(Title of the Thesis)*
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

National Parks and Wildlife Act 1972.PDF
Version: 1.7.2015 South Australia National Parks and Wildlife Act 1972 An Act to provide for the establishment and management of reserves for public benefit and enjoyment; to provide for the conservation of wildlife in a natural environment; and for other purposes. Contents Part 1—Preliminary 1 Short title 5 Interpretation Part 2—Administration Division 1—General administrative powers 6 Constitution of Minister as a corporation sole 9 Power of acquisition 10 Research and investigations 11 Wildlife Conservation Fund 12 Delegation 13 Information to be included in annual report 14 Minister not to administer this Act Division 2—The Parks and Wilderness Council 15 Establishment and membership of Council 16 Terms and conditions of membership 17 Remuneration 18 Vacancies or defects in appointment of members 19 Direction and control of Minister 19A Proceedings of Council 19B Conflict of interest under Public Sector (Honesty and Accountability) Act 19C Functions of Council 19D Annual report Division 3—Appointment and powers of wardens 20 Appointment of wardens 21 Assistance to warden 22 Powers of wardens 23 Forfeiture 24 Hindering of wardens etc 24A Offences by wardens etc 25 Power of arrest 26 False representation [3.7.2015] This version is not published under the Legislation Revision and Publication Act 2002 1 National Parks and Wildlife Act 1972—1.7.2015 Contents Part 3—Reserves and sanctuaries Division 1—National parks 27 Constitution of national parks by statute 28 Constitution of national parks by proclamation 28A Certain co-managed national -

Discovering Marsupials Snout, and Enlarged Front Legs
Discovered by Mark Harvey, Western Australian Museum ustralia, New Guinea and some They delved deeper into nearby islands are home to the the status of the quenda, which Avast majority of living marsupials, is endemic to south-western a group of mammals that separated from Australia. It has usually been other mammals in the Mesozoic, about considered as a subspecies 160 million years ago. Elsewhere, marsupials known as Isoodon obesulus are only known from the Americas, although fusciventer, with the other fossils have been found in Asia and Europe. subspecies found in south- Many Australian marsupials have become eastern Australia. However, iconic and are much-loved symbols of Kenny and Matthew raised our nation. Sadly, many are imperilled by it to a full species, finding land clearing and threats from introduced consistent differences in predators. the shape of some of the One of the most distinctive (and teeth and some previously perhaps cute) marsupial groups comprises published molecular data. the bandicoots and bilbies which This species is now known as are classified in their own order, the Isoodon fusciventer. Peramelemorphia. They have long ears and They then turned their Discovering marsupials snout, and enlarged front legs. There are attention to the other only about 22 living species of bandicoots genus, Perameles. Previous and they are only found in the Australo- classifications recognised only Papuan region. Their size and relatively a few modern species, including the long- Above A western barred bandicoot. placid nature has made them vulnerable nosed bandicoot (P. nasuta) from eastern Photo – Jiri Lochman to land-clearing and introduced predators Australia, the eastern barred bandicoot such as feral cats and foxes, and many (P. -

Decline and Extinction of Australian Mammals Since European Settlement
Ongoing unraveling of a continental fauna: Decline FEATURE ARTICLE and extinction of Australian mammals since European settlement John C. Z. Woinarskia,b,1, Andrew A. Burbidgec, and Peter L. Harrisond aNorthern Australian Hub of National Environmental Research Program and bThreatened Species Recovery Hub of National Environmental Science Program, SEE COMMENTARY Charles Darwin University, Darwin, NT 0909, Australia; cResearch Fellow, Department of Parks and Wildlife, Wanneroo, WA 6069, Australia; and dMarine Ecology Research Centre, School of Environment, Science and Engineering, Southern Cross University, Lismore, NSW 2480, Australia This Feature Article is part of a series identified by the Editorial Board as reporting findings of exceptional significance. Edited by William J. Bond, University of Cape Town, Cape Town, South Africa, and approved January 13, 2015 (received for review September 10, 2014) The highly distinctive and mostly endemic Australian land mam- than previously recognized and that many surviving Australian mal fauna has suffered an extraordinary rate of extinction (>10% native mammal species are in rapid decline, notwithstanding the of the 273 endemic terrestrial species) over the last ∼200 y: in generally low level in Australia of most of the threats that are comparison, only one native land mammal from continental North typically driving biodiversity decline elsewhere in the world. America became extinct since European settlement. A further 21% of Australian endemic land mammal species are now assessed to Earlier Losses be threatened, indicating that the rate of loss (of one to two European settlement at 1788 marks a particularly profound extinctions per decade) is likely to continue. Australia’s marine historical landmark for the Australian environment, the opening mammals have fared better overall, but status assessment for up of the continent to a diverse array of new factors, and an ap- them is seriously impeded by lack of information. -

