Copula Regression Models for the Analysis of Correlated Data with Missing Values
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Estimation of Common Location and Scale Parameters in Nonregular Cases Ahmad Razmpour Iowa State University
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 1982 Estimation of common location and scale parameters in nonregular cases Ahmad Razmpour Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Statistics and Probability Commons Recommended Citation Razmpour, Ahmad, "Estimation of common location and scale parameters in nonregular cases " (1982). Retrospective Theses and Dissertations. 7528. https://lib.dr.iastate.edu/rtd/7528 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. INFORMATION TO USERS This reproduction was made from a copy of a document sent to us for microfilming. While the most advanced technology has been used to photograph and reproduce this document, the quality of the reproduction is heavily dependent upon the quality of the material submitted. The following explanation of techniques is provided to help clarify markings or notations which may appear on this reproduction. 1. The sign or "target" for pages apparently lacking from the document photographed is "Missing Page(s)". If it was possible to obtain the missing page(s) or section, they are spliced into the film along with adjacent pages. This may have necessitated cutting through an image and duplicating adjacent pages to assure complete continuity. 2. When an image on the film is obliterated with a round black mark, it is an indication of either blurred copy because of movement during exposure, duplicate copy, or copyrighted materials that should not have been filmed. -

A Comparison of Unbiased and Plottingposition Estimators of L
WATER RESOURCES RESEARCH, VOL. 31, NO. 8, PAGES 2019-2025, AUGUST 1995 A comparison of unbiased and plotting-position estimators of L moments J. R. M. Hosking and J. R. Wallis IBM ResearchDivision, T. J. Watson ResearchCenter, Yorktown Heights, New York Abstract. Plotting-positionestimators of L momentsand L moment ratios have several disadvantagescompared with the "unbiased"estimators. For generaluse, the "unbiased'? estimatorsshould be preferred. Plotting-positionestimators may still be usefulfor estimatingextreme upper tail quantilesin regional frequencyanalysis. Probability-Weighted Moments and L Moments •r+l-" (--1)r • P*r,k Olk '- E p *r,!•[J!•. Probability-weightedmoments of a randomvariable X with k=0 k=0 cumulativedistribution function F( ) and quantile function It is convenient to define dimensionless versions of L mo- x( ) were definedby Greenwoodet al. [1979]to be the quan- tities ments;this is achievedby dividingthe higher-orderL moments by the scale measure h2. The L moment ratios •'r, r = 3, Mp,ra= E[XP{F(X)}r{1- F(X)} s] 4, '", are definedby ßr-" •r/•2 ß {X(u)}PUr(1 -- U)s du. L momentratios measure the shapeof a distributionindepen- dently of its scaleof measurement.The ratios *3 ("L skew- ness")and *4 ("L kurtosis")are nowwidely used as measures Particularlyuseful specialcases are the probability-weighted of skewnessand kurtosis,respectively [e.g., Schaefer,1990; moments Pilon and Adamowski,1992; Royston,1992; Stedingeret al., 1992; Vogeland Fennessey,1993]. 12•r= M1,0, r = •01 (1 - u)rx(u) du, Estimators Given an ordered sample of size n, Xl: n • X2:n • ''' • urx(u) du. X.... there are two establishedways of estimatingthe proba- /3r--- Ml,r, 0 =f01 bility-weightedmoments and L moments of the distribution from whichthe samplewas drawn. -

The Asymmetric T-Copula with Individual Degrees of Freedom
The Asymmetric t-Copula with Individual Degrees of Freedom d-fine GmbH Christ Church University of Oxford A thesis submitted for the degree of MSc in Mathematical Finance Michaelmas 2012 Abstract This thesis investigates asymmetric dependence structures of multivariate asset returns. Evidence of such asymmetry for equity returns has been reported in the literature. In order to model the dependence structure, a new t-copula approach is proposed called the skewed t-copula with individ- ual degrees of freedom (SID t-copula). This copula provides the flexibility to assign an individual degree-of-freedom parameter and an individual skewness parameter to each asset in a multivariate setting. Applying this approach to GARCH residuals of bivariate equity index return data and using maximum likelihood estimation, we find significant asymmetry. By means of the Akaike information criterion, it is demonstrated that the SID t-copula provides the best model for the market data compared to other copula approaches without explicit asymmetry parameters. In addition, it yields a better fit than the conventional skewed t-copula with a single degree-of-freedom parameter. In a model impact study, we analyse the errors which can occur when mod- elling asymmetric multivariate SID-t returns with the symmetric multi- variate Gauss or standard t-distribution. The comparison is done in terms of the risk measures value-at-risk and expected shortfall. We find large deviations between the modelled and the true VaR/ES of a spread posi- tion composed of asymmetrically distributed risk factors. Going from the bivariate case to a larger number of risk factors, the model errors increase. -

