UC Riverside UC Riverside Previously Published Works
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Development and Evaluation of Rrna Targeted in Situ Probes and Phylogenetic Relationships of Freshwater Fungi
Development and evaluation of rRNA targeted in situ probes and phylogenetic relationships of freshwater fungi vorgelegt von Diplom-Biologin Christiane Baschien aus Berlin Von der Fakultät III - Prozesswissenschaften der Technischen Universität Berlin zur Erlangung des akademischen Grades Doktorin der Naturwissenschaften - Dr. rer. nat. - genehmigte Dissertation Promotionsausschuss: Vorsitzender: Prof. Dr. sc. techn. Lutz-Günter Fleischer Berichter: Prof. Dr. rer. nat. Ulrich Szewzyk Berichter: Prof. Dr. rer. nat. Felix Bärlocher Berichter: Dr. habil. Werner Manz Tag der wissenschaftlichen Aussprache: 19.05.2003 Berlin 2003 D83 Table of contents INTRODUCTION ..................................................................................................................................... 1 MATERIAL AND METHODS .................................................................................................................. 8 1. Used organisms ............................................................................................................................. 8 2. Media, culture conditions, maintenance of cultures and harvest procedure.................................. 9 2.1. Culture media........................................................................................................................... 9 2.2. Culture conditions .................................................................................................................. 10 2.3. Maintenance of cultures.........................................................................................................10 -

Fungal Evolution: Major Ecological Adaptations and Evolutionary Transitions
Biol. Rev. (2019), pp. 000–000. 1 doi: 10.1111/brv.12510 Fungal evolution: major ecological adaptations and evolutionary transitions Miguel A. Naranjo-Ortiz1 and Toni Gabaldon´ 1,2,3∗ 1Department of Genomics and Bioinformatics, Centre for Genomic Regulation (CRG), The Barcelona Institute of Science and Technology, Dr. Aiguader 88, Barcelona 08003, Spain 2 Department of Experimental and Health Sciences, Universitat Pompeu Fabra (UPF), 08003 Barcelona, Spain 3ICREA, Pg. Lluís Companys 23, 08010 Barcelona, Spain ABSTRACT Fungi are a highly diverse group of heterotrophic eukaryotes characterized by the absence of phagotrophy and the presence of a chitinous cell wall. While unicellular fungi are far from rare, part of the evolutionary success of the group resides in their ability to grow indefinitely as a cylindrical multinucleated cell (hypha). Armed with these morphological traits and with an extremely high metabolical diversity, fungi have conquered numerous ecological niches and have shaped a whole world of interactions with other living organisms. Herein we survey the main evolutionary and ecological processes that have guided fungal diversity. We will first review the ecology and evolution of the zoosporic lineages and the process of terrestrialization, as one of the major evolutionary transitions in this kingdom. Several plausible scenarios have been proposed for fungal terrestralization and we here propose a new scenario, which considers icy environments as a transitory niche between water and emerged land. We then focus on exploring the main ecological relationships of Fungi with other organisms (other fungi, protozoans, animals and plants), as well as the origin of adaptations to certain specialized ecological niches within the group (lichens, black fungi and yeasts). -

Metabolites from Nematophagous Fungi and Nematicidal Natural Products from Fungi As an Alternative for Biological Control