How to Cite Complete Issue More Information About This Article
Therya ISSN: 2007-3364 Centro de Investigaciones Biológicas del Noroeste Woinarski, John C. Z.; Burbidge, Andrew A.; Harrison, Peter L. A review of the conservation status of Australian mammals Therya, vol. 6, no. 1, January-April, 2015, pp. 155-166 Centro de Investigaciones Biológicas del Noroeste DOI: 10.12933/therya-15-237 Available in: http://www.redalyc.org/articulo.oa?id=402336276010 How to cite Complete issue Scientific Information System Redalyc More information about this article Network of Scientific Journals from Latin America and the Caribbean, Spain and Portugal Journal's homepage in redalyc.org Project academic non-profit, developed under the open access initiative THERYA, 2015, Vol. 6 (1): 155-166 DOI: 10.12933/therya-15-237, ISSN 2007-3364 Una revisión del estado de conservación de los mamíferos australianos A review of the conservation status of Australian mammals John C. Z. Woinarski1*, Andrew A. Burbidge2, and Peter L. Harrison3 1National Environmental Research Program North Australia and Threatened Species Recovery Hub of the National Environmental Science Programme, Charles Darwin University, NT 0909. Australia. E-mail: [email protected] (JCZW) 2Western Australian Wildlife Research Centre, Department of Parks and Wildlife, PO Box 51, Wanneroo, WA 6946, Australia. E-mail: [email protected] (AAB) 3Marine Ecology Research Centre, School of Environment, Science and Engineering, Southern Cross University, PO Box 157, Lismore, NSW 2480, Australia. E-mail: [email protected] (PLH) *Corresponding author Introduction: This paper provides a summary of results from a recent comprehensive review of the conservation status of all Australian land and marine mammal species and subspecies. -
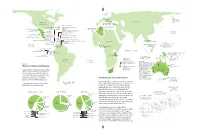
New World Extinctions: Where Next? Mammals: the Recently Departed
GERMANY BERING SEA Bavarian Commander I. pine vole 1962 EURASIA Stellerís sea cow NORTH 1768 CORSICA & SARDINIA AMERICA Sardinian pika Late 18th C. WEST INDIES CUBA MEDITERRANEAN Cuban spiny rats (2 species)* HAITI/ DOMINICAN REPUBLIC ALGERIA Hutia (6 species)* San Pedro Nolasco I. Cuban island shrews (4 species)* CANARY ISLANDS Red gazelle Arredondoís solenodon* Quemi* Volcano mouse* Late 19th c. Pembertonís Hispaniolan spiny rats (2 species)* deer mouse 1931 CAYMAN IS. Hispaniolan island shrews (3 species)* PACIFIC Cayman island shrews Marcanoís solenodon* OCEAN (2 species)* Maria Madre I. PUERTO RICO Nelsonís rice rat 1897 Cayman coneys (two species)* Puerto Rican spiny rats (2 species)* Cayman hutia* MEXICO Omilteme cottontail 1991 BARBUDA Barbuda muskrat* CARIBBEAN Little Swan I. Caribbean MARTINIQUE Martinique muskrat 1902 Little Swan Island coney 1950s-60s PHILIPPINES: Ilin I. Monk seal BARBADOS Barbados rice rat 1847-1890 AFRICA J AMAICA 1950s Small Ilin cloud rat 1953 Jamaican rice rat 1877 ST. VINCENT GUAM Jamaican monkey* St. Vincent rice rat 1890s Guam flying fox 1967 PACIFIC ST. LUCIA PHILIPPINES: Negros I. GHANA Philippine bare- OCEAN St. Lucia muskrat Groove-toothed forest CAROLINES: Palau I. pre-1881 backed fruit bat Palau flying fox 1874 mouse 1890 1975 GAL¡PAGOS ISLANDS San Salvador I. INDIAN OCEAN San Salvador rice rat 1965 Santa Cruz I. Curioís large rice rat* SOLOMON ISLANDS: Guadalcanal Isabela I. Gal·pagos rice rat Little pig rat 1887 Large rice rat* 1930s TANZANIA Giant naked-tailed rat 1960s Small rice rats (2 species)* Painted bat 1878 NEW GUINEA CHRISTMAS I. Large-eared Maclearís rat 1908 nyctophilus (bat) 1890 MADAGASCAR Bulldog rat 1908 Santa Cruz I. -

