Molecular Control of Retinal Ganglion Cell Specification and Differentiation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Genome-Wide DNA Methylation Analysis of KRAS Mutant Cell Lines Ben Yi Tew1,5, Joel K
www.nature.com/scientificreports OPEN Genome-wide DNA methylation analysis of KRAS mutant cell lines Ben Yi Tew1,5, Joel K. Durand2,5, Kirsten L. Bryant2, Tikvah K. Hayes2, Sen Peng3, Nhan L. Tran4, Gerald C. Gooden1, David N. Buckley1, Channing J. Der2, Albert S. Baldwin2 ✉ & Bodour Salhia1 ✉ Oncogenic RAS mutations are associated with DNA methylation changes that alter gene expression to drive cancer. Recent studies suggest that DNA methylation changes may be stochastic in nature, while other groups propose distinct signaling pathways responsible for aberrant methylation. Better understanding of DNA methylation events associated with oncogenic KRAS expression could enhance therapeutic approaches. Here we analyzed the basal CpG methylation of 11 KRAS-mutant and dependent pancreatic cancer cell lines and observed strikingly similar methylation patterns. KRAS knockdown resulted in unique methylation changes with limited overlap between each cell line. In KRAS-mutant Pa16C pancreatic cancer cells, while KRAS knockdown resulted in over 8,000 diferentially methylated (DM) CpGs, treatment with the ERK1/2-selective inhibitor SCH772984 showed less than 40 DM CpGs, suggesting that ERK is not a broadly active driver of KRAS-associated DNA methylation. KRAS G12V overexpression in an isogenic lung model reveals >50,600 DM CpGs compared to non-transformed controls. In lung and pancreatic cells, gene ontology analyses of DM promoters show an enrichment for genes involved in diferentiation and development. Taken all together, KRAS-mediated DNA methylation are stochastic and independent of canonical downstream efector signaling. These epigenetically altered genes associated with KRAS expression could represent potential therapeutic targets in KRAS-driven cancer. Activating KRAS mutations can be found in nearly 25 percent of all cancers1. -

SUPPLEMENTARY MATERIAL Bone Morphogenetic Protein 4 Promotes
www.intjdevbiol.com doi: 10.1387/ijdb.160040mk SUPPLEMENTARY MATERIAL corresponding to: Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells SUMIYO MIMURA, MIKA SUGA, KAORI OKADA, MASAKI KINEHARA, HIROKI NIKAWA and MIHO K. FURUE* *Address correspondence to: Miho Kusuda Furue. Laboratory of Stem Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, 7-6-8, Saito-Asagi, Ibaraki, Osaka 567-0085, Japan. Tel: 81-72-641-9819. Fax: 81-72-641-9812. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.160040mk TABLE S1 PRIMER LIST FOR QRT-PCR Gene forward reverse AP2α AATTTCTCAACCGACAACATT ATCTGTTTTGTAGCCAGGAGC CDX2 CTGGAGCTGGAGAAGGAGTTTC ATTTTAACCTGCCTCTCAGAGAGC DLX1 AGTTTGCAGTTGCAGGCTTT CCCTGCTTCATCAGCTTCTT FOXD3 CAGCGGTTCGGCGGGAGG TGAGTGAGAGGTTGTGGCGGATG GAPDH CAAAGTTGTCATGGATGACC CCATGGAGAAGGCTGGGG MSX1 GGATCAGACTTCGGAGAGTGAACT GCCTTCCCTTTAACCCTCACA NANOG TGAACCTCAGCTACAAACAG TGGTGGTAGGAAGAGTAAAG OCT4 GACAGGGGGAGGGGAGGAGCTAGG CTTCCCTCCAACCAGTTGCCCCAAA PAX3 TTGCAATGGCCTCTCAC AGGGGAGAGCGCGTAATC PAX6 GTCCATCTTTGCTTGGGAAA TAGCCAGGTTGCGAAGAACT p75 TCATCCCTGTCTATTGCTCCA TGTTCTGCTTGCAGCTGTTC SOX9 AATGGAGCAGCGAAATCAAC CAGAGAGATTTAGCACACTGATC SOX10 GACCAGTACCCGCACCTG CGCTTGTCACTTTCGTTCAG Suppl. Fig. S1. Comparison of the gene expression profiles of the ES cells and the cells induced by NC and NC-B condition. Scatter plots compares the normalized expression of every gene on the array (refer to Table S3). The central line -