Appl Microbiol Biotechnol (2016) 100:3799–3812 DOI 10.1007/s00253-015-7233-6 MINI-REVIEW Metabolites from nematophagous fungi and nematicidal natural products from fungi as an alternative for biological control. Part I: metabolites from nematophagous ascomycetes Thomas Degenkolb1 & Andreas Vilcinskas1,2 Received: 4 October 2015 /Revised: 29 November 2015 /Accepted: 2 December 2015 /Published online: 29 December 2015 # The Author(s) 2015. This article is published with open access at Springerlink.com Abstract Plant-parasitic nematodes are estimated to cause Keywords Phytoparasitic nematodes . Nematicides . global annual losses of more than US$ 100 billion. The num- Oligosporon-type antibiotics . Nematophagous fungi . ber of registered nematicides has declined substantially over Secondary metabolites . Biocontrol the last 25 years due to concerns about their non-specific mechanisms of action and hence their potential toxicity and likelihood to cause environmental damage. Environmentally Introduction beneficial and inexpensive alternatives to chemicals, which do not affect vertebrates, crops, and other non-target organisms, Nematodes as economically important crop pests are therefore urgently required. Nematophagous fungi are nat- ural antagonists of nematode parasites, and these offer an eco- Among more than 26,000 known species of nematodes, 8000 physiological source of novel biocontrol strategies. In this first are parasites of vertebrates (Hugot et al. 2001), whereas 4100 section of a two-part review article, we discuss 83 nematicidal are parasites of plants, mostly soil-borne root pathogens and non-nematicidal primary and secondary metabolites (Nicol et al. 2011). Approximately 100 species in this latter found in nematophagous ascomycetes. Some of these sub- group are considered economically important phytoparasites stances exhibit nematicidal activities, namely oligosporon, of crops. -

Pulmonary Mycobiome of Patients with Suspicion of Respiratory Fungal Infection – an Exploratory Study Mariana Oliveira 1,2 , Miguel Pinto 3, C
#138: Pulmonary mycobiome of patients with suspicion of respiratory fungal infection – an exploratory study Mariana Oliveira 1,2 , Miguel Pinto 3, C. Veríssimo 2, R. Sabino 2,* . 1Faculty of Sciences of the University of Lisbon, Portugal; 2Department of Infectious Diseases of the National Institute of Health Doutor Ricardo Jorge, Lisbon, Portugal; 3 Bioinformatics Unit, Infectious Diseases Department, National Institute of Health Dr. Ricardo Jorge, Avenida Padre Cruz, 1600-560 Lisboa, Portugal. ABSTRACT INTRODUCTION This pilot study aimed to characterize the pulmonary mycobiome of patients with suspicion of fungal infection of the The possibility of knowing and comparing the mycobiome of healthy individuals with respiratory tract as well as to identify potentially pathogenic fungi infecting their lungs. the mycobiome of patients with different pathologies, as well as the capacity to quickly and DNA was extracted from the respiratory samples of a cohort of 10 patients with suspicion of respiratory fungal infection. The specifically detect and identify potentially pathogenic fungi present in the pulmonary internal transcribed spacer 1 (ITS1) region and the calmodulin (CMD) gene were amplified by PCR and the resulting amplicons were sequenced through next generation sequencing (NGS) techniques. The DNA sequences obtained were taxonomically mycobiome of patients makes NGS techniques useful for the laboratory diagnosis of identified using the PIPITS and bowtie2 platforms. fungal infections. Thus, the aim of this exploratory study was to optimize the procedure for Twenty-four different OTU (grouped in 17 phylotypes) were considered as part of the pulmonary mycobiome. Twelve genera the detection of fungi through NGS techniques. A metagenomic analysis was performed in of fungi were identified. -