Checklist of the Mammals of Western Australia
Records ofthe Western Australian Museum Supplement No. 63: 91-98 (2001). Checklist of the mammals of Western Australia R.A. How, N.K. Cooper and J.L. Bannister Western Australian Museum, Francis Street, Perth, Western Australia 6000, Australia INTRODUCTION continued collection of species across their range. The Checklist ofthe Mammals ofWestern Australia is Where the level of taxonomic uncertainty is being a collation of the most recent systematic information formally resolved, footnotes to the individual taxon on Western Australian mammal taxa, incorporating appear at the end of the family listings. the list of taxa compiled from the Western Numerous taxa have become extinct on a national Australian Museum's mammal database and the or state level since European settlement and there literature. The Checklist presents the nomenclature have been several recent attempts to reintroduce accepted by the Western Australian Museum in regionally extinct taxa to former areas. The present maintaining the state's mammal collection and status of these taxa is indicated by symbols in the database. Listed are those species probably extant Checklist. at the time of arrival of Europeans to Western Australia. Symbols used Nomenclature, in general, follows the Zoological t Denotes extinct taxon. Catalogue ofAustralia, Volume 5, Mammalia (1988). * Denotes taxon extinct in Western Australia but Consideration has been given to the nomenclatural extant in other parts of Australia. decisions in The 1996 Action Plan of Australian $ Denotes taxon extinct on Western Australian Marsupials and Monotremes (Maxwell, Burbidge and mainland and recently reintroduced from other Morris, 1996) and The Action Plan for Australian Bats parts of Australia or translocated from islands (Reardon, 1999a). -

Chapter 1: Introduction 1
Chapter 1: Introduction 1 Chapter 1 Introduction 1.1 General introduction Forty-two species of mammals worldwide have suffered extinction since 1600 (IUCN 2003). Almost half of the world’s mammal extinctions in the last two hundred years have occurred in Australia, coinciding with European settlement of the continent (Short and Smith 1994). Seventeen species are extinct, 10 survive only as island populations, and a further 17 have been reduced to remnant populations of less than 10% of their range prior to European settlement. The decline and extinction of these species has been documented by a number of early authors (e.g. Finlayson 1935, 1961; Jones 1923-25; Krefft 1866; Shortridge 1909), and has typically been associated with the impact of European settlement of Australia. Soulé (1983, p. 112) noted: "As far as I know, no biologist has documented the extinction of a continental species of plant or animal caused solely by non- human agencies such as competition, disease or environmental perturbation in situations unaffected by man". Possible causes for the losses in our native mammal fauna have been widely debated over the last century, yet the causal factors for the decline and extinction of individual species are rarely known (Caughley and Gunn 1996). In many cases, the biology of a species has been so poorly known, that any threatening processes identified have been rarely more than a guess. Predation by introduced feral cats Felis catus and the European fox Vulpes vulpes, habitat modification and fragmentation (destruction by clearing for agriculture, the impacts of introduced livestock and European rabbits Oryctolagus cuniculus, changes in fire regimes), competition from introduced species, hunting pressure, pest control, pollution, and disease, or a combination of any of these factors, have commonly been implicated in the loss of mammal species in Australia (e.g. -