A Bayesian Hierarchical Spatial Copula Model: an Application to Extreme Temperatures in Extremadura (Spain)
atmosphere Article A Bayesian Hierarchical Spatial Copula Model: An Application to Extreme Temperatures in Extremadura (Spain) J. Agustín García 1,† , Mario M. Pizarro 2,*,† , F. Javier Acero 1,† and M. Isabel Parra 2,† 1 Departamento de Física, Universidad de Extremadura, Avenida de Elvas, 06006 Badajoz, Spain; [email protected] (J.A.G.); [email protected] (F.J.A.) 2 Departamento de Matemáticas, Universidad de Extremadura, Avenida de Elvas, 06006 Badajoz, Spain; [email protected] * Correspondence: [email protected] † These authors contributed equally to this work. Abstract: A Bayesian hierarchical framework with a Gaussian copula and a generalized extreme value (GEV) marginal distribution is proposed for the description of spatial dependencies in data. This spatial copula model was applied to extreme summer temperatures over the Extremadura Region, in the southwest of Spain, during the period 1980–2015, and compared with the spatial noncopula model. The Bayesian hierarchical model was implemented with a Monte Carlo Markov Chain (MCMC) method that allows the distribution of the model’s parameters to be estimated. The results show the GEV distribution’s shape parameter to take constant negative values, the location parameter to be altitude dependent, and the scale parameter values to be concentrated around the same value throughout the region. Further, the spatial copula model chosen presents lower deviance information criterion (DIC) values when spatial distributions are assumed for the GEV distribution’s Citation: García, J.A.; Pizarro, M.M.; location and scale parameters than when the scale parameter is taken to be constant over the region. Acero, F.J.; Parra, M.I. A Bayesian Hierarchical Spatial Copula Model: Keywords: Bayesian hierarchical model; extreme temperature; Gaussian copula; generalized extreme An Application to Extreme value distribution Temperatures in Extremadura (Spain). -

Procedures for Estimation of Weibull Parameters James W
United States Department of Agriculture Procedures for Estimation of Weibull Parameters James W. Evans David E. Kretschmann David W. Green Forest Forest Products General Technical Report February Service Laboratory FPL–GTR–264 2019 Abstract Contents The primary purpose of this publication is to provide an 1 Introduction .................................................................. 1 overview of the information in the statistical literature on 2 Background .................................................................. 1 the different methods developed for fitting a Weibull distribution to an uncensored set of data and on any 3 Estimation Procedures .................................................. 1 comparisons between methods that have been studied in the 4 Historical Comparisons of Individual statistics literature. This should help the person using a Estimator Types ........................................................ 8 Weibull distribution to represent a data set realize some advantages and disadvantages of some basic methods. It 5 Other Methods of Estimating Parameters of should also help both in evaluating other studies using the Weibull Distribution .......................................... 11 different methods of Weibull parameter estimation and in 6 Discussion .................................................................. 12 discussions on American Society for Testing and Materials Standard D5457, which appears to allow a choice for the 7 Conclusion ................................................................ -
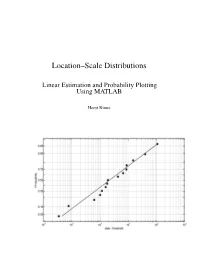
Location-Scale Distributions
Location–Scale Distributions Linear Estimation and Probability Plotting Using MATLAB Horst Rinne Copyright: Prof. em. Dr. Horst Rinne Department of Economics and Management Science Justus–Liebig–University, Giessen, Germany Contents Preface VII List of Figures IX List of Tables XII 1 The family of location–scale distributions 1 1.1 Properties of location–scale distributions . 1 1.2 Genuine location–scale distributions — A short listing . 5 1.3 Distributions transformable to location–scale type . 11 2 Order statistics 18 2.1 Distributional concepts . 18 2.2 Moments of order statistics . 21 2.2.1 Definitions and basic formulas . 21 2.2.2 Identities, recurrence relations and approximations . 26 2.3 Functions of order statistics . 32 3 Statistical graphics 36 3.1 Some historical remarks . 36 3.2 The role of graphical methods in statistics . 38 3.2.1 Graphical versus numerical techniques . 38 3.2.2 Manipulation with graphs and graphical perception . 39 3.2.3 Graphical displays in statistics . 41 3.3 Distribution assessment by graphs . 43 3.3.1 PP–plots and QQ–plots . 43 3.3.2 Probability paper and plotting positions . 47 3.3.3 Hazard plot . 54 3.3.4 TTT–plot . 56 4 Linear estimation — Theory and methods 59 4.1 Types of sampling data . 59 IV Contents 4.2 Estimators based on moments of order statistics . 63 4.2.1 GLS estimators . 64 4.2.1.1 GLS for a general location–scale distribution . 65 4.2.1.2 GLS for a symmetric location–scale distribution . 71 4.2.1.3 GLS and censored samples . -