Proquest Dissertations
RICE UNIVERSITY Molecular Basis of Gene Dosage Sensitivity by Jianping Chen A THESIS SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE Doctor of Philosophy APPROVED, THESIS COMMITTEE: Ariel Fernandez, Chair Karl F. Hasselmann Professor Department of Bioengineering Rice University rn'idtfP h.tQsJUW*- Michael W. Deem, John W. Cox Professor Department of Bioengineering Department of Physics and Astronomy Rice Universit ^engineering at Rice University Jiochemistry at Baylor College of Medicine Axxttig- «J^gb/uv Laura Segaton, T.N. Law Assistant Professor Chemical and Biomolecular Engineering Rice University HOUSTON, TEXAS JANUARY 2009 UMI Number: 3362141 INFORMATION TO USERS The quality of this reproduction is dependent upon the quality of the copy submitted. Broken or indistinct print, colored or poor quality illustrations and photographs, print bleed-through, substandard margins, and improper alignment can adversely affect reproduction. In the unlikely event that the author did not send a complete manuscript and there are missing pages, these will be noted. Also, if unauthorized copyright material had to be removed, a note will indicate the deletion. UMI® UMI Microform 3362141 Copyright 2009 by ProQuest LLC All rights reserved. This microform edition is protected against unauthorized copying under Title 17, United States Code. ProQuest LLC 789 East Eisenhower Parkway P.O. Box 1346 Ann Arbor, Ml 48106-1346 ABSTRACT Molecular Basis of Gene Dosage Sensitivity by JianpingChen Deviation of gene expression from normal levels has been associated with diseases. Both under- and overexpression of genes could lead to deleterious biological consequences. Dosage balance has been proposed to be a key issue of determining gene expression pheno- type. -

A Three-Dimensional Organoid Model Recapitulates Tumorigenic Aspects
www.nature.com/scientificreports OPEN A three-dimensional organoid model recapitulates tumorigenic aspects and drug responses of Received: 22 June 2018 Accepted: 10 October 2018 advanced human retinoblastoma Published: xx xx xxxx Duangporn Saengwimol1, Duangnate Rojanaporn2, Vijender Chaitankar3, Pamorn Chittavanich4, Rangsima Aroonroch5, Tatpong Boontawon4, Weerin Thammachote4, Natini Jinawath4, Suradej Hongeng6 & Rossukon Kaewkhaw4 Persistent or recurrent retinoblastoma (RB) is associated with the presence of vitreous or/and subretinal seeds in advanced RB and represents a major cause of therapeutic failure. This necessitates the development of novel therapies and thus requires a model of advanced RB for testing candidate therapeutics. To this aim, we established and characterized a three-dimensional, self-organizing organoid model derived from chemotherapy-naïve tumors. The responses of organoids to drugs were determined and compared to relate organoid model to advanced RB, in terms of drug sensitivities. We found that organoids had histological features resembling retinal tumors and seeds and retained DNA copy-number alterations as well as gene and protein expression of the parental tissue. Cone signal circuitry (M/L+ cells) and glial tumor microenvironment (GFAP+ cells) were primarily present in organoids. Topotecan alone or the combined drug regimen of topotecan and melphalan efectively targeted proliferative tumor cones (RXRγ+ Ki67+) in organoids after 24-h drug exposure, blocking mitotic entry. In contrast, methotrexate showed the least efcacy against tumor cells. The drug responses of organoids were consistent with those of tumor cells in advanced disease. Patient-derived organoids enable the creation of a faithful model to use in examining novel therapeutics for RB. Retinoblastoma (RB) is a serious childhood retinal tumor that, if lef untreated, can cause death within 1–2 years. -

Single Cell Derived Clonal Analysis of Human Glioblastoma Links
SUPPLEMENTARY INFORMATION: Single cell derived clonal analysis of human glioblastoma links functional and genomic heterogeneity ! Mona Meyer*, Jüri Reimand*, Xiaoyang Lan, Renee Head, Xueming Zhu, Michelle Kushida, Jane Bayani, Jessica C. Pressey, Anath Lionel, Ian D. Clarke, Michael Cusimano, Jeremy Squire, Stephen Scherer, Mark Bernstein, Melanie A. Woodin, Gary D. Bader**, and Peter B. Dirks**! ! * These authors contributed equally to this work.! ** Correspondence: [email protected] or [email protected]! ! Supplementary information - Meyer, Reimand et al. Supplementary methods" 4" Patient samples and fluorescence activated cell sorting (FACS)! 4! Differentiation! 4! Immunocytochemistry and EdU Imaging! 4! Proliferation! 5! Western blotting ! 5! Temozolomide treatment! 5! NCI drug library screen! 6! Orthotopic injections! 6! Immunohistochemistry on tumor sections! 6! Promoter methylation of MGMT! 6! Fluorescence in situ Hybridization (FISH)! 7! SNP6 microarray analysis and genome segmentation! 7! Calling copy number alterations! 8! Mapping altered genome segments to genes! 8! Recurrently altered genes with clonal variability! 9! Global analyses of copy number alterations! 9! Phylogenetic analysis of copy number alterations! 10! Microarray analysis! 10! Gene expression differences of TMZ resistant and sensitive clones of GBM-482! 10! Reverse transcription-PCR analyses! 11! Tumor subtype analysis of TMZ-sensitive and resistant clones! 11! Pathway analysis of gene expression in the TMZ-sensitive clone of GBM-482! 11! Supplementary figures and tables" 13" "2 Supplementary information - Meyer, Reimand et al. Table S1: Individual clones from all patient tumors are tumorigenic. ! 14! Fig. S1: clonal tumorigenicity.! 15! Fig. S2: clonal heterogeneity of EGFR and PTEN expression.! 20! Fig. S3: clonal heterogeneity of proliferation.! 21! Fig. -

Pou4f1 and Pou4f2 Are Dispensable for the Long-Term Survival of Adult Retinal Ganglion Cells in Mice
Pou4f1 and Pou4f2 Are Dispensable for the Long-Term Survival of Adult Retinal Ganglion Cells in Mice Liang Huang1,2, Fang Hu1,2, Xiaoling Xie1, Jeffery Harder1, Kimberly Fernandes1, Xiang-yun Zeng2, Richard Libby1, Lin Gan1,3* 1 Flaum Eye Institute and Department of Ophthalmology, University of Rochester School of Medicine and Dentistry, Rochester, New York, United States of America, 2 Department of Ophthalmology, First Affiliated Hospital of Gannan Medical University, Ganzhou, Jiangxi, China, 3 College of Life and Environmental Sciences, Hangzhou Normal University, Hangzhou, Zhejiang, China Abstract Purpose: To investigate the role of Pou4f1 and Pou4f2 in the survival of adult retinal ganglion cells (RGCs). Methods: Conditional alleles of Pou4f1 and Pou4f2 were generated (Pou4f1loxP and Pou4f2loxP respectively) for the removal of Pou4f1 and Pou4f2 in adult retinas. A tamoxifen-inducible Cre was used to delete Pou4f1 and Pou4f2 in adult mice and retinal sections and flat mounts were subjected to immunohistochemistry to confirm the deletion of both alleles and to quantify the changes in the number of RGCs and other retinal neurons. To determine the effect of loss of Pou4f1 and Pou4f2 on RGC survival after axonal injury, controlled optic nerve crush (CONC) was performed and RGC death was assessed. Results: Pou4f1 and Pou4f2 were ablated two weeks after tamoxifen treatment. Retinal interneurons and Mu¨ller glial cells are not affected by the ablation of Pou4f1 or Pou4f2 or both. Although the deletion of both Pou4f1 and Pou4f2 slightly delays the death of RGCs at 3 days post-CONC in adult mice, it does not affect the cell death progress afterwards. -
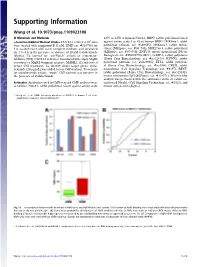
Supporting Information
Supporting Information Wang et al. 10.1073/pnas.1109023108 SI Materials and Methods 2278 to 2470 of human Notch1; RBPJ, rabbit polyclonal raised γ-Secretase-Inhibitor Washout Studies. CUTLL1 cells (2 × 105/mL) against amino acids 1 to 48 of human RBPJ; H3K4me1, rabbit were treated with compound E (1 μM, EMD cat. #565790) for polyclonal (Abcam, cat. #ab8895); H3K4me3, rabbit mono- 3 d, washed twice with warm complete medium, and incubated clonal (Millipore, cat. #04–745); H3K27me3, rabbit polyclonal for 2 to 4 h in the presence or absence of 20 μM cycloheximide (Millipore, cat. #07–449); ZNF143, mouse monoclonal (Novus (Sigma). To control for “off-Notch” effects of γ-secretase- Biologicals, cat. #H00007702-M01); GABPA, rabbit polyclonal inhibitor (GSI), CUTLL1 cells were transduced with empty MigRI (Santa Cruz Biotechnology, cat. #sc-22810); RUNX1, rabbit retrovirus or MigRI-dominant negative MAML1 (1) and sorted polyclonal (Abcam, cat. #ab23980); ETS1, rabbit polyclon- before GSI treatment. To identify direct target genes, cyclo- al (Santa Cruz Biotechnology, cat. #sc-350); CREB, rabbit heximide (20 μg/mL) was added before GSI washout. To control monoclonal (Cell Signaling Technology, cat. #9197); REST, for cycloheximide effects, “mock” GSI washout was perform in rabbit polyclonal (Santa Cruz Biotechnology, cat. #sc-25398); the presence of cycloheximide. mouse nonimmune IgG (Millipore, cat. #12–371). Western blot analysis was performed with the antibodies above or rabbit an- Antibodies. Antibodies used in ChIP-seq and ChIP analyses were tiactivated Notch1 (Cell Signaling Technology, cat. #2412), and as follows: Notch1, rabbit polyclonal raised against amino acids mouse anti–β-actin (Sigma). -