Checklist of the Lichens and Allied Fungi of Kathy Stiles Freeland Bibb County Glades Preserve, Alabama, U.S.A
Opuscula Philolichenum, 18: 420–434. 2019. *pdf effectively published online 2December2019 via (http://sweetgum.nybg.org/philolichenum/) Checklist of the lichens and allied fungi of Kathy Stiles Freeland Bibb County Glades Preserve, Alabama, U.S.A. J. KEVIN ENGLAND1, CURTIS J. HANSEN2, JESSICA L. ALLEN3, SEAN Q. BEECHING4, WILLIAM R. BUCK5, VITALY CHARNY6, JOHN G. GUCCION7, RICHARD C. HARRIS8, MALCOLM HODGES9, NATALIE M. HOWE10, JAMES C. LENDEMER11, R. TROY MCMULLIN12, ERIN A. TRIPP13, DENNIS P. WATERS14 ABSTRACT. – The first checklist of lichenized, lichenicolous and lichen-allied fungi from the Kathy Stiles Freeland Bibb County Glades Preserve in Bibb County, Alabama, is presented. Collections made during the 2017 Tuckerman Workshop and additional records from herbaria and online sources are included. Two hundred and thirty-eight taxa in 115 genera are enumerated. Thirty taxa of lichenized, lichenicolous and lichen-allied fungi are newly reported for Alabama: Acarospora fuscata, A. novomexicana, Circinaria contorta, Constrictolumina cinchonae, Dermatocarpon dolomiticum, Didymocyrtis cladoniicola, Graphis anfractuosa, G. rimulosa, Hertelidea pseudobotryosa, Heterodermia pseudospeciosa, Lecania cuprea, Marchandiomyces lignicola, Minutoexcipula miniatoexcipula, Monoblastia rappii, Multiclavula mucida, Ochrolechia trochophora, Parmotrema subsumptum, Phaeographis brasiliensis, Phaeographis inusta, Piccolia nannaria, Placynthiella icmalea, Porina scabrida, Psora decipiens, Pyrenographa irregularis, Ramboldia blochiana, Thyrea confusa, Trichothelium -

Digestive Diseases
Progress Report 進度報告 2019 Progress Report 進度報告 2019 DIGESTIVE Research Progress Summary Colorectal Cancer: DISEASES Gut microbiota: 1. The team led by Professor Jun Yu demonstrated tumour-associated neutrophils, which are that Peptostreptococcus anaerobius, an anaerobic associated with chronic infl ammation and tumour gut bacterium, could adhere to colon mucosa progression were observed in P. anaerobius- Min/+ and accelerates CRC development in mice. They treated Apc mice. Blockade of integrin α2/ϐ1 by further identifi ed that a P. anaerobius surface RGDS peptide, small interfering RNA or antibodies protein, putative cell wall binding repeat 2 all impaired P. anaerobius attachment and 01 (PCWBR2), directly interacts with colonic cell abolished P. anaerobius-mediated oncogenic Principal Investigator response in vitro and in vivo. They determined lines via α2/ϐ1 integrin. Interaction between PCWBR2 and integrin α /ϐ induced the activation that P. anaerobius drives CRC via a PCWBR2- Professor Jun Yu 2 1 of the PI3K–Akt pathway in CRC cells, leading to integrin α2/ϐ1-PI3K–Akt–NF-κB signalling axis increased cell proliferation and nuclear factor and that the PCWBR2-integrin α2/ϐ1 axis is a Team kappa-light-chain-enhancer of activated B potential therapeutic target for CRC (Nature cells (NF-κB) activation. Signifi cant expansion Communication 2019 ). Joseph Sung | Francis Chan | Henry Chan | Vincent Wong | of myeloid-derived suppressor cells, tumour- Dennis Wong | Jessie Liang | Olabisi Coker associated macrophages and granulocytic 84 85 Progress -

A Higher-Level Phylogenetic Classification of the Fungi
