Copy Number Variants and Infantile Spasms: Evidence for Abnormalities in Ventral Forebrain Development and Pathways of Synaptic Function
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genome Wide Association Study of Response to Interval and Continuous Exercise Training: the Predict‑HIIT Study Camilla J
Williams et al. J Biomed Sci (2021) 28:37 https://doi.org/10.1186/s12929-021-00733-7 RESEARCH Open Access Genome wide association study of response to interval and continuous exercise training: the Predict-HIIT study Camilla J. Williams1†, Zhixiu Li2†, Nicholas Harvey3,4†, Rodney A. Lea4, Brendon J. Gurd5, Jacob T. Bonafglia5, Ioannis Papadimitriou6, Macsue Jacques6, Ilaria Croci1,7,20, Dorthe Stensvold7, Ulrik Wislof1,7, Jenna L. Taylor1, Trishan Gajanand1, Emily R. Cox1, Joyce S. Ramos1,8, Robert G. Fassett1, Jonathan P. Little9, Monique E. Francois9, Christopher M. Hearon Jr10, Satyam Sarma10, Sylvan L. J. E. Janssen10,11, Emeline M. Van Craenenbroeck12, Paul Beckers12, Véronique A. Cornelissen13, Erin J. Howden14, Shelley E. Keating1, Xu Yan6,15, David J. Bishop6,16, Anja Bye7,17, Larisa M. Haupt4, Lyn R. Grifths4, Kevin J. Ashton3, Matthew A. Brown18, Luciana Torquati19, Nir Eynon6 and Jef S. Coombes1* Abstract Background: Low cardiorespiratory ftness (V̇O2peak) is highly associated with chronic disease and mortality from all causes. Whilst exercise training is recommended in health guidelines to improve V̇O2peak, there is considerable inter-individual variability in the V̇O2peak response to the same dose of exercise. Understanding how genetic factors contribute to V̇O2peak training response may improve personalisation of exercise programs. The aim of this study was to identify genetic variants that are associated with the magnitude of V̇O2peak response following exercise training. Methods: Participant change in objectively measured V̇O2peak from 18 diferent interventions was obtained from a multi-centre study (Predict-HIIT). A genome-wide association study was completed (n 507), and a polygenic predictor score (PPS) was developed using alleles from single nucleotide polymorphisms= (SNPs) signifcantly associ- –5 ated (P < 1 10 ) with the magnitude of V̇O2peak response. -

