TRANSCRIPTIONAL REGULATION of Hur in RENAL STRESS
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
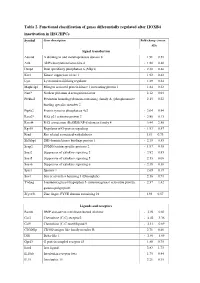
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

Podocyte Specific Knockdown of Klf15 in Podocin-Cre Klf15flox/Flox Mice Was Confirmed
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1: Podocyte specific knockdown of Klf15 in Podocin-Cre Klf15flox/flox mice was confirmed. (A) Primary glomerular epithelial cells (PGECs) were isolated from 12-week old Podocin-Cre Klf15flox/flox and Podocin-Cre Klf15+/+ mice and cultured at 37°C for 1 week. Real-time PCR was performed for Nephrin, Podocin, Synaptopodin, and Wt1 mRNA expression (n=6, ***p<0.001, Mann-Whitney test). (B) Real- time PCR was performed for Klf15 mRNA expression (n=6, *p<0.05, Mann-Whitney test). (C) Protein was also extracted and western blot analysis for Klf15 was performed. The representative blot of three independent experiments is shown in the top panel. The bottom panel shows the quantification of Klf15 by densitometry (n=3, *p<0.05, Mann-Whitney test). (D) Immunofluorescence staining for Klf15 and Wt1 was performed in 12-week old Podocin-Cre Klf15flox/flox and Podocin-Cre Klf15+/+ mice. Representative images from four mice in each group are shown in the left panel (X 20). Arrows show colocalization of Klf15 and Wt1. Arrowheads show a lack of colocalization. Asterisk demonstrates nonspecific Wt1 staining. “R” represents autofluorescence from RBCs. In the right panel, a total of 30 glomeruli were selected in each mouse and quantification of Klf15 staining in the podocytes was determined by the ratio of Klf15+ and Wt1+ cells to Wt1+ cells (n=6 mice, **p<0.01, unpaired t test). Supplementary Figure 2: LPS treated Podocin-Cre Klf15flox/flox mice exhibit a lack of recovery in proteinaceous casts and tubular dilatation after DEX administration. -

TBX15/Mir-152/KIF2C Pathway Regulates Breast Cancer Doxorubicin Resistance Via Preventing PKM2 Ubiquitination
TBX15/miR-152/KIF2C Pathway Regulates Breast Cancer Doxorubicin Resistance via Preventing PKM2 Ubiquitination Cheng-Fei Jiang Nanjing Medical University Yun-Xia Xie Zhengzhou University Ying-Chen Qian Nanjing Medical University Min Wang Nanjing Medical University Ling-Zhi Liu Thomas Jefferson University - Center City Campus: Thomas Jefferson University Yong-Qian Shu Nanjing Medical University Xiao-Ming Bai ( [email protected] ) Nanjing Medical University Bing-Hua Jiang Thomas Jefferson University Research Keywords: TBX15, miR-152, KIF2C, PKM2, Doxorubicin resistance, breast cancer Posted Date: December 28th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-130874/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/29 Abstract Background Chemoresistance is a critical risk problem for breast cancer treatment. However, mechanisms by which chemoresistance arises remains to be elucidated. The expression of T-box transcription factor 15 (TBX- 15) was found downregulated in some cancer tissues. However, role and mechanism of TBX15 in breast cancer chemoresistance is unknown. Here we aimed to identify the effects and mechanisms of TBX15 in doxorubicin resistance in breast cancer. Methods As measures of Drug sensitivity analysis, MTT and IC50 assays were used in DOX-resistant breast cancer cells. ECAR and OCR assays were used to analyze the glycolysis level, while Immunoblotting and Immunouorescence assays were used to analyze the autophagy levels in vitro. By using online prediction software, luciferase reporter assays, co-Immunoprecipitation, Western blotting analysis and experimental animals models, we further elucidated the mechanisms. Results We found TBX15 expression levels were decreased in Doxorubicin (DOX)-resistant breast cancer cells. -

The Krüppel-Like Factors in Female Reproductive System Pathologies
R C M SIMMEN and others KLFs in female reproductive 54:2 R89–R101 Review diseases The Kru¨ppel-like factors in female reproductive system pathologies Rosalia C M Simmen, Melissa E Heard, Angela M Simmen1, Maria Theresa M Montales, Meera Marji, Samantha Scanlon and John Mark P Pabona2 Department of Physiology and Biophysics, University of Arkansas for Medical Sciences, Little Rock, Correspondence Arkansas 72205, USA should be addressed 1Department of Obstetrics and Gynecology, University of Michigan Health System, Ann Arbor, Michigan 48109, USA to R C M Simmen 2Department of Internal Medicine, Harlem Hospital Center, Columbia University Medical Center, Email New York, New York 10037, USA [email protected] Abstract Female reproductive tract pathologies arise largely from dysregulation of estrogen and Key Words progesterone receptor signaling, leading to aberrant cell proliferation, survival, and " KLF differentiation. The signaling pathways orchestrated by these nuclear receptors are complex, " endometrial pathologies require the participation of many nuclear proteins serving as key binding partners or targets, " progesterone and involve a range of paracrine and autocrine regulatory circuits. The members of the " Notch Kru¨ ppel-like factor (KLF) family of transcription factors are ubiquitously expressed in " Wnt reproductive tissues and have been increasingly implicated as critical co-regulators and integrators of steroid hormone actions. Herein, we explore the involvement of KLF family members in uterine pathology, describe their currently known molecular mechanisms, Journal of Molecular and discuss their potential as targets for therapeutic intervention. Endocrinology (2015) 54, R89–R101 Journal of Molecular Endocrinology Introduction The human uterus has a unique role in the successful transcriptional pathways involve interactions with transmission of germ line DNA to guarantee the numerous nuclear co-regulators (Dasgupta & O’Malley propagation of the human species. -

Mediator of DNA Damage Checkpoint 1 (MDC1) Is a Novel Estrogen Receptor Co-Regulator in Invasive 6 Lobular Carcinoma of the Breast 7 8 Evelyn K
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.16.423142; this version posted December 16, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 1 Running Title: MDC1 co-regulates ER in ILC 2 3 Research article 4 5 Mediator of DNA damage checkpoint 1 (MDC1) is a novel estrogen receptor co-regulator in invasive 6 lobular carcinoma of the breast 7 8 Evelyn K. Bordeaux1+, Joseph L. Sottnik1+, Sanjana Mehrotra1, Sarah E. Ferrara2, Andrew E. Goodspeed2,3, James 9 C. Costello2,3, Matthew J. Sikora1 10 11 +EKB and JLS contributed equally to this project. 12 13 Affiliations 14 1Dept. of Pathology, University of Colorado Anschutz Medical Campus 15 2Biostatistics and Bioinformatics Shared Resource, University of Colorado Comprehensive Cancer Center 16 3Dept. of Pharmacology, University of Colorado Anschutz Medical Campus 17 18 Corresponding author 19 Matthew J. Sikora, PhD.; Mail Stop 8104, Research Complex 1 South, Room 5117, 12801 E. 17th Ave.; Aurora, 20 CO 80045. Tel: (303)724-4301; Fax: (303)724-3712; email: [email protected]. Twitter: 21 @mjsikora 22 23 Authors' contributions 24 MJS conceived of the project. MJS, EKB, and JLS designed and performed experiments. JLS developed models 25 for the project. EKB, JLS, SM, and AEG contributed to data analysis and interpretation. SEF, AEG, and JCC 26 developed and performed informatics analyses. MJS wrote the draft manuscript; all authors read and revised the 27 manuscript and have read and approved of this version of the manuscript. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Modeling Non-Linear Relationships Between
Georgia State University ScholarWorks @ Georgia State University Public Health Theses School of Public Health Spring 5-13-2016 Modeling Non-Linear Relationships Between DNA Methylation And Age: The Application of Regularization Methods To Predict Human Age And The Implication Of DNA Methylation In Immunosenescence Nicholas Johnson Follow this and additional works at: https://scholarworks.gsu.edu/iph_theses Recommended Citation Johnson, Nicholas, "Modeling Non-Linear Relationships Between DNA Methylation And Age: The Application of Regularization Methods To Predict Human Age And The Implication Of DNA Methylation In Immunosenescence." Thesis, Georgia State University, 2016. https://scholarworks.gsu.edu/iph_theses/473 This Thesis is brought to you for free and open access by the School of Public Health at ScholarWorks @ Georgia State University. It has been accepted for inclusion in Public Health Theses by an authorized administrator of ScholarWorks @ Georgia State University. For more information, please contact [email protected]. P a g e | 1 ABSTRACT Modeling Non-Linear Relationships Between DNA Methylation And Age: The Application of Regularization Methods To Predict Human Age And The Implication Of DNA Methylation In Immunosenescence By Nicholas David Johnson May 4, 2016 Background: Gene expression is regulated via highly coordinated epigenetic changes, the most studied of which is DNA methylation (DNAm). Many studies have shown that DNAm is linearly associated with age, and some have even used DNAm data to build predictive models of human age, which are immensely important considering that DNAm can predict health outcomes, such as all-cause mortality, better than chronological age. Nevertheless, few studies have investigated non-linear relationships between DNAm and age, which could potentially improve these predictive models. -

KLF14 Potentiates Oxidative Adaptation Via Modulating HO-1 Signaling in Castrate-Resistant Prostate Cancer
26 1 Endocrine-Related X Luo, J Liu, B Wang et al. Role of KLF14 in CRPC 26:1 181–195 Cancer RESEARCH KLF14 potentiates oxidative adaptation via modulating HO-1 signaling in castrate-resistant prostate cancer Xiao-hui Luo1,*, Jian-zhou Liu1,*, Bo Wang1,*, Qun-li Men1, Yu-quan Ju1, Feng-yan Yin1, Chao Zheng1 and Wei Li2 1Department of Urology, Baoji Center Hospital, Baoji, Shaanxi Province, People’s Republic of China 2Department of Human Anatomy, Histology and Embryology, Fourth Military Medical University, Xi’an, Shaanxi Province, People’s Republic of China Correspondence should be addressed to X Luo or W Li: [email protected] or [email protected] *(X Luo, J Liu and B Wang contributed equally to this work) Abstract Insights into the mechanisms by which key factors stimulate cell growth under androgen- Key Words depleted conditions is a premise to the development of effective treatments with clinically f Krüppel-like factor 14 significant activity in patients with castration-resistant prostate cancer (CRPC). Herein, (KLF14) we report that, the expression of Krüppel-like factor 14 (KLF14), a master transcription f CRPC factor in the regulation of lipid metabolism, was significantly induced in castration- f oxidative stress insensitive PCa cells and tumor tissues from a mouse xenograft model of CRPC. KLF14 f heme oxygenase-1 (HO-1) upregulation in PCa cells, which was stimulated upstream by oxidative stress, was f transcriptional regulation dependent on multiple pathways including PI3K/AKT, p42/p44 MAPK, AMPK and PKC pathways. By means of ectopic overexpression and genetic inactivation, we further show that KLF14 promoted cell growth via positive regulation of the antioxidant response under androgen-depleted conditions. -

Supplementary Table S5. Differentially Expressed Gene Lists of PD-1High CD39+ CD8 Tils According to 4-1BB Expression Compared to PD-1+ CD39- CD8 Tils
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Immunother Cancer Supplementary Table S5. Differentially expressed gene lists of PD-1high CD39+ CD8 TILs according to 4-1BB expression compared to PD-1+ CD39- CD8 TILs Up- or down- regulated genes in Up- or down- regulated genes Up- or down- regulated genes only PD-1high CD39+ CD8 TILs only in 4-1BBneg PD-1high CD39+ in 4-1BBpos PD-1high CD39+ CD8 compared to PD-1+ CD39- CD8 CD8 TILs compared to PD-1+ TILs compared to PD-1+ CD39- TILs CD39- CD8 TILs CD8 TILs IL7R KLRG1 TNFSF4 ENTPD1 DHRS3 LEF1 ITGA5 MKI67 PZP KLF3 RYR2 SIK1B ANK3 LYST PPP1R3B ETV1 ADAM28 H2AC13 CCR7 GFOD1 RASGRP2 ITGAX MAST4 RAD51AP1 MYO1E CLCF1 NEBL S1PR5 VCL MPP7 MS4A6A PHLDB1 GFPT2 TNF RPL3 SPRY4 VCAM1 B4GALT5 TIPARP TNS3 PDCD1 POLQ AKAP5 IL6ST LY9 PLXND1 PLEKHA1 NEU1 DGKH SPRY2 PLEKHG3 IKZF4 MTX3 PARK7 ATP8B4 SYT11 PTGER4 SORL1 RAB11FIP5 BRCA1 MAP4K3 NCR1 CCR4 S1PR1 PDE8A IFIT2 EPHA4 ARHGEF12 PAICS PELI2 LAT2 GPRASP1 TTN RPLP0 IL4I1 AUTS2 RPS3 CDCA3 NHS LONRF2 CDC42EP3 SLCO3A1 RRM2 ADAMTSL4 INPP5F ARHGAP31 ESCO2 ADRB2 CSF1 WDHD1 GOLIM4 CDK5RAP1 CD69 GLUL HJURP SHC4 GNLY TTC9 HELLS DPP4 IL23A PITPNC1 TOX ARHGEF9 EXO1 SLC4A4 CKAP4 CARMIL3 NHSL2 DZIP3 GINS1 FUT8 UBASH3B CDCA5 PDE7B SOGA1 CDC45 NR3C2 TRIB1 KIF14 TRAF5 LIMS1 PPP1R2C TNFRSF9 KLRC2 POLA1 CD80 ATP10D CDCA8 SETD7 IER2 PATL2 CCDC141 CD84 HSPA6 CYB561 MPHOSPH9 CLSPN KLRC1 PTMS SCML4 ZBTB10 CCL3 CA5B PIP5K1B WNT9A CCNH GEM IL18RAP GGH SARDH B3GNT7 C13orf46 SBF2 IKZF3 ZMAT1 TCF7 NECTIN1 H3C7 FOS PAG1 HECA SLC4A10 SLC35G2 PER1 P2RY1 NFKBIA WDR76 PLAUR KDM1A H1-5 TSHZ2 FAM102B HMMR GPR132 CCRL2 PARP8 A2M ST8SIA1 NUF2 IL5RA RBPMS UBE2T USP53 EEF1A1 PLAC8 LGR6 TMEM123 NEK2 SNAP47 PTGIS SH2B3 P2RY8 S100PBP PLEKHA7 CLNK CRIM1 MGAT5 YBX3 TP53INP1 DTL CFH FEZ1 MYB FRMD4B TSPAN5 STIL ITGA2 GOLGA6L10 MYBL2 AHI1 CAND2 GZMB RBPJ PELI1 HSPA1B KCNK5 GOLGA6L9 TICRR TPRG1 UBE2C AURKA Leem G, et al. -

Ten Commandments for a Good Scientist
Unravelling the mechanism of differential biological responses induced by food-borne xeno- and phyto-estrogenic compounds Ana María Sotoca Covaleda Wageningen 2010 Thesis committee Thesis supervisors Prof. dr. ir. Ivonne M.C.M. Rietjens Professor of Toxicology Wageningen University Prof. dr. Albertinka J. Murk Personal chair at the sub-department of Toxicology Wageningen University Thesis co-supervisor Dr. ir. Jacques J.M. Vervoort Associate professor at the Laboratory of Biochemistry Wageningen University Other members Prof. dr. Michael R. Muller, Wageningen University Prof. dr. ir. Huub F.J. Savelkoul, Wageningen University Prof. dr. Everardus J. van Zoelen, Radboud University Nijmegen Dr. ir. Toine F.H. Bovee, RIKILT, Wageningen This research was conducted under the auspices of the Graduate School VLAG Unravelling the mechanism of differential biological responses induced by food-borne xeno- and phyto-estrogenic compounds Ana María Sotoca Covaleda Thesis submitted in fulfillment of the requirements for the degree of doctor at Wageningen University by the authority of the Rector Magnificus Prof. dr. M.J. Kropff, in the presence of the Thesis Committee appointed by the Academic Board to be defended in public on Tuesday 14 September 2010 at 4 p.m. in the Aula Unravelling the mechanism of differential biological responses induced by food-borne xeno- and phyto-estrogenic compounds. Ana María Sotoca Covaleda Thesis Wageningen University, Wageningen, The Netherlands, 2010, With references, and with summary in Dutch. ISBN: 978-90-8585-707-5 “Caminante no hay camino, se hace camino al andar. Al andar se hace camino, y al volver la vista atrás se ve la senda que nunca se ha de volver a pisar” - Antonio Machado – A mi madre. -

Supplementary Table 1. Pain and PTSS Associated Genes (N = 604
Supplementary Table 1. Pain and PTSS associated genes (n = 604) compiled from three established pain gene databases (PainNetworks,[61] Algynomics,[52] and PainGenes[42]) and one PTSS gene database (PTSDgene[88]). These genes were used in in silico analyses aimed at identifying miRNA that are predicted to preferentially target this list genes vs. a random set of genes (of the same length). ABCC4 ACE2 ACHE ACPP ACSL1 ADAM11 ADAMTS5 ADCY5 ADCYAP1 ADCYAP1R1 ADM ADORA2A ADORA2B ADRA1A ADRA1B ADRA1D ADRA2A ADRA2C ADRB1 ADRB2 ADRB3 ADRBK1 ADRBK2 AGTR2 ALOX12 ANO1 ANO3 APOE APP AQP1 AQP4 ARL5B ARRB1 ARRB2 ASIC1 ASIC2 ATF1 ATF3 ATF6B ATP1A1 ATP1B3 ATP2B1 ATP6V1A ATP6V1B2 ATP6V1G2 AVPR1A AVPR2 BACE1 BAMBI BDKRB2 BDNF BHLHE22 BTG2 CA8 CACNA1A CACNA1B CACNA1C CACNA1E CACNA1G CACNA1H CACNA2D1 CACNA2D2 CACNA2D3 CACNB3 CACNG2 CALB1 CALCRL CALM2 CAMK2A CAMK2B CAMK4 CAT CCK CCKAR CCKBR CCL2 CCL3 CCL4 CCR1 CCR7 CD274 CD38 CD4 CD40 CDH11 CDK5 CDK5R1 CDKN1A CHRM1 CHRM2 CHRM3 CHRM5 CHRNA5 CHRNA7 CHRNB2 CHRNB4 CHUK CLCN6 CLOCK CNGA3 CNR1 COL11A2 COL9A1 COMT COQ10A CPN1 CPS1 CREB1 CRH CRHBP CRHR1 CRHR2 CRIP2 CRYAA CSF2 CSF2RB CSK CSMD1 CSNK1A1 CSNK1E CTSB CTSS CX3CL1 CXCL5 CXCR3 CXCR4 CYBB CYP19A1 CYP2D6 CYP3A4 DAB1 DAO DBH DBI DICER1 DISC1 DLG2 DLG4 DPCR1 DPP4 DRD1 DRD2 DRD3 DRD4 DRGX DTNBP1 DUSP6 ECE2 EDN1 EDNRA EDNRB EFNB1 EFNB2 EGF EGFR EGR1 EGR3 ENPP2 EPB41L2 EPHB1 EPHB2 EPHB3 EPHB4 EPHB6 EPHX2 ERBB2 ERBB4 EREG ESR1 ESR2 ETV1 EZR F2R F2RL1 F2RL2 FAAH FAM19A4 FGF2 FKBP5 FLOT1 FMR1 FOS FOSB FOSL2 FOXN1 FRMPD4 FSTL1 FYN GABARAPL1 GABBR1 GABBR2 GABRA2 GABRA4