mycological research 111 (2007) 509–547 available at www.sciencedirect.com journal homepage: www.elsevier.com/locate/mycres A higher-level phylogenetic classification of the Fungi David S. HIBBETTa,*, Manfred BINDERa, Joseph F. BISCHOFFb, Meredith BLACKWELLc, Paul F. CANNONd, Ove E. ERIKSSONe, Sabine HUHNDORFf, Timothy JAMESg, Paul M. KIRKd, Robert LU¨ CKINGf, H. THORSTEN LUMBSCHf, Franc¸ois LUTZONIg, P. Brandon MATHENYa, David J. MCLAUGHLINh, Martha J. POWELLi, Scott REDHEAD j, Conrad L. SCHOCHk, Joseph W. SPATAFORAk, Joost A. STALPERSl, Rytas VILGALYSg, M. Catherine AIMEm, Andre´ APTROOTn, Robert BAUERo, Dominik BEGEROWp, Gerald L. BENNYq, Lisa A. CASTLEBURYm, Pedro W. CROUSl, Yu-Cheng DAIr, Walter GAMSl, David M. GEISERs, Gareth W. GRIFFITHt,Ce´cile GUEIDANg, David L. HAWKSWORTHu, Geir HESTMARKv, Kentaro HOSAKAw, Richard A. HUMBERx, Kevin D. HYDEy, Joseph E. IRONSIDEt, Urmas KO˜ LJALGz, Cletus P. KURTZMANaa, Karl-Henrik LARSSONab, Robert LICHTWARDTac, Joyce LONGCOREad, Jolanta MIA˛ DLIKOWSKAg, Andrew MILLERae, Jean-Marc MONCALVOaf, Sharon MOZLEY-STANDRIDGEag, Franz OBERWINKLERo, Erast PARMASTOah, Vale´rie REEBg, Jack D. ROGERSai, Claude ROUXaj, Leif RYVARDENak, Jose´ Paulo SAMPAIOal, Arthur SCHU¨ ßLERam, Junta SUGIYAMAan, R. Greg THORNao, Leif TIBELLap, Wendy A. UNTEREINERaq, Christopher WALKERar, Zheng WANGa, Alex WEIRas, Michael WEISSo, Merlin M. WHITEat, Katarina WINKAe, Yi-Jian YAOau, Ning ZHANGav aBiology Department, Clark University, Worcester, MA 01610, USA bNational Library of Medicine, National Center for Biotechnology Information, -

Generic Classification of the Verrucariaceae TAXON 58 (1) • February 2009: 184–208
Gueidan & al. • Generic classification of the Verrucariaceae TAXON 58 (1) • February 2009: 184–208 TAXONOMY Generic classification of the Verrucariaceae (Ascomycota) based on molecular and morphological evidence: recent progress and remaining challenges Cécile Gueidan1,16, Sanja Savić2, Holger Thüs3, Claude Roux4, Christine Keller5, Leif Tibell2, Maria Prieto6, Starri Heiðmarsson7, Othmar Breuss8, Alan Orange9, Lars Fröberg10, Anja Amtoft Wynns11, Pere Navarro-Rosinés12, Beata Krzewicka13, Juha Pykälä14, Martin Grube15 & François Lutzoni16 1 Centraalbureau voor Schimmelcultures, P.O. Box 85167, 3508 AD Utrecht, the Netherlands. c.gueidan@ cbs.knaw.nl (author for correspondence) 2 Uppsala University, Evolutionary Biology Centre, Department of Systematic Botany, Norbyvägen 18D, 752 36 Uppsala, Sweden 3 Botany Department, Natural History Museum, Cromwell Road, London, SW7 5BD, U.K. 4 Chemin des Vignes vieilles, 84120 Mirabeau, France 5 Swiss Federal Institute for Forest, Snow and Landscape Research WSL, Zürcherstrasse 111, 8903 Birmensdorf, Switzerland 6 Universidad Rey Juan Carlos, ESCET, Área de Biodiversidad y Conservación, c/ Tulipán s/n, 28933 Móstoles, Madrid, Spain 7 Icelandic Institute of Natural History, Akureyri division, P.O. Box 180, 602 Akureyri, Iceland 8 Naturhistorisches Museum Wien, Botanische Abteilung, Burgring 7, 1010 Wien, Austria 9 Department of Biodiversity and Systematic Biology, National Museum of Wales, Cathays Park, Cardiff CF10 3NP, U.K. 10 Botanical Museum, Östra Vallgatan 18, 223 61 Lund, Sweden 11 Institute for Ecology, Department of Zoology, Copenhagen University, Thorvaldsensvej 40, 1871 Frederiksberg C, Denmark 12 Departament de Biologia Vegetal (Botànica), Facultat de Biologia, Universitat de Barcelona, Diagonal 645, 08028 Barcelona, Spain 13 Laboratory of Lichenology, Institute of Botany, Polish Academy of Sciences, Lubicz 46, 31-512 Kraków, Poland 14 Finnish Environment Institute, Research Programme for Biodiversity, P.O. -

Australia Biodiversity of Biodiversity Taxonomy and and Taxonomy Plant Pathogenic Fungi Fungi Plant Pathogenic