Circular RNA Hsa Circ 0005114‑Mir‑142‑3P/Mir‑590‑5P‑ Adenomatous
ONCOLOGY LETTERS 21: 58, 2021 Circular RNA hsa_circ_0005114‑miR‑142‑3p/miR‑590‑5p‑ adenomatous polyposis coli protein axis as a potential target for treatment of glioma BO WEI1*, LE WANG2* and JINGWEI ZHAO1 1Department of Neurosurgery, China‑Japan Union Hospital of Jilin University, Changchun, Jilin 130033; 2Department of Ophthalmology, The First Hospital of Jilin University, Jilin University, Changchun, Jilin 130021, P.R. China Received September 12, 2019; Accepted October 22, 2020 DOI: 10.3892/ol.2020.12320 Abstract. Glioma is the most common type of brain tumor APC expression with a good overall survival rate. UALCAN and is associated with a high mortality rate. Despite recent analysis using TCGA data of glioblastoma multiforme and the advances in treatment options, the overall prognosis in patients GSE25632 and GSE103229 microarray datasets showed that with glioma remains poor. Studies have suggested that circular hsa‑miR‑142‑3p/hsa‑miR‑590‑5p was upregulated and APC (circ)RNAs serve important roles in the development and was downregulated. Thus, hsa‑miR‑142‑3p/hsa‑miR‑590‑5p‑ progression of glioma and may have potential as therapeutic APC‑related circ/ceRNA axes may be important in glioma, targets. However, the expression profiles of circRNAs and their and hsa_circ_0005114 interacted with both of these miRNAs. functions in glioma have rarely been studied. The present study Functional analysis showed that hsa_circ_0005114 was aimed to screen differentially expressed circRNAs (DECs) involved in insulin secretion, while APC was associated with between glioma and normal brain tissues using sequencing the Wnt signaling pathway. In conclusion, hsa_circ_0005114‑ data collected from the Gene Expression Omnibus database miR‑142‑3p/miR‑590‑5p‑APC ceRNA axes may be potential (GSE86202 and GSE92322 datasets) and explain their mecha‑ targets for the treatment of glioma. -
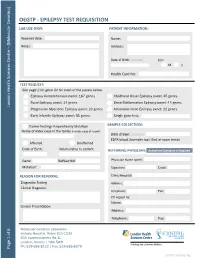
Oegtp - Epilepsy Test Requisition Lab Use Only: Patient Information
OEGTP - EPILEPSY TEST REQUISITION LAB USE ONLY: PATIENT INFORMATION: Received date: Name: Notes: Address: Date of Birth: YY/MM/DD Sex: M F Health Card No: TEST REQUEST: See page 2 for gene list for each of the panels below Epilepsy Comprehensive panel: 167 genes Childhood Onset Epilepsy panel: 45 genes Focal Epilepsy panel: 14 genes Brain Malformation Epilepsy panel: 44 genes London Health Sciences Centre – (Molecular Genetics) London Health Sciences Centre Progressive Myoclonic Epilepsy panel: 20 genes Actionable Gene Epilepsy panel: 22 genes Early Infantile Epilepsy panel: 51 genes Single gene test: Carrier Testing/ KnownFamily Mutation SAMPLE COLLECTION: Name of index case in the family (include copy of report) Date drawn: YY/MM/DD EDTA blood (lavender top) (5ml at room temp) Affected Unaffected Date of Birth: Relationship to patient: REFERRING PHYSICIAN: Authorized Signature is Required Gene: RefSeq:NM Physician Name (print): Mutation: Signature: Email: REASON FOR REFERRAL: Clinic/Hospital: Diagnostic Testing Address: Clinical Diagnosis: Telephone: Fax: CC report to: Name: Clinical Presentation: Address: Telephone: Fax: Molecular Genetics Laboratory Victoria Hospital, Room B10-123A 800 Commissioners Rd. E. London, Ontario | N6A 5W9 Pathology and Laboratory Medicine Ph: 519-685-8122 | Fax: 519-685-8279 Page 1 of 6 Page OEGTP (2021/05/28) OEGTP - EPILEPSY TEST PANELS Patient Identifier: COMPREHENSIVE EPILEPSY PANEL: 167 Genes ACTB, ACTG1, ADSL, AKT3, ALDH7A1, AMT, AP3B2, ARFGEF2, ARHGEF9, ARV1, ARX, ASAH1, ASNS, ATP1A3, ATP6V0A2, ATP7A, -

Characterization of Zebrafish GABAA Receptor Subunits
www.nature.com/scientificreports OPEN Characterization of zebrafsh GABAA receptor subunits Kenichiro Sadamitsu, Leona Shigemitsu, Marina Suzuki, Daishi Ito, Makoto Kashima & Hiromi Hirata* γ-Aminobutyric acid (GABA), the major inhibitory neurotransmitter in the central nervous system, exerts its efect through the activation of GABA receptors. GABAA receptors are ligand-gated chloride channels composed of fve subunit proteins. Mammals have 19 diferent GABAA receptor subunits (α1–6, β1–3, γ1–3, δ, ε, π, θ, and ρ1–3), the physiological properties of which have been assayed by electrophysiology. However, the evolutionary conservation of the physiological characteristics of diverged GABAA receptor subunits remains unclear. Zebrafsh have 23 subunits (α1, α2a, α2b, α3–5, α6a, α6b, β1–4, γ1–3, δ, π, ζ, ρ1, ρ2a, ρ2b, ρ3a, and ρ3b), but the electrophysiological properties of these subunits have not been explored. In this study, we cloned the coding sequences for zebrafsh GABAA receptor subunits and investigated their expression patterns in larval zebrafsh by whole- mount in situ hybridization. We also performed electrophysiological recordings of GABA-evoked currents from Xenopus oocytes injected with one or multiple zebrafsh GABAA receptor subunit cRNAs and calculated the half-maximal efective concentrations (EC50s) for each. Our results revealed the spatial expressions and electrophysiological GABA sensitivities of zebrafsh GABAA receptors, suggesting that the properties of GABAA receptor subunits are conserved among vertebrates. γ-Aminobutyric acid (GABA), the major inhibitory neurotransmitter in the central nervous system of vertebrates, 1 controls the excitability of neural networks mainly through GABA A receptors . Te GABAA receptor mediates two types of inhibition, known as phasic and tonic inhibition2. -