Identifying the Risky SNP of Osteoporosis with ID3-PEP Decision Tree Algorithm
Hindawi Complexity Volume 2017, Article ID 9194801, 8 pages https://doi.org/10.1155/2017/9194801 Research Article Identifying the Risky SNP of Osteoporosis with ID3-PEP Decision Tree Algorithm Jincai Yang,1 Huichao Gu,1 Xingpeng Jiang,1 Qingyang Huang,2 Xiaohua Hu,1 and Xianjun Shen1 1 School of Computer Science, Central China Normal University, Wuhan 430079, China 2School of Life Science, Central China Normal University, Wuhan 430079, China Correspondence should be addressed to Jincai Yang; [email protected] Received 31 March 2017; Revised 26 May 2017; Accepted 8 June 2017; Published 7 August 2017 Academic Editor: Fang-Xiang Wu Copyright © 2017 Jincai Yang et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. In the past 20 years, much progress has been made on the genetic analysis of osteoporosis. A number of genes and SNPs associated with osteoporosis have been found through GWAS method. In this paper, we intend to identify the suspected risky SNPs of osteoporosis with computational methods based on the known osteoporosis GWAS-associated SNPs. The process includes two steps. Firstly, we decided whether the genes associated with the suspected risky SNPs are associated with osteoporosis by using random walk algorithm on the PPI network of osteoporosis GWAS-associated genes and the genes associated with the suspected risky SNPs. In order to solve the overfitting problem in ID3 decision tree algorithm, we then classified the SNPs with positive results based on their features of position and function through a simplified classification decision tree which was constructed by ID3 decision tree algorithm with PEP (Pessimistic-Error Pruning). -

BMC Biology Biomed Central
BMC Biology BioMed Central Research article Open Access Classification and nomenclature of all human homeobox genes PeterWHHolland*†1, H Anne F Booth†1 and Elspeth A Bruford2 Address: 1Department of Zoology, University of Oxford, South Parks Road, Oxford, OX1 3PS, UK and 2HUGO Gene Nomenclature Committee, European Bioinformatics Institute (EMBL-EBI), Wellcome Trust Genome Campus, Hinxton, Cambridgeshire, CB10 1SA, UK Email: Peter WH Holland* - [email protected]; H Anne F Booth - [email protected]; Elspeth A Bruford - [email protected] * Corresponding author †Equal contributors Published: 26 October 2007 Received: 30 March 2007 Accepted: 26 October 2007 BMC Biology 2007, 5:47 doi:10.1186/1741-7007-5-47 This article is available from: http://www.biomedcentral.com/1741-7007/5/47 © 2007 Holland et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: The homeobox genes are a large and diverse group of genes, many of which play important roles in the embryonic development of animals. Increasingly, homeobox genes are being compared between genomes in an attempt to understand the evolution of animal development. Despite their importance, the full diversity of human homeobox genes has not previously been described. Results: We have identified all homeobox genes and pseudogenes in the euchromatic regions of the human genome, finding many unannotated, incorrectly annotated, unnamed, misnamed or misclassified genes and pseudogenes. -