Taxonomy and biodiversity of plant pathogenic fungi from Australia Yu Pei Tan 2019 Tan Pei Yu Australia and biodiversity of plant pathogenic fungi from Taxonomy Taxonomy and biodiversity of plant pathogenic fungi from Australia Australia Bipolaris Botryosphaeriaceae Yu Pei Tan Curvularia Diaporthe Taxonomy and biodiversity of plant pathogenic fungi from Australia Yu Pei Tan Yu Pei Tan Taxonomy and biodiversity of plant pathogenic fungi from Australia PhD thesis, Utrecht University, Utrecht, The Netherlands (2019) ISBN: 978-90-393-7126-8 Cover and invitation design: Ms Manon Verweij and Ms Yu Pei Tan Layout and design: Ms Manon Verweij Printing: Gildeprint The research described in this thesis was conducted at the Department of Agriculture and Fisheries, Ecosciences Precinct, 41 Boggo Road, Dutton Park, Queensland, 4102, Australia. Copyright © 2019 by Yu Pei Tan ([email protected]) All rights reserved. No parts of this thesis may be reproduced, stored in a retrieval system or transmitted in any other forms by any means, without the permission of the author, or when appropriate of the publisher of the represented published articles. Front and back cover: Spatial records of Bipolaris, Curvularia, Diaporthe and Botryosphaeriaceae across the continent of Australia, sourced from the Atlas of Living Australia (http://www.ala. org.au). Accessed 12 March 2019. Taxonomy and biodiversity of plant pathogenic fungi from Australia Taxonomie en biodiversiteit van plantpathogene schimmels van Australië (met een samenvatting in het Nederlands) Proefschrift ter verkrijging van de graad van doctor aan de Universiteit Utrecht op gezag van de rector magnificus, prof. dr. H.R.B.M. Kummeling, ingevolge het besluit van het college voor promoties in het openbaar te verdedigen op donderdag 9 mei 2019 des ochtends te 10.30 uur door Yu Pei Tan geboren op 16 december 1980 te Singapore, Singapore Promotor: Prof. -

<I>Heteroplacidium Compactum</I>
MYCOTAXON ISSN (print) 0093-4666 (online) 2154-8889 Mycotaxon, Ltd. ©2019 April–June 2019—Volume 134, pp. 369–376 https://doi.org/10.5248/134.369 Heteroplacidium compactum reported as a genus new to China Xiang-Min Cheng1,2, Da-Le Liu1,3, Xin-Li Wei*1,2, Jiang-Chun Wei1,2 1 State Key Laboratory of Mycology, Institute of Microbiology, Chinese Academy of Sciences, Beijing 100101, China 2 University of Chinese Academy of Sciences, Beijing 100049, China 3 College of Life Sciences, Southwest Forestry University, Kunming 650224, China * Correspondence to: [email protected] Abstract—Three specimens of Heteroplacidium compactum (Verrucariaceae), isolated from arid areas in Northwest China, were identified by morphological and phylogenetic comparisons with other species, including species of Endocarpon and Placidium (common verrucariaceous genera in the same area). This is the first report of Heteroplacidium from China. Key words—desert, ITS, lichen, morphology, taxonomy Introduction Breuss (1996) separated four genera—Anthracocarpon, Clavascidium, Heteroplacidium, and Involucropyrenium—from Catapyrenium Flot. s. lat. (Verrucariaceae) based on morphological characters. Neocatapyrenium H. Harada, Placidium A. Massal., and Scleropyrenium H. Harada are also morphologically related (Prieto & al. 2010b) but are independent monophyletic lineages within Verrucariaceae, according to phylogenetic analyses (Gueidan & al. 2007, 2009; Prieto & al. 2010b). Endocarpon Hedw. (Verrucariaceae) was relatively common during our field survey in the arid region of Northwest China (Yang & Wei 2008, Zhang & al. 2017a), but no species of Heteroplacidium Breuss was recorded. Heteroplacidium, which grows primarily on soil and rock in warm temperate 370 ... Cheng & al. Fig. 1. Heteroplacidium compactum collection sites in China. regions, is characterized by a crustose-areolate to squamulose thallus, a paraplectenchymatous or subparaplectenchymatous cortex, and clavate asci with biseriate ascospores (Prieto & al. -

Erstnachweise Und Weitere Bemerkenswerte Funde Pyrenocarper Flechten in Der Schweiz