The Landscape of Genomic Imprinting Across Diverse Adult Human Tissues
Downloaded from genome.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press Research The landscape of genomic imprinting across diverse adult human tissues Yael Baran,1 Meena Subramaniam,2 Anne Biton,2 Taru Tukiainen,3,4 Emily K. Tsang,5,6 Manuel A. Rivas,7 Matti Pirinen,8 Maria Gutierrez-Arcelus,9 Kevin S. Smith,5,10 Kim R. Kukurba,5,10 Rui Zhang,10 Celeste Eng,2 Dara G. Torgerson,2 Cydney Urbanek,11 the GTEx Consortium, Jin Billy Li,10 Jose R. Rodriguez-Santana,12 Esteban G. Burchard,2,13 Max A. Seibold,11,14,15 Daniel G. MacArthur,3,4,16 Stephen B. Montgomery,5,10 Noah A. Zaitlen,2,19 and Tuuli Lappalainen17,18,19 1The Blavatnik School of Computer Science, Tel-Aviv University, Tel Aviv 69978, Israel; 2Department of Medicine, University of California San Francisco, San Francisco, California 94158, USA; 3Analytic and Translational Genetics Unit, Massachusetts General Hospital, Boston, Massachusetts 02114, USA; 4Program in Medical and Population Genetics, Broad Institute of Harvard and MIT, Cambridge, Massachusetts 02142, USA; 5Department of Pathology, Stanford University, Stanford, California 94305, USA; 6Biomedical Informatics Program, Stanford University, Stanford, California 94305, USA; 7Wellcome Trust Center for Human Genetics, Nuffield Department of Clinical Medicine, University of Oxford, Oxford, OX3 7BN, United Kingdom; 8Institute for Molecular Medicine Finland, University of Helsinki, 00014 Helsinki, Finland; 9Department of Genetic Medicine and Development, University of Geneva, 1211 Geneva, Switzerland; -

Probedealer Is a Convenient Tool for Designing Probes for Highly Multiplexed Fuorescence in Situ Hybridization Mengwei Hu1,9, Bing Yang1,9, Yubao Cheng1, Jonathan S
www.nature.com/scientificreports OPEN ProbeDealer is a convenient tool for designing probes for highly multiplexed fuorescence in situ hybridization Mengwei Hu1,9, Bing Yang1,9, Yubao Cheng1, Jonathan S. D. Radda1, Yanbo Chen1, Miao Liu1 & Siyuan Wang1,2,3,4,5,6,7,8* Fluorescence in situ hybridization (FISH) is a powerful method to visualize the spatial positions of specifc genomic loci and RNA species. Recent technological advances have leveraged FISH to visualize these features in a highly multiplexed manner. Notable examples include chromatin tracing, RNA multiplexed error-robust FISH (MERFISH), multiplexed imaging of nucleome architectures (MINA), and sequential single-molecule RNA FISH. However, one obstacle to the broad adoption of these methods is the complexity of the multiplexed FISH probe design. In this paper, we introduce an easy-to-use, versatile, and all-in-one application called ProbeDealer to design probes for a variety of multiplexed FISH techniques and their combinations. ProbeDealer ofers a one-stop shop for multiplexed FISH design needs of the research community. Fluorescence in situ hybridization (FISH) can be used to visualize spatial locations of DNA regions and RNA molecules in a sequence-specifc manner. Recently, FISH has been extended with multiplicity to profle chroma- tin folding pattern and the abundance of numerous transcripts in several methods. Tese include multiplexed sequential DNA FISH 1–6 (termed chromatin tracing), multiplexed error-robust FISH (MERFISH)7–10 and similar methods11,12, and sequential single-molecule RNA FISH (sequentially imaging individual RNA species without combinatorial barcoding)4,9. Chromatin tracing has been combined with single-molecule RNA FISH to study the association between gene expression regulation and chromatin folding 4,5. -

Research Article Microarray-Based Comparisons of Ion Channel Expression Patterns: Human Keratinocytes to Reprogrammed Hipscs To
Hindawi Publishing Corporation Stem Cells International Volume 2013, Article ID 784629, 25 pages http://dx.doi.org/10.1155/2013/784629 Research Article Microarray-Based Comparisons of Ion Channel Expression Patterns: Human Keratinocytes to Reprogrammed hiPSCs to Differentiated Neuronal and Cardiac Progeny Leonhard Linta,1 Marianne Stockmann,1 Qiong Lin,2 André Lechel,3 Christian Proepper,1 Tobias M. Boeckers,1 Alexander Kleger,3 and Stefan Liebau1 1 InstituteforAnatomyCellBiology,UlmUniversity,Albert-EinsteinAllee11,89081Ulm,Germany 2 Institute for Biomedical Engineering, Department of Cell Biology, RWTH Aachen, Pauwelstrasse 30, 52074 Aachen, Germany 3 Department of Internal Medicine I, Ulm University, Albert-Einstein Allee 11, 89081 Ulm, Germany Correspondence should be addressed to Alexander Kleger; [email protected] and Stefan Liebau; [email protected] Received 31 January 2013; Accepted 6 March 2013 Academic Editor: Michael Levin Copyright © 2013 Leonhard Linta et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Ion channels are involved in a large variety of cellular processes including stem cell differentiation. Numerous families of ion channels are present in the organism which can be distinguished by means of, for example, ion selectivity, gating mechanism, composition, or cell biological function. To characterize the distinct expression of this group of ion channels we have compared the mRNA expression levels of ion channel genes between human keratinocyte-derived induced pluripotent stem cells (hiPSCs) and their somatic cell source, keratinocytes from plucked human hair. This comparison revealed that 26% of the analyzed probes showed an upregulation of ion channels in hiPSCs while just 6% were downregulated. -