© Cambridge University Press Cambridge
Cambridge University Press 978-0-521-86620-0 - Continent of Curiosities: A Journey Through Australian Natural History Danielle Clode Index More information INDEX Page numbers in italics refer to illustrations, Bare-Backed Fruit Bat (Dobsonia moluccensis), plates and their accompanying captions. 92 Baret, Jeanne, 31 A barking spider (Selenocosima crassipes), 38 Acacia (Wattles), 66, 67 Bass, George, 30 Forest Wattle (Acacia frigiscens), 70 Batavia (ship), 18 Mountain Hickory Wattle (Acacia Baudin expedition, 89–92 obliquinervia), 70 Bennett, George, 14 Silver Wattle (Acacia dealbata), 70 biogeography, 137–8, 154–8 Acheron River, 61 dispersal, 154 adaptation vicariance, 156 of behaviour, 102 Wallace develops, 138–46 to cold, 110 Bird Rock Cliffs (Victoria), 95 to fire, 67 birds, classification of, 143–4 Agassiz, Louis, 94–5 Australian Bustard (Ardeotis australis), 34 American Mink (Mustela vison), 102, 102 Brolga (Grus rubicundus), 34 Antilopine Wallaroo (Macropus antilopinus), Cassowary (Casuarius spp.), facing 148 facing 36 Chestnut Quail-Thrush (Cinclosoma Antarctic Icefish (Champocephalus gunnari), 110 castanotus), facing 133 apes Great Pampa-Finch (Embernagra platensis), classification, 123–6 3,5 Museum Victoria display, 115, 124, 125 Mrs Gould’s Sunbird (Aethopyga gouldiae), archaebacteria, 164–5, see also bacteria facing 36 Asian dragon (Calotes spp.), 105 Pied Goose (Anseranas semipalmata), 34 Astacopsidrilus, 158 Piopio (Turnagra capensis), 155 atlases, see maps Powerful Owl (Ninox strenua), 61 Austral Grass-tree (Xanthorrhoea australis), -

Finding out Bout Desert Mammals by Andrew Burbidge and Phillip Uller Western Australian Wildlife Research Centre
Finding out bout Desert Mammals by Andrew Burbidge and Phillip uller Western Australian Wildlife Research Centre & Phil Fuller and Aboriginal informant Mi cky Robinson at Tjirrkali, Gibson Desert. Mr Fuller is holding a Oalgyte skin. Photo: R. Southgate Until quite recently surprisingly little was known about the Research Centre started vertebrate mammals of the interior of Western Australia. Because of the surveys in Western Australia and the inaccessibility of the region there were few systematic attempts to Northern Territory Government document the fauna of central Australia and by the time four-wheel appointed biologists to study drive vehicles became available it appeared that many species had endangered species there. either become extremely rare or were extinct. It soon became apparent that many That we knew anything about the the region, obtaining much of the species which Finlayson found mammals of the deserts in past times information and help from them. to be abundant and widespread were was due largely to the efforts of one Finlayson's studies were concentrated now rare or extinct. The species most person, H . H. Finlayson of Adelaide. in northern South Australia but he affected were those of intermediate Prior to his work our knowledge was also recorded information from size between the large kangaroos and confined to a few scattered records Western Australia and the Northern the small native mice, dunnarts and from explorers and missionaries and Territory. His unequalled knowledge bats. Recent analyses of the decline the only extensive scientific study, of the distribution and status of of mammals in Western Australia by that of the Horn Scientific central Australian mammals was the authors and N. -