Lecture 22 - 11/19/2019 Lecture 22: Robust Location Estimation Lecturer: Jiantao Jiao Scribe: Vignesh Subramanian
EE290 Mathematics of Data Science Lecture 22 - 11/19/2019 Lecture 22: Robust Location Estimation Lecturer: Jiantao Jiao Scribe: Vignesh Subramanian In this lecture, we get a historical perspective into the robust estimation problem and discuss Huber's work [1] for robust estimation of a location parameter. The Huber loss function is given by, ( 1 2 2 t ; jtj ≤ k ρHuber(t) = 1 2 : (1) k jtj − 2 k ; jtj > k Here k is a parameter and the idea behind the loss function is to penalize outliers (beyond k) linearly instead of quadratically. Figure 1 shows the Huber loss function for k = 1. In this lecture we will get an intuitive 1 2 Figure 1: The green line plots the Huber-loss function for k = 1, and the blue line plots the quadratic function 2 t . understanding for the reasons behind the particular form of this function, quadratic in interior, linear in exterior and convex and will see that this loss function is optimal for one dimensional robust mean estimation for Gaussian location model. First we describe the problem setting. 1 Problem Setting Suppose we observe X1;X2;:::;Xn where Xi − µ ∼ F 2 F are i.i.d. Here, F = fF j F = (1 − )G + H; H 2 Mg; (2) where G 2 M is some fixed distribution function which is usually assumed to have zero mean, and M denotes the space of all probability measures. This describes the corruption model where the observed distribution is a convex combination of the true distribution G(x) and an arbitrary corruption distribution H. -

Robust Estimator of Location Parameter1)
The Korean Communications in Statistics Vol. 11 No. 1, 2004 pp. 153-160 Robust Estimator of Location Parameter1) Dongryeon Park2) Abstract In recent years, the size of data set which we usually handle is enormous, so a lot of outliers could be included in data set. Therefore the robust procedures that automatically handle outliers become very importance issue. We consider the robust estimation problem of location parameter in the univariate case. In this paper, we propose a new method for defining robustness weights for the weighted mean based on the median distance of observations and compare its performance with several existing robust estimators by a simulation study. It turns out that the proposed method is very competitive. Keywords : Location parameter, Median distance, Robust estimator, Robustness weight 1. Introduction It is often assumed in the social sciences that data conform to a normal distribution. When estimating the location of a normal distribution, a sample mean is well known to be the best estimator according to many criteria. However, numerous studies (Hample et al., 1986; Hoaglin et al., 1976; Rousseeuw and Leroy, 1987) have strongly questioned normal assumption in real world data sets. In fact, a few large errors might infect the data set so that the tails of the underlying distribution are heavier than those of the normal distribution. In this situation, the sample mean is no longer a good estimate for the center of symmetry because all the observations equally contribute to the value of the sample mean, so the estimators which are insensitive to extreme values should have better performance. -

Maximum Likelihood Characterization of Distributions
Bernoulli 20(2), 2014, 775–802 DOI: 10.3150/13-BEJ506 Maximum likelihood characterization of distributions MITIA DUERINCKX1,*, CHRISTOPHE LEY1,** and YVIK SWAN2 1D´epartement de Math´ematique, Universit´elibre de Bruxelles, Campus Plaine CP 210, Boule- vard du triomphe, B-1050 Bruxelles, Belgique. * ** E-mail: [email protected]; [email protected] 2 Universit´edu Luxembourg, Campus Kirchberg, UR en math´ematiques, 6, rue Richard Couden- hove-Kalergi, L-1359 Luxembourg, Grand-Duch´ede Luxembourg. E-mail: [email protected] A famous characterization theorem due to C.F. Gauss states that the maximum likelihood es- timator (MLE) of the parameter in a location family is the sample mean for all samples of all sample sizes if and only if the family is Gaussian. There exist many extensions of this re- sult in diverse directions, most of them focussing on location and scale families. In this paper, we propose a unified treatment of this literature by providing general MLE characterization theorems for one-parameter group families (with particular attention on location and scale pa- rameters). In doing so, we provide tools for determining whether or not a given such family is MLE-characterizable, and, in case it is, we define the fundamental concept of minimal necessary sample size at which a given characterization holds. Many of the cornerstone references on this topic are retrieved and discussed in the light of our findings, and several new characterization theorems are provided. Of particular interest is that one part of our work, namely the intro- duction of so-called equivalence classes for MLE characterizations, is a modernized version of Daniel Bernoulli’s viewpoint on maximum likelihood estimation. -