©Österreichische Mykologische Gesellschaft, Austria, download unter www.biologiezentrum.at Österr. Z. Pilzk. 22 (2013) 93 Erstnachweise und weitere bemerkenswerte Funde pyrenocarper Flechten in der Schweiz OTHMAR BREUSS Naturhistorisches Museum Wien Botanische Abteilung Burgring 7 A-1010 Wien, Österreich Email: [email protected] PHILIPPE CLERC Conservatoire et Jardin Botaniques de la Ville de Genève Ch. de Impératrice, 1 CP 60 CH-1292 Chambésy, Schweiz Email: [email protected] Angenommen am 11. 10. 2013 Key words: Lichenized Ascomycotina, pyrenocarpous lichens, Verrucariaceae. – New records. – Alps, Switzerland. Abstract: Five species (Endocarpon adsurgens, E. loscosii, Involucropyrenium squamulosum, Placidiopsis tiroliensis, and Placidium adami-borosi) and one variety (Clavascidium lacinulatum var. atrans) of pyrenocarpous lichens are reported from Switzerland for the first time. Several additional species, previously already known from Swiss records, are reported from additional provinces. Brief notes on characteristics, ecology and distribution of the taxa are given. Zusammenfassung: Fünf Arten (Endocarpon adsurgens, E. loscosii, Involucropyrenium squamulo- sum, Placidiopsis tiroliensis und Placidium adami-borosi) und eine Varietät (Clavascidium lacinula- tum var. atrans) pyrenocarper Flechten werden erstmals aus der Schweiz gemeldet. Von einigen weite- ren aus der Schweiz bereits bekannten Arten werden zusätzliche Vorkommen in anderen Kantonen dokumentiert. Die Taxa werden hinsichtlich Charakteristika, Ökologie und Verbreitung kurz kommentiert. Trotz des wachsenden Bewusstseins um den Schutz bedrohter Arten hat die terricole Flechtenflora in der Schweiz kaum Beachtung gefunden. VUST (2011) legte die erste umfassende Studie über Ökologie und geografische Verbreitung sowie den Gefährdungsgrad erdbewohnender Flechten in der Schweiz vor. Durchgeführt wurde sie im Rahmen eines staatlichen Projekts zur Erstellung einer Roten Liste der epiphyti- schen und epigäischen Arten (SCHEIDEGGER & al. -
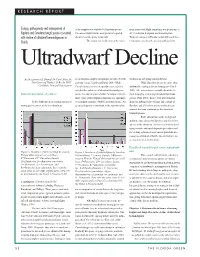
RESEARCH REPORT Ecology, Pathogenicity and Management Of
RESEARCH REPORT Ecology, pathogenicity and management of of the samples was attributed to Bipolaris species. from zoysia with blight symptoms, was pathogenic at Bipolaris and Curvularia fungal species associated Curvularia blight was the most prevalent reported 30° C in zoysia, bentgrass, and bermudagrass. with decline of ultradwarf bermudagrasses in disease from late spring to late fall. Turfgrass canopies in Florida consistently reach these Florida. The fungus was isolated off of the major- temperatures used for the previous pathogenicity Ultradwarf Decline By Dr. Lawrence E. Datnoff, Dr. Carol Stiles, Dr. ity of turfgrass samples brought into the labs (Stowell, studies from late spring through the fall. John Cisar and Matthew O. Brecht, Ph.D. personal comm.; Unruh and Davis, 2001). While While Bipolaris species are more often Candidate, Principal Investigators Curvularia species were frequently recovered from attributed to causing a disease in turfgrass (Couch both healthy and diseased ultradwarf bermudagrass 1995), little information is available about the role Rationale/description of problem: tissue, it is often unclear whether the fungus is the pri- these fungi play in affecting ultradwarf bermuda- mary cause of the turfgrass symptoms or a saprophyt- grasses (Pratt, 2001). In fact, very little is known In the Southeast, an increasing number of ic secondary organism (Stowell, personal comm.). An about the pathogenicity, etiology, and ecology of putting greens consist of the new ultradwarf accurate diagnosis is important to the superintendent Bipolaris and Curvularia species and no disease research has been conducted on the ultradwarf bermudagrasses. Basic information on the biology and ability to cause disease by Bipolaris and Curvularia species in the ultradwarf cultivars is critical for devel- oping accurate and rapid diagnostic procedures and for creating optimum, long-term integrated disease- management strategies that the superintendents can use to please their membership.