Biological Survey Anangu Pitjantjatjara Lands
VEGETATION MAPPING By S. J. Pillman1, P. D. Canty1, A. C. Robinson1, and P. J. Lang1 INTRODUCTION plains, Eucalyptus microtheca [= coolabah] Low Only very generalised vegetation mapping was Woodland on floodplains, Eucalyptus gamophylla +/ available for the Anangu Pitjantjatjara Lands (AP E. oxymitra Open Scrub on dunes and Atriplex Lands) prior to this survey. rhagodioides [= nummularia] Low Shrubland on plains. Specht (1972) mapped four general communities: EPHEMERAL COMMUNITIES, Far North-west In his update of the work of Davies, Neagle (1995) Ranges, TALL SHRUBLAND, Acacia aneura- A. added data on additions to the National Parks and brachystachya, HUMMOCK GRASSLAND to LOW Wildlife Conservation Reserves system and on the OPEN WOODLAND Triodia basedowii (+/ many Heritage Agreement Areas proclaimed since Casuarina [= Allocasuarina] decaisneana in north, Davies’ work, but this had no effect in the AP Lands in LOW SHRUBLAND, Atriplex rhagodioides [= terms of re-defining the threatened vegetation nummularia] and LOW SHRUBLAND Atriplex communities. vesicaria- Bassia [Sclerolaena] sp. Broad-scale vegetation mapping has also been A map produced by the Pastoral Board (1973) showed completed in those areas of Western Australia and the four general communities: MALLEE - NATIVE PINE Northern Territory, which adjoin the AP Lands. - NATIVE FIG – CORKWOOD – BLOODWOOD SPINIFEX ASSOCIATION on the ranges, DESERT In Western Australia, Beard and Webb (1974) OAK - DESERT KURRAJONG – MULGA described the techniques used in the 1:250 000 SPINIFEX ASSOCIATION and a MULGA – vegetation mapping to develop the 1:1 000 000 CORKWOOD - BLOODWOOD- IRONWOOD vegetation map series for Western Australia. The ASSOCIATION on the surrounding plains. second of these published maps covered the Great Victoria Desert (Beard 1974) and included the area The map by Boomsma and Lewis (1980) incorporating adjacent to the AP Lands in Western Australia. -

Tackling Feral Cats and Their Impacts
What is a feral cat? ................................................................................................................... 2 Why are feral cats such a big problem? ................................................................................. 2 What is being done to reduce feral cats? ............................................................................... 3 How many feral cats are there in Australia? ........................................................................... 3 How was the two million feral cats culled target determined? .............................................. 4 What do you mean by humane, effective and justifiable? ..................................................... 4 How does Eradicat® and 1080 work? ..................................................................................... 4 How does Curiosity® work? Is it humane? ............................................................................ 5 Are these feral cat baits poisoning the environment? ........................................................... 6 Is feral cat control a threat to my pet cat? .............................................................................. 6 Why don’t You make desexing of all cats mandatory? .......................................................... 6 Habitat loss is the most important driver of extinction, why are you focusing on feral cats? .......................................................................................................................................... 7 If feral -

Invasive Species – a Leading Threat to Australia's Wildlife
2017 INVASIVE FEBRUARY SPECIES A leading threat to Australia’s wildlife INVASIVE SPECIES A leading threat to Australia’s wildlife Report author: Tim Low © 2017 Invasive Species Council Report design: John Sampson, Ecotype. Cover photos: Feral cat graphic based on photo by Mick Davis, Invasive Animals CRC; cassowary chick, Emmagen Creek (Dan Gordon - Flickr | CC BY-NC 2.0). Contact Andrew Cox, CEO Email: [email protected] Phone: 0438 588 040 All material in this publication is licensed under a Creative Commons Attribution 3.0 Australia licence, save for the Invasive Species Council logo and third party content. You are free to use this publication in accordance with the licence terms, attributing the Invasive Species Council and keeping intact the original copyright notice. 2 Invasive Species: A leading threat to Australia’s wildlife INVASIVE Introduction Report author Tim Low SPECIES ustralia has an invidious reputation as Habitat loss is often assumed to be the main a place of extinction, having lost more threatening process in Australia, but the evidence mammal species than any other country. indicates that invasive species have caused the A leading threat to Australia’s wildlife AFoxes and feral cats are blamed for most of the most animal extinctions, and pose the main losses. Australia’s smaller mammals proved very threat to some animal groups. The evidence vulnerable after evolving in isolation on a continent for this comes from a number of sources and is lacking carnivores that operate like these. The same summarised here, firstly as it applies to threatened isolation has left Australia’s wildlife susceptible to species (drawing upon three studies), secondly to any number of introduced organisms, including threatened ecological communities, and thirdly to cane toads with their unusual toxins, trout released extinct animals.