Parametric Forecasting of Value at Risk Using Heavy-Tailed Distribution
REVISTA INVESTIGACIÓN OPERACIONAL _____ Vol., 30, No.1, 32-39, 2009 DEPENDENCE BETWEEN VOLATILITY PERSISTENCE, KURTOSIS AND DEGREES OF FREEDOM Ante Rozga1 and Josip Arnerić, 2 Faculty of Economics, University of Split, Croatia ABSTRACT In this paper the dependence between volatility persistence, kurtosis and degrees of freedom from Student’s t-distribution will be presented in estimation alternative risk measures on simulated returns. As the most used measure of market risk is standard deviation of returns, i.e. volatility. However, based on volatility alternative risk measures can be estimated, for example Value-at-Risk (VaR). There are many methodologies for calculating VaR, but for simplicity they can be classified into parametric and nonparametric models. In category of parametric models the GARCH(p,q) model is used for modeling time-varying variance of returns. KEY WORDS: Value-at-Risk, GARCH(p,q), T-Student MSC 91B28 RESUMEN En este trabajo la dependencia de la persistencia de la volatilidad, kurtosis y grados de liberad de una Distribución T-Student será presentada como una alternativa para la estimación de medidas de riesgo en la simulación de los retornos. La medida más usada de riesgo de mercado es la desviación estándar de los retornos, i.e. volatilidad. Sin embargo, medidas alternativas de la volatilidad pueden ser estimadas, por ejemplo el Valor-al-Riesgo (Value-at-Risk, VaR). Existen muchas metodologías para calcular VaR, pero por simplicidad estas pueden ser clasificadas en modelos paramétricos y no paramétricos . En la categoría de modelos paramétricos el modelo GARCH(p,q) es usado para modelar la varianza de retornos que varían en el tiempo. -

Sufficient Conditions for Monotone Hazard Rate an Application to Latency-Probability Curves
JOURNAL OF MATHEMATICAL PSYCHOLOGY 8, 303-332 (1971) Sufficient Conditions for Monotone Hazard Rate An Application to Latency-Probability Curves EWART A. C. THOMAS The University of Michigan, Ann Arbor, Michigan 48104 It is assumed that when a subject makes a response after comparing a random variable with a fixed criterion, his response latency is inversely related to the distance between the value of the variable and the criterion. This paper then examines the relationship to response probability of (a) response latency, (b) difference between two latencies and (c) dispersion of latency, and also some properties of the latency distribu- tion. It is shown that the latency-probability curve is decreasing if and only if the hazard function of the underlying distribution is increasing and, by using a fundamental lemma, sufficient conditions are obtained for monotone hazard rate. An inequality is established which is satisfied by the slope of the Receiver Operating Characteristic under these sufficient conditions. Finally, a Latency Operating Charac- teristic is defined and it is suggested that such a plot can be useful in assessing the consistency between latency data and theory. I. INTRODUCTION Many authors have remarked that an analysis of the response times from a signal detection experiment can yield valuable information about the nature of the detection process (e.g. Carterette, 1966; Green and Lute, 1967; Laming, 1969). The method of such an analysis depends on the assumed detection and response time models, and on the response time statistic being observed, though the dependence on the former is more fundamental than that on the latter. -

Functions by Name
Title stata.com Functions by name abbrev(s,n) name s, abbreviated to a length of n abs(x) the absolute value of x acos(x) the radian value of the arccosine of x acosh(x) the inverse hyperbolic cosine of x age(ed DOB,ed ,snl ) the age in integer years on ed for date of birth ed DOB with snl the nonleap-year birthday for 29feb birthdates age frac(ed DOB,ed ,snl ) the age in years, including the fractional part, on ed for date of birth ed DOB with snl the nonleap-year birthday for 29feb birthdates asin(x) the radian value of the arcsine of x asinh(x) the inverse hyperbolic sine of x atan(x) the radian value of the arctangent of x atan2(y, x) the radian value of the arctangent of y=x, where the signs of the parameters y and x are used to determine the quadrant of the answer atanh(x) the inverse hyperbolic tangent of x autocode(x,n,x0,x1) partitions the interval from x0 to x1 into n equal-length intervals and returns the upper bound of the interval that contains x or the upper bound of the first or last interval if x < x0 or x > x1, respectively betaden(a,b,x) the probability density of the beta distribution, where a and b are the shape parameters; 0 if x < 0 or x > 1 binomial(n,k,θ) the probability of observing floor(k) or fewer successes in floor(n) trials when the probability of a success on one trial is θ; 0 if k < 0; or 1 if k > n binomialp(n,k,p) the probability of observing floor(k) successes in floor(n) trials when the probability of a success on one trial is p binomialtail(n,k,θ) the probability of observing floor(k) or more successes