Assessment of Molecular Action of Direct Gating and Allosteric Modulatory Effects of Carisoprodol (Somartm) on GABA a Receptors
Graduate Theses, Dissertations, and Problem Reports 2015 Assessment of molecular action of direct gating and allosteric modulatory effects of carisoprodol (SomaRTM) on GABA A receptors Manoj Kumar Follow this and additional works at: https://researchrepository.wvu.edu/etd Recommended Citation Kumar, Manoj, "Assessment of molecular action of direct gating and allosteric modulatory effects of carisoprodol (SomaRTM) on GABA A receptors" (2015). Graduate Theses, Dissertations, and Problem Reports. 6022. https://researchrepository.wvu.edu/etd/6022 This Dissertation is protected by copyright and/or related rights. It has been brought to you by the The Research Repository @ WVU with permission from the rights-holder(s). You are free to use this Dissertation in any way that is permitted by the copyright and related rights legislation that applies to your use. For other uses you must obtain permission from the rights-holder(s) directly, unless additional rights are indicated by a Creative Commons license in the record and/ or on the work itself. This Dissertation has been accepted for inclusion in WVU Graduate Theses, Dissertations, and Problem Reports collection by an authorized administrator of The Research Repository @ WVU. For more information, please contact [email protected]. ASSESSMENT OF MOLECULAR ACTION OF DIRECT GATING AND ALLOSTERIC MODULATORY EFFECTS OF MEPROBAMATE (MILTOWN®) ON GABAA RECEPTORS Manish Kumar, MD, MS Dissertation submitted to the School of Pharmacy at West Virginia University in partial fulfillment of Requirements -

Exploiting the Genome
Exploiting the Genome Lead Author G.Joyce Contributors S. Block J. Cornwall F. Dyson S. Koonin N. Lewis R. Schwitters September 1998 JSR-98-315 Approved for public release; distribution unlimited. JASON The MITRE Corporation 1820 Dolley Madison Boulevard McLean. Virginia 22102-3481 (703) 883-6997 Form Approved REPORT DOCUMENTATION PAGE OMS No. 0704-0188 Public reporting burden lor this collection 01 Inlormation estimated to average 1 hour per response, including the time lor review Instructions, searc hlng eldsting data sources, gathering and maintaining the data needed, and completing and reviewing the collection 01 Inlormatlon. Send comments regarding this burden estimate or any other aspect 01 this collection 01 Inlormation, Including suggestions for reducing this burden, to Washington Headquarters Services, Directorate lor Information Operations and Reports, 1215 Jefferson Davis Highway, SuHe 1204, Arlington, VA 22202·4302, and to the Office 01 Management and Budget. Paperwori< Reduction Project (0704·0188). Washington. DC 20503. 1. AGENCY USE ONLY (Leave blank) 12. REPORT DATE r' REPORT TYPE AND DATES COVERED September 11, 1998 4. TITLE AND SUBTITLE 5. FUNDING NUMBERS Exploiting the Genome 13-958534-04 6. AUTHOR(S) S. Block, J. Cornwall, F. Dyson, G. Joyce, S. Koonin, N. Lewis, R. Schwitters 7. PERFORMING ORGANIZATION NAME(S) AND ADDRESS(ES) 8. PERFORMING ORGANIZATION The MITRE Corporation REPORT NUMBER JASON Program Office 1820 Dolley Madison Blvd JSR-98-315 McLean, Virginia 22102 9. SPONSORINGIMONITORING AGENCY NAME(S) AND ADDRESS(ES) 10. SPONSORING/MONITORING AGENCY REPORT NUMBER US Department of Energy Biological and Environmental Research 1901 Germantown Road JSR-98-315 Ge~town,~ 20874-1290 11. -

