thresholds of three odorants, and general olfactory acuity. -

Environmental Influences on Endothelial Gene Expression
ENDOTHELIAL CELL GENE EXPRESSION John Matthew Jeff Herbert Supervisors: Prof. Roy Bicknell and Dr. Victoria Heath PhD thesis University of Birmingham August 2012 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. ABSTRACT Tumour angiogenesis is a vital process in the pathology of tumour development and metastasis. Targeting markers of tumour endothelium provide a means of targeted destruction of a tumours oxygen and nutrient supply via destruction of tumour vasculature, which in turn ultimately leads to beneficial consequences to patients. Although current anti -angiogenic and vascular targeting strategies help patients, more potently in combination with chemo therapy, there is still a need for more tumour endothelial marker discoveries as current treatments have cardiovascular and other side effects. For the first time, the analyses of in-vivo biotinylation of an embryonic system is performed to obtain putative vascular targets. Also for the first time, deep sequencing is applied to freshly isolated tumour and normal endothelial cells from lung, colon and bladder tissues for the identification of pan-vascular-targets. Integration of the proteomic, deep sequencing, public cDNA libraries and microarrays, delivers 5,892 putative vascular targets to the science community. -

Copy Number Variation in Fetal Alcohol Spectrum Disorder
Biochemistry and Cell Biology Copy number variation in fetal alcohol spectrum disorder Journal: Biochemistry and Cell Biology Manuscript ID bcb-2017-0241.R1 Manuscript Type: Article Date Submitted by the Author: 09-Nov-2017 Complete List of Authors: Zarrei, Mehdi; The Centre for Applied Genomics Hicks, Geoffrey G.; University of Manitoba College of Medicine, Regenerative Medicine Reynolds, James N.; Queen's University School of Medicine, Biomedical and Molecular SciencesDraft Thiruvahindrapuram, Bhooma; The Centre for Applied Genomics Engchuan, Worrawat; Hospital for Sick Children SickKids Learning Institute Pind, Molly; University of Manitoba College of Medicine, Regenerative Medicine Lamoureux, Sylvia; The Centre for Applied Genomics Wei, John; The Centre for Applied Genomics Wang, Zhouzhi; The Centre for Applied Genomics Marshall, Christian R.; The Centre for Applied Genomics Wintle, Richard; The Centre for Applied Genomics Chudley, Albert; University of Manitoba Scherer, Stephen W.; The Centre for Applied Genomics Is the invited manuscript for consideration in a Special Fetal Alcohol Spectrum Disorder Issue? : Keyword: Fetal alcohol spectrum disorder, FASD, copy number variations, CNV https://mc06.manuscriptcentral.com/bcb-pubs Page 1 of 354 Biochemistry and Cell Biology 1 Copy number variation in fetal alcohol spectrum disorder 2 Mehdi Zarrei,a Geoffrey G. Hicks,b James N. Reynolds,c,d Bhooma Thiruvahindrapuram,a 3 Worrawat Engchuan,a Molly Pind,b Sylvia Lamoureux,a John Wei,a Zhouzhi Wang,a Christian R. 4 Marshall,a Richard F. Wintle,a Albert E. Chudleye,f and Stephen W. Scherer,a,g 5 aThe Centre for Applied Genomics and Program in Genetics and Genome Biology, The Hospital 6 for Sick Children, Toronto, Ontario, Canada 7 bRegenerative Medicine Program, University of Manitoba, Winnipeg, Canada 8 cCentre for Neuroscience Studies, Queen's University, Kingston, Ontario, Canada. -

LETTER Doi:10.1038/Nature09515
LETTER doi:10.1038/nature09515 Distant metastasis occurs late during the genetic evolution of pancreatic cancer Shinichi Yachida1*, Siaˆn Jones2*, Ivana Bozic3, Tibor Antal3,4, Rebecca Leary2, Baojin Fu1, Mihoko Kamiyama1, Ralph H. Hruban1,5, James R. Eshleman1, Martin A. Nowak3, Victor E. Velculescu2, Kenneth W. Kinzler2, Bert Vogelstein2 & Christine A. Iacobuzio-Donahue1,5,6 Metastasis, the dissemination and growth of neoplastic cells in an were present in the primary pancreatic tumours from which the meta- organ distinct from that in which they originated1,2, is the most stases arose. A small number of these samples of interest were cell lines common cause of death in cancer patients. This is particularly true or xenografts, similar to the index lesions, whereas the majority were for pancreatic cancers, where most patients are diagnosed with fresh-frozen tissues that contained admixed neoplastic, stromal, metastatic disease and few show a sustained response to chemo- inflammatory, endothelial and normal epithelial cells (Fig. 1a). Each therapy or radiation therapy3. Whether the dismal prognosis of tissue sample was therefore microdissected to minimize contaminat- patients with pancreatic cancer compared to patients with other ing non-neoplastic elements before purifying DNA. types of cancer is a result of late diagnosis or early dissemination of Two categories of mutations were identified (Fig. 1b). The first and disease to distant organs is not known. Here we rely on data gen- largest category corresponded to those mutations present in all samples erated by sequencing the genomes of seven pancreatic cancer meta- from a given patient (‘founder’ mutations, mean of 64%, range 48–83% stases to evaluate the clonal relationships among primary and of all mutations per patient; Fig. -

Crosstalk in Competing Endogenous RNA Network Reveals the Complex Molecular Mechanism Underlying Lung Cancer
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 53), pp: 91270-91280 Research Paper Crosstalk in competing endogenous RNA network reveals the complex molecular mechanism underlying lung cancer Xiang Jin1, Yinghui Guan1, Hui Sheng1 and Yang Liu1 1Department of Respiration, The First Hospital of Jilin University, Changchun City, Jilin Province, 130021, China Correspondence to: Yinghui Guan, email: [email protected] Keywords: crosstalk, ceRNA network, lung cancer, molecular mechanism, circRNA-seq Received: March 29, 2017 Accepted: July 16, 2017 Published: August 24, 2017 Copyright: Jin et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT We investigated the transcriptional mechanism underlying lung cancer development. RNA sequencing analysis was performed on blood samples from lung cancer cases and healthy controls. Differentially expressed microRNAs (miRNAs), circular RNAs (circRNAs), mRNAs (genes), and long non-coding RNAs (lncRNA) were identified, followed by pathway enrichment analysis. Based on miRNA target interactions, a competing endogenous network was established and significant nodes were screened. Differentially expressed transcriptional factors were retrieved from the TRRUST database and the transcriptional factor regulatory network was constructed. The expression of 59 miRNAs, 18,306 genes,232 lncRNAs, and 292 circRNAs were greatly altered in patients with lung cancer. miRNAs were closely associated with cancer-related pathways, such as pathways in cancer, colorectal cancer, and transcriptional misregulation in cancer. Two novel pathways, olfactory transduction and neuroactive ligand-receptor interactions, were significantly enriched by differentially expressed genes. -

OR2A5 (NM 012365) Human Tagged ORF Clone – RC220078L3
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC220078L3 OR2A5 (NM_012365) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: OR2A5 (NM_012365) Human Tagged ORF Clone Tag: Myc-DDK Symbol: OR2A5 Synonyms: OR2A8; OR2A11P; OR2A26; OR7-138; OR7-141 Vector: pLenti-C-Myc-DDK-P2A-Puro (PS100092) E. coli Selection: Chloramphenicol (34 ug/mL) Cell Selection: Puromycin ORF Nucleotide The ORF insert of this clone is exactly the same as(RC220078). Sequence: Restriction Sites: SgfI-MluI Cloning Scheme: ACCN: NM_012365 ORF Size: 933 bp This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 OR2A5 (NM_012365) Human Tagged ORF Clone – RC220078L3 OTI Disclaimer: The molecular sequence of this clone aligns with the gene accession number as a point of reference only. However, individual transcript sequences of the same gene can differ through naturally occurring variations (e.g. polymorphisms), each with its own valid existence. This clone is substantially in agreement with the reference, but a complete review of all prevailing variants is recommended prior to use. More info OTI Annotation: This clone was engineered to express the complete ORF with an expression tag. Expression varies depending on the nature of the gene. RefSeq: NM_012365.1, NP_036497.1 RefSeq Size: 936 bp RefSeq ORF: 936 bp Locus ID: 393046 UniProt ID: Q96R48, A0A126GW49 Protein Pathways: Olfactory transduction MW: 35 kDa Gene Summary: Olfactory receptors interact with odorant molecules in the nose, to initiate a neuronal response that triggers the perception of a smell. -

WO 2019/068007 Al Figure 2
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/068007 Al 04 April 2019 (04.04.2019) W 1P O PCT (51) International Patent Classification: (72) Inventors; and C12N 15/10 (2006.01) C07K 16/28 (2006.01) (71) Applicants: GROSS, Gideon [EVIL]; IE-1-5 Address C12N 5/10 (2006.0 1) C12Q 1/6809 (20 18.0 1) M.P. Korazim, 1292200 Moshav Almagor (IL). GIBSON, C07K 14/705 (2006.01) A61P 35/00 (2006.01) Will [US/US]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., C07K 14/725 (2006.01) P.O. Box 4044, 7403635 Ness Ziona (TL). DAHARY, Dvir [EilL]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., P.O. (21) International Application Number: Box 4044, 7403635 Ness Ziona (IL). BEIMAN, Merav PCT/US2018/053583 [EilL]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., P.O. (22) International Filing Date: Box 4044, 7403635 Ness Ziona (E.). 28 September 2018 (28.09.2018) (74) Agent: MACDOUGALL, Christina, A. et al; Morgan, (25) Filing Language: English Lewis & Bockius LLP, One Market, Spear Tower, SanFran- cisco, CA 94105 (US). (26) Publication Language: English (81) Designated States (unless otherwise indicated, for every (30) Priority Data: kind of national protection available): AE, AG, AL, AM, 62/564,454 28 September 2017 (28.09.2017) US AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, 62/649,429 28 March 2018 (28.03.2018) US CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, (71) Applicant: IMMP ACT-BIO LTD. -

Olfactory Receptors in Non-Chemosensory Tissues
BMB Reports Invited Mini Review Olfactory receptors in non-chemosensory tissues NaNa Kang & JaeHyung Koo* Department of Brain Science, Daegu Gyeongbuk Institute of Science and Technology (DGIST), Daegu 711-873, Korea Olfactory receptors (ORs) detect volatile chemicals that lead to freezing behavior (3-5). the initial perception of smell in the brain. The olfactory re- ORs are localized in the cilia of olfactory sensory neurons ceptor (OR) is the first protein that recognizes odorants in the (OSNs) in the olfactory epithelium (OE) and are activated by olfactory signal pathway and it is present in over 1,000 genes chemical cues, typically odorants at the molecular level, in mice. It is also the largest member of the G protein-coupled which lead to the perception of smell in the brain (6). receptors (GPCRs). Most ORs are extensively expressed in the Tremendous research was conducted since Buck and Axel iso- nasal olfactory epithelium where they perform the appropriate lated ORs as an OE-specific expression in 1991 (7). OR genes, physiological functions that fit their location. However, recent the largest family among the G protein-coupled receptors whole-genome sequencing shows that ORs have been found (GPCRs) (8), constitute more than 1,000 genes on the mouse outside of the olfactory system, suggesting that ORs may play chromosome (9, 10) and more than 450 genes in the human an important role in the ectopic expression of non-chemo- genome (11, 12). sensory tissues. The ectopic expressions of ORs and their phys- Odorant activation shows a distinct signal transduction iological functions have attracted more attention recently since pathway for odorant perception. -

Sean Raspet – Molecules
1. Commercial name: Fructaplex© IUPAC Name: 2-(3,3-dimethylcyclohexyl)-2,5,5-trimethyl-1,3-dioxane SMILES: CC1(C)CCCC(C1)C2(C)OCC(C)(C)CO2 Molecular weight: 240.39 g/mol Volume (cubic Angstroems): 258.88 Atoms number (non-hydrogen): 17 miLogP: 4.43 Structure: Biological Properties: Predicted Druglikenessi: GPCR ligand -0.23 Ion channel modulator -0.03 Kinase inhibitor -0.6 Nuclear receptor ligand 0.15 Protease inhibitor -0.28 Enzyme inhibitor 0.15 Commercial name: Fructaplex© IUPAC Name: 2-(3,3-dimethylcyclohexyl)-2,5,5-trimethyl-1,3-dioxane SMILES: CC1(C)CCCC(C1)C2(C)OCC(C)(C)CO2 Predicted Olfactory Receptor Activityii: OR2L13 83.715% OR1G1 82.761% OR10J5 80.569% OR2W1 78.180% OR7A2 77.696% 2. Commercial name: Sylvoxime© IUPAC Name: N-[4-(1-ethoxyethenyl)-3,3,5,5tetramethylcyclohexylidene]hydroxylamine SMILES: CCOC(=C)C1C(C)(C)CC(CC1(C)C)=NO Molecular weight: 239.36 Volume (cubic Angstroems): 252.83 Atoms number (non-hydrogen): 17 miLogP: 4.33 Structure: Biological Properties: Predicted Druglikeness: GPCR ligand -0.6 Ion channel modulator -0.41 Kinase inhibitor -0.93 Nuclear receptor ligand -0.17 Protease inhibitor -0.39 Enzyme inhibitor 0.01 Commercial name: Sylvoxime© IUPAC Name: N-[4-(1-ethoxyethenyl)-3,3,5,5tetramethylcyclohexylidene]hydroxylamine SMILES: CCOC(=C)C1C(C)(C)CC(CC1(C)C)=NO Predicted Olfactory Receptor Activity: OR52D1 71.900% OR1G1 70.394% 0R52I2 70.392% OR52I1 70.390% OR2Y1 70.378% 3. Commercial name: Hyperflor© IUPAC Name: 2-benzyl-1,3-dioxan-5-one SMILES: O=C1COC(CC2=CC=CC=C2)OC1 Molecular weight: 192.21 g/mol Volume -
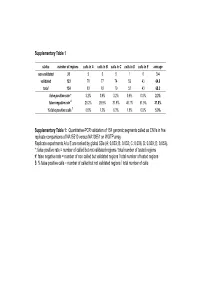
Quantitative-PCR Validation of 154 Genomic Segments Called As Cnvs in Five Replicat
Supplementary Table 1 status number of regions calls in A calls in B calls in C calls in D calls in E average non validated 31 5 6 5 1 0 3.4 validated 123 78 77 74 52 43 64.8 total 154 83 83 79 53 43 68.2 false positive rate * 3.2% 3.9% 3.2% 0.6% 0.0% 2.2% false negative rate # 29.2% 29.9% 31.8% 46.1% 51.9% 37.8% % false positive calls $ 6.0% 7.2% 6.3% 1.9% 0.0% 5.0% Supplementary Table 1: Quantitative-PCR validation of 154 genomic segments called as CNVs in five replicate comparisons of NA15510 versus NA10851 on WGTP array Replicate experiments A to E are ranked by global SDe (A: 0.033; B: 0.033; C: 0.036; D: 0.039; E: 0.053). *: false positive rate = number of called but not validated regions / total number of tested regions #: false negative rate = number of non called but validated regions / total number of tested regions $: % false positive calls = number of called but not validated regions / total number of calls False positive estimates for 500K EA CNV calls Total Rep1 Rep2 Rep3 Avg (unique) Validated 33 28 32 31 38 Not validated 2 2 2 2 5 Total 35 30 34 33 43 % False positive 5.71% 6.67% 5.88% 6.09% - % False negative 13.16% 26.32% 15.79% 18.42% - Supplementary Table 2A : Quantitative PCR validation of 43 unique CNV regions called as CNVs in three replicate comparisons of NA15510 versus NA10851 using the 500K EA array. -

Supplemental Files 1-5.Pdf
Supplemental file 1: The whole list of tumor aberrations by time tumor biopsy ctDNA 2002 (T1) 2012 (M1) 2015 (M2) 08/2015 09/2015 11/2015 02/2016 05/2016 07/2016 11/2016 02/2017 05/2017 07/2017 11/2017 PIK3CA Q546R PIK3CA Q546R PIK3CA Q546R PIK3CA Q546R 0.30% PIK3CA Q546R 1.30% PIK3CA Q546R 5.40% PIK3CA Q546R 12.50% PIK3CA Q546R 9.50% PIK3CA Q546R 32.58% PIK3CA Q546R 44.30% PIK3CA Q546R 15.40% PIK3CA Q546R 41.00% TP53 E180K TP53 E180K TP53 E180K TP53 E180K 0.30% TP53 E180K 1.00% TP53 E180K 4.80% PIK3CA M1043I 0.20% PIK3CA M1043I 0.70% PIK3CA E453K 0.69% PIK3CA E453K 3.20% PIK3CA E453K 0.55% PIK3CA M1043I 0.20% PIK3R2 S688* PIK3R2 S688* ATR E2579K VHL E173Q 0.40% PTEN Q245* 0.20% PTEN Q245* 0.30% PIK3CA E726K 1.50% PIK3CA E245Q 0.10% PIK3CA M1043I 0.36% PIK3CA M1043I 1.20% TP53 E180K 11.80% PIK3CA E453K 0.25% PLCG2 P737T PLCG2 P737T PIK3CA E726K VHL E173Q 1.10% RB1 R556* 3.10% TP53 E180K 11.00% PIK3CA E726K 0.20% PIK3CA E726K 0.23% PIK3CA E726K 0.44% TP53 I195F 1.20% PIK3CA E39Q 0.53% CDKN1B Q163* PIK3CA M1004I MET S1061F MET L229F 0.30% CCND1 E9K 0.10% PTEN Q245* 0.80% TP53 E180K 7.30% TP53 E180K 24.28% TP53 E180K 35.40% TP53 L137Q 1.30% TP53 E180K 38.40% ERBB2 R487W PTEN S229* TBX3 S435* MET W112C 1.20% NF1 D1556N 1.30% TP53 I195F 0.20% TP53 R175G 0.20% TP53 I195F 1.70% ESR1 E380Q 7.10% TP53 I195F 0.56% ALK E1407K RB1 Q217* SPEN Q743* NF1 R1176T 1.00% NF1 E76K 0.30% ESR1 E380Q 0.90% TP53 I195F 1.37% TP53 L137Q 2.10% ESR1 L391V 0.41% TP53 I254M 0.25% APC E2637K PIK3CA D1017N PTEN I253M VHL E173Q 1.20% RB1 R556* 7.20% NF1 R1176T 0.50% TP53 -

Supplementary Material: Human Adaptation to Arsenic-Rich Environments
Supplementary material: Human Adaptation to Arsenic-Rich Environments 1 15 Total SAC Sub SAC 10 density 5 0 0.0 0.2 0.4 0.6 iAs/AsAdj 25 20 15 density 10 5 0 0.00 0.05 0.10 0.15 0.20 MMA/AsAdj 3.0 2.0 density 1.0 0.0 0.0 0.5 1.0 1.5 2.0 DMA/AsAdj Figure S1: Phenotype density comparisons between the entire study population (385 women, in green) and the sample used for the association study and selection study (124 women, in blue), for 3 types of metabolites: inorganic arsenic (top panel), MMA (middle panel) and DMA (bottom panel), adjusted for the total arsenic content in urine. 2 Figure S2: Manhattan plots (FDR-corrected q-values) for %MMA (upper panel), %DMA (middle panel) and %inorganic arsenic from GWAS analyses. 3 Figure S3: Zoom-in of chromosome 10 region associated with %MMA and %DMA. 4 MMA Chr 3 MMA Chr 6 8 8 LOC440970 CADM2 LAMA2 BC035400 ARHGAP18C6orf191 L3MBTL3 SAMD3 TMEM200A 6 6 ● ● ● ● ● ● ● ● ● (P − value) ●●● ● (P − value) 4 ●● ●● ● 4 ● 10 ●● 10 ● ●●●●● ● ● ● ●●●●●●●●●●● ● ● ●●●● ● ●●●● ● ●●● ● ●●● ● ● ● ●●●● ●●● ● ● ●●●●● ● ●●●●● ● −log ● ● −log ● ●● ● ●● ●● ● ●●● ● ● ● ●●● ●●● ● ● ● ● ●●●●● ●●● ● ●● ● ● ● ● ●●● ●● ● ● ● ● ●●●●●●● ●● ● ●● ● ●●● ●● ● ●●● ● ● ● ● ●●● ● ●●● ●● ● ● ● ● ● ● ● ● 2 ● 2 ●●● ●●● ●●● ● ● ● ● ●●●●● ● ● ●● ● ● ●●● ● ●●● ●● ●●●●●●●●●● ● ● ●●● ●●● ●● ● ●●● ● ●● ●● ● ● ● ●●●●●●●● ● ●● ● ●● ● ●●● ●●●● ●●● ●● ● ●●● ● ● ● ● ● ● ●●●● ●● ●●●●● ●●● ● ● ● ● ● ● ●● ● ● ● ● ●●●● ● ●● ●● ● ● ●● ● ●●●● ● ●● ● ● ● ● ● ●● ● ● ● ● ● ●●●● ●●●● ● ●● ●● ●●●●●● ● ● ●●●●● ● ●●● ●● ● ● ● ● ●● ● ●● ●● ● ● ● ●●● ● ●●●● ●●