Genetic Interplay Between Transcription Factor Pou4f1/Brn3a and Neurotrophin Receptor Ret in Retinal Ganglion Cell Type Specification
bioRxiv preprint doi: https://doi.org/10.1101/2020.03.23.004242; this version posted April 1, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. This article is a US Government work. It is not subject to copyright under 17 USC 105 and is also made available for use under a CC0 license. Genetic Interplay Between Transcription Factor Pou4f1/Brn3a and Neurotrophin Receptor Ret In Retinal Ganglion Cell Type Specification Vladimir Vladimirovich Muzyka1,@, Tudor Constantin Badea1,@ 1: Retinal Circuit Development & Genetics Unit, Neurobiology-Neurodegeneration & Repair Laboratory, National Eye Institute, NIH, Bethesda, MD @: corresponding author: Tudor C. Badea / Vladimir V. Muzyka Retinal Circuit Development & Genetics Unit Building 6, Room 331 6 Center Drive Bethesda, MD 20892-0610 301-496-3978 [email protected] or [email protected] / [email protected] Figures 9 Tables 2 Suplementary Figures 2 Supplementary Tables 2 bioRxiv preprint doi: https://doi.org/10.1101/2020.03.23.004242; this version posted April 1, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. This article is a US Government work. It is not subject to copyright under 17 USC 105 and is also made available for use under a CC0 license. Abstract While the transcriptional code governing retinal ganglion cell (RGC) type specification begins to be understood, its interplay with neurotrophic signaling is largely unexplored. Using sparse random recombination, we show that mosaic gene dosage manipulation of the transcription factor Brn3a/Pou4f1 in neurotrophic receptor Ret heterozygote RGCs results in altered cell fate decisions and/or morphological dendritic defects. -

Development in the Mammalian Auditory System Depends on Transcription Factors
International Journal of Molecular Sciences Review Development in the Mammalian Auditory System Depends on Transcription Factors Karen L. Elliott 1, Gabriela Pavlínková 2 , Victor V. Chizhikov 3, Ebenezer N. Yamoah 4 and Bernd Fritzsch 1,* 1 Department of Biology, University of Iowa, Iowa City, IA 52242, USA; [email protected] 2 Institute of Biotechnology of the Czech Academy of Sciences, 25250 Vestec, Czechia; [email protected] 3 Department of Anatomy and Neurobiology, The University of Tennessee Health Science Center, Memphis, TN 38163, USA; [email protected] 4 Department of Physiology and Cell Biology, School of Medicine, University of Nevada, Reno, NV 89557, USA; [email protected] * Correspondence: [email protected] Abstract: We review the molecular basis of several transcription factors (Eya1, Sox2), including the three related genes coding basic helix–loop–helix (bHLH; see abbreviations) proteins (Neurog1, Neurod1, Atoh1) during the development of spiral ganglia, cochlear nuclei, and cochlear hair cells. Neuronal development requires Neurog1, followed by its downstream target Neurod1, to cross- regulate Atoh1 expression. In contrast, hair cells and cochlear nuclei critically depend on Atoh1 and require Neurod1 expression for interactions with Atoh1. Upregulation of Atoh1 following Neurod1 loss changes some vestibular neurons’ fate into “hair cells”, highlighting the significant interplay between the bHLH genes. Further work showed that replacing Atoh1 by Neurog1 rescues some hair cells from complete absence observed in Atoh1 null mutants, suggesting that bHLH genes can partially replace Citation: Elliott, K.L.; Pavlínková, G.; one another. The inhibition of Atoh1 by Neurod1 is essential for proper neuronal cell fate, and in Chizhikov, V.V.; Yamoah, E.N.; the absence of Neurod1, Atoh1 is upregulated, resulting in the formation of “intraganglionic” HCs.