Epileptic Mechanisms Shared by Alzheimer's Disease
International Journal of Molecular Sciences Review Epileptic Mechanisms Shared by Alzheimer’s Disease: Viewed via the Unique Lens of Genetic Epilepsy Jing-Qiong Kang 1,2 1 Department of Neurology & Pharmacology, Vanderbilt University Medical Center, Nashville, TN 37232-8552, USA; [email protected]; Tel.: +1-615-936-8399; Fax: +1-615-322-5517 2 Vanderbilt Kennedy Center of Human Development, Vanderbilt Brain Institute, Vanderbilt University, Nashville, TN 37232-8522, USA Abstract: Our recent work on genetic epilepsy (GE) has identified common mechanisms between GE and neurodegenerative diseases including Alzheimer’s disease (AD). Although both disorders are seemingly unrelated and occur at opposite ends of the age spectrum, it is likely there are shared mechanisms and studies on GE could provide unique insights into AD pathogenesis. Neurodegen- erative diseases are typically late-onset disorders, but the underlying pathology may have already occurred long before the clinical symptoms emerge. Pathophysiology in the early phase of these diseases is understudied but critical for developing mechanism-based treatment. In AD, increased seizure susceptibility and silent epileptiform activity due to disrupted excitatory/inhibitory (E/I) balance has been identified much earlier than cognition deficit. Increased epileptiform activity is likely a main pathology in the early phase that directly contributes to impaired cognition. It is an enormous challenge to model the early phase of pathology with conventional AD mouse models due to the chronic disease course, let alone the complex interplay between subclinical nonconvulsive epileptiform activity, AD pathology, and cognition deficit. We have extensively studied GE, especially with gene mutations that affect the GABA pathway such as mutations in GABAA receptors and GABA transporter Citation: Kang, J.-Q. -

Expression of GABAA Α2-, Β1- And
OPEN Citation: Transl Psychiatry (2013) 3, e303; doi:10.1038/tp.2013.64 & 2013 Macmillan Publishers Limited All rights reserved 2158-3188/13 www.nature.com/tp ORIGINAL ARTICLE Expression of GABAA a2-, b1- and e-receptors are altered significantly in the lateral cerebellum of subjects with schizophrenia, major depression and bipolar disorder SH Fatemi1,2,3, TD Folsom1, RJ Rooney4 and PD Thuras5 There is abundant evidence that dysfunction of the g-aminobutyric acid (GABA)ergic signaling system is implicated in the pathology of schizophrenia and mood disorders. Less is known about the alterations in protein expression of GABA receptor subunits in brains of subjects with schizophrenia and mood disorders. We have previously demonstrated reduced expression of GABAB receptor subunits 1 and 2 (GABBR1 and GABBR2) in the lateral cerebella of subjects with schizophrenia, bipolar disorder and major depressive disorder. In the current study, we have expanded these studies to examine the mRNA and protein expression of 12 GABAA subunit proteins (a1, a2, a3, a5, a6, b1, b2, b3, d, e, g2 and g3) in the lateral cerebella from the same set of subjects with schizophrenia (N ¼ 9–15), bipolar disorder (N ¼ 10–15) and major depression (N ¼ 12–15) versus healthy controls (N ¼ 10–15). We found significant group effects for protein levels of the a2-, b1- and e-subunits across treatment groups. We also found a significant group effect for mRNA levels of the a1-subunit across treatment groups. New avenues for treatment, such as the use of neurosteroids to promote GABA modulation, could potentially ameliorate GABAergic dysfunction in these disorders. -

Ion Channels
UC Davis UC Davis Previously Published Works Title THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels. Permalink https://escholarship.org/uc/item/1442g5hg Journal British journal of pharmacology, 176 Suppl 1(S1) ISSN 0007-1188 Authors Alexander, Stephen PH Mathie, Alistair Peters, John A et al. Publication Date 2019-12-01 DOI 10.1111/bph.14749 License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: Ion channels. British Journal of Pharmacology (2019) 176, S142–S228 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels Stephen PH Alexander1 , Alistair Mathie2 ,JohnAPeters3 , Emma L Veale2 , Jörg Striessnig4 , Eamonn Kelly5, Jane F Armstrong6 , Elena Faccenda6 ,SimonDHarding6 ,AdamJPawson6 , Joanna L Sharman6 , Christopher Southan6 , Jamie A Davies6 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 3Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 4Pharmacology and Toxicology, Institute of Pharmacy, University of Innsbruck, A-6020 Innsbruck, Austria 5School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 6Centre for Discovery Brain Science, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties.