Pdf/Infopackage Kinex.Pdf for a Com- Domain Inhibition (15)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Human RET Kinase Protein (His Tag)
Human RET Kinase Protein (His Tag) Catalog Number: 11997-H08H1 General Information SDS-PAGE: Gene Name Synonym: CDHF12; CDHR16; HSCR1; MEN2A; MEN2B; MTC1; PTC; RET-ELE1; RET51 Protein Construction: A DNA sequence encoding the extracellular domain of human RET (P07949-1) (Met 1-Arg 635) was fused with a polyhistidine tag at the N- terminus. Source: Human Expression Host: HEK293 Cells QC Testing Purity: > 92 % as determined by SDS-PAGE Protein Description Endotoxin: RET proto-oncogene, also known as RET, is a cell-surface molecule that < 1.0 EU per μg of the protein as determined by the LAL method transduce signals for cell growth and differentiation. It contains 1 cadherin domain and 1 protein kinase domain. RET proto-oncogene belongs to the Stability: protein kinase superfamily, tyr protein kinase family. RET proto-oncogene is involved in numerous cellular mechanisms including cell proliferation, ℃ Samples are stable for up to twelve months from date of receipt at -70 neuronal navigation, cell migration, and cell differentiation upon binding with glial cell derived neurotrophic factor family ligands. It phosphorylates Leu 29 Predicted N terminal: PTK2/FAK1 and regulates both cell death/survival balance and positional Molecular Mass: information. RET is required for the molecular mechanisms orchestration during intestine organogenesis; involved in the development of enteric The recombinant human RET consists of 618 amino acids and has a nervous system and renal organogenesis during embryonic life; promotes calculated molecular mass of 69.1 kDa. The apparent molecular mass of the formation of Peyer's patch-like structures; modulates cell adhesion via the protein is approximately 110-120 kDa in SDS-PAGE under reducing its cleavage; involved in the development of the neural crest. -

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

The Regulatory Roles of Phosphatases in Cancer
Oncogene (2014) 33, 939–953 & 2014 Macmillan Publishers Limited All rights reserved 0950-9232/14 www.nature.com/onc REVIEW The regulatory roles of phosphatases in cancer J Stebbing1, LC Lit1, H Zhang, RS Darrington, O Melaiu, B Rudraraju and G Giamas The relevance of potentially reversible post-translational modifications required for controlling cellular processes in cancer is one of the most thriving arenas of cellular and molecular biology. Any alteration in the balanced equilibrium between kinases and phosphatases may result in development and progression of various diseases, including different types of cancer, though phosphatases are relatively under-studied. Loss of phosphatases such as PTEN (phosphatase and tensin homologue deleted on chromosome 10), a known tumour suppressor, across tumour types lends credence to the development of phosphatidylinositol 3--kinase inhibitors alongside the use of phosphatase expression as a biomarker, though phase 3 trial data are lacking. In this review, we give an updated report on phosphatase dysregulation linked to organ-specific malignancies. Oncogene (2014) 33, 939–953; doi:10.1038/onc.2013.80; published online 18 March 2013 Keywords: cancer; phosphatases; solid tumours GASTROINTESTINAL MALIGNANCIES abs in sera were significantly associated with poor survival in Oesophageal cancer advanced ESCC, suggesting that they may have a clinical utility in Loss of PTEN (phosphatase and tensin homologue deleted on ESCC screening and diagnosis.5 chromosome 10) expression in oesophageal cancer is frequent, Cao et al.6 investigated the role of protein tyrosine phosphatase, among other gene alterations characterizing this disease. Zhou non-receptor type 12 (PTPN12) in ESCC and showed that PTPN12 et al.1 found that overexpression of PTEN suppresses growth and protein expression is higher in normal para-cancerous tissues than induces apoptosis in oesophageal cancer cell lines, through in 20 ESCC tissues. -

Role of Focal Adhesion Kinase in Small-Cell Lung Cancer and Its Potential As a Therapeutic Target
cancers Review Role of Focal Adhesion Kinase in Small-Cell Lung Cancer and Its Potential as a Therapeutic Target Frank Aboubakar Nana 1,2 , Marie Vanderputten 1 and Sebahat Ocak 1,3,* 1 Institut de Recherche Expérimentale et Clinique (IREC), Pôle de Pneumologie, ORL et Dermatologie (PNEU), Université catholique de Louvain (UCLouvain), 1200 Brussels, Belgium; [email protected] (F.A.N.); [email protected] (M.V.) 2 Division of Pneumology, Cliniques Universitaires St-Luc, UCL, 1200 Brussels, Belgium 3 Division of Pneumology, CHU UCL Namur (Godinne Site), UCL, 5530 Yvoir, Belgium * Correspondence: [email protected]; Tel.: +32-2-764-9448; Fax: +32-2-764-9440 Received: 15 September 2019; Accepted: 24 October 2019; Published: 29 October 2019 Abstract: Small-cell lung cancer (SCLC) represents 15% of all lung cancers and it is clinically the most aggressive type, being characterized by a tendency for early metastasis, with two-thirds of the patients diagnosed with an extensive stage (ES) disease and a five-year overall survival (OS) as low as 5%. There are still no effective targeted therapies in SCLC despite improved understanding of the molecular steps leading to SCLC development and progression these last years. After four decades, the only modest improvement in OS of patients suffering from ES-SCLC has recently been shown in a trial combining atezolizumab, an anti-PD-L1 immune checkpoint inhibitor, with carboplatin and etoposide, chemotherapy agents. This highlights the need to pursue research efforts in this field. Focal adhesion kinase (FAK) is a non-receptor protein tyrosine kinase that is overexpressed and activated in several cancers, including SCLC, and contributing to cancer progression and metastasis through its important role in cell proliferation, survival, adhesion, spreading, migration, and invasion. -
Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach
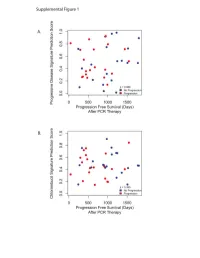
Cancers 2019, 11, x S1 of S9 Supplementary Materials: Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach Young Rae Kim, Yong Wan Kim, Suh Eun Lee, Hye Won Yang and Sung Young Kim Table S1. Characteristics of individual studies. Sample Size Origin of Cancer Drug Dataset Platform S AR (Cell Lines) Lung cancer Agilent-014850 Whole Human GSE34228 26 26 (PC9) Genome Microarray 4x44K Gefitinib Epidermoid carcinoma Affymetrix Human Genome U133 GSE10696 3 3 (A431) Plus 2.0 Head and neck cancer Illumina HumanHT-12 V4.0 GSE62061 12 12 (Cal-27, SSC-25, FaDu, expression beadchip SQ20B) Erlotinib Head and neck cancer Illumina HumanHT-12 V4.0 GSE49135 3 3 (HN5) expression beadchip Lung cancer (HCC827, Illumina HumanHT-12 V3.0 GSE38310 3 6 ER3, T15-2) expression beadchip Lung cancer Illumina HumanHT-12 V3.0 GSE62504 1 2 (HCC827) expression beadchip Afatinib Lung cancer * Illumina HumanHT-12 V4.0 GSE75468 1 3 (HCC827) expression beadchip Head and neck cancer Affymetrix Human Genome U133 Cetuximab GSE21483 3 3 (SCC1) Plus 2.0 Array GEO, gene expression omnibus; GSE, gene expression series; S, sensitive; AR, acquired EGFR-TKI resistant; * Lung Cancer Cells Derived from Tumor Xenograft Model. Table S2. The performances of four penalized regression models. Model ACC precision recall F1 MCC AUROC BRIER Ridge 0.889 0.852 0.958 0.902 0.782 0.964 0.129 Lasso 0.944 0.957 0.938 0.947 0.889 0.991 0.042 Elastic Net 0.978 0.979 0.979 0.979 0.955 0.999 0.023 EPSGO Elastic Net 0.989 1.000 0.979 0.989 0.978 1.000 0.018 AUROC, area under curve of receiver operating characteristic; ACC, accuracy; MCC, Matthews correlation coefficient; EPSGO, Efficient Parameter Selection via Global Optimization algorithm. -

A Homozygous Deletion Within the Carbonic Anhydrase-Like Domain of the Ptprg Gene in Murine L-Cells1
(CANCER RESEARCH 53. 14«)»-1502.April1. 1993| Advances in Brief A Homozygous Deletion within the Carbonic Anhydrase-like Domain of the Ptprg Gene in Murine L-Cells1 Kishore K. Wary, Zhuangwei Lou, Arthur M. Buchberg, Linda D. Siracusa, Teresa Druck, Sal LaForgia, and Kay Huebner2 Jefferson Cancer Instilare. Thomas Jefferson Medical College, Philadelphia. Pennsylvania ¡9ÃŒ07 Abstract and murine predicted amino acid sequences for this isoform are greater than 90% identical: the Ptprg gene contains a single trans- Protein tyrosine phosphatases, on purely theoretical grounds, were sug membrane domain and two tandem tyrosine phosphatase domains. gested as possible tumor suppressor genes, and receptor protein tyrosine The extracellular region contains one fibronectin type repeat and. like phosphatase 7 (PTPRG ) has been proposed, on the basis of its location at the PTPRZ gene (7).' an NH2-terminal region of 266 amino acids with human chromosome region 3pl4.2, specifically as a tumor suppressor gene —¿30%sequence similarity to members of the carbonic anhydrase for renal cell carcinoma. We have isolated murine genomic and complementary DNA clones for enzyme family (8). analysis and mapping of the murine Ptprg locus; interspecific backcross During an investigation of the genomic organization of human and analysis showed that the Ptprg locus maps to the centramene region of mouse Ptprg loci, we noted extensive polymorphism of the murine mouse chromosome 14. We also observed a homozygous, intragenic dele PÃprglocus5 and a homozygous deletion of a portion of the CA-like tion in the Ptprg gene in all donai derivatives of the original I.-cell strain, domain in murine L-cell lines, which we undertook to describe in a methylcholanthrene-treated mouse connective tissue cell line which pro detail, since homozygous deletion of a gene is one of the hallmarks of duces sarcomas in syngeneic mice. -
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

The Extracellular Matrix for Bone Regeneration
This chapter is based on human mesenchymal stromal cells in adhesion to cell-derived 5 extracellular matrix and titanium: a comparative kinome profi le analysis Marta Baroncelli1^, Gwenny M. Fuhler2^, Jeroen van de Peppel1, Willian F. Zambuzzi3, Johannes P.T.M van Leeuwen1, Bram C.J van der Eerden1°, Maikel P. Peppelenbosch2° 1Department of Internal Medicine, Erasmus MC. University Medical Center Rotterdam, Wytemaweg 80, 3015 CN, Rotterdam, The Netherlands 2Department of Gastroenterology and Hepatology, Erasmus MC, University Medical Center Rotterdam, Wytemaweg 80, 3015 CN, Rotterdam, The Netherlands 3Laboratorio de Bioensaios e Dinâmica Celular, Dep.to de Quimica e Bioquimica, Instituto de Biociências, Universidade Estadual Paulista- UNESP, Campus Botucatu, São Paulo, SP Brazil. ^, °: both authors contributed equally to this work Submitted Chapter 5 ABSTRACT The extracellular matrix (ECM) is an essential component of tissue architecture that physically supports cells and actively influences stem cell behaviour, by modulating kinase-mediated signaling cascades that are the key regulators of signal transduc- tion. Cell-derived ECMs have recently emerged in the context of bone regeneration, as they reproduce physiological tissue-architecture and ameliorate the promising properties of mesenchymal stromal cells (MSCs). Titanium scaffolds show good mechanical properties and porosity to facilitate cell adhesion, and thus have been routinely used for bone tissue engineering applications. The aim of this study was to analyze the kinomic signature of human MSCs in adhesion to an osteoblast-derived ECM that we have previously shown to be osteopromotive, and to compare it to MSCs on titanium. PamChip kinase array analysis revealed 63 phosphorylated peptides on ECM and 59 on titanium, with MSCs on ECM exhibiting significant higher levels of kinase activity than those on titanium. -

Wrangling Phosphoproteomic Data to Elucidate Cancer Signaling Pathways
University of Montana ScholarWorks at University of Montana Biological Sciences Faculty Publications Biological Sciences 1-3-2013 Wrangling Phosphoproteomic Data to Elucidate Cancer Signaling Pathways Mark L. Grimes University of Montana - Missoula, [email protected] Wan-Jui Lee Laurens van der Maaten Paul Shannon Follow this and additional works at: https://scholarworks.umt.edu/biosci_pubs Part of the Biology Commons Let us know how access to this document benefits ou.y Recommended Citation Grimes, Mark L.; Lee, Wan-Jui; van der Maaten, Laurens; and Shannon, Paul, "Wrangling Phosphoproteomic Data to Elucidate Cancer Signaling Pathways" (2013). Biological Sciences Faculty Publications. 178. https://scholarworks.umt.edu/biosci_pubs/178 This Article is brought to you for free and open access by the Biological Sciences at ScholarWorks at University of Montana. It has been accepted for inclusion in Biological Sciences Faculty Publications by an authorized administrator of ScholarWorks at University of Montana. For more information, please contact [email protected]. Wrangling Phosphoproteomic Data to Elucidate Cancer Signaling Pathways Mark L. Grimes1*, Wan-Jui Lee2, Laurens van der Maaten2, Paul Shannon3 1 Division of Biological Sciences, University of Montana, Missoula, Montana, United States of America, 2 Pattern Recognition and Bioinformatics Group, Delft University of Technology, CD Delft, The Netherlands, 3 Fred Hutchison Cancer Research Institute, Seattle, Washington, United States of America Abstract The interpretation of biological data sets is essential for generating hypotheses that guide research, yet modern methods of global analysis challenge our ability to discern meaningful patterns and then convey results in a way that can be easily appreciated. Proteomic data is especially challenging because mass spectrometry detectors often miss peptides in complex samples, resulting in sparsely populated data sets. -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942 -

Rabbit Anti-Phospho-Ret-SL3384R-FITC
SunLong Biotech Co.,LTD Tel: 0086-571- 56623320 Fax:0086-571- 56623318 E-mail:[email protected] www.sunlongbiotech.com Rabbit Anti-Phospho-Ret SL3384R-FITC Product Name: Anti-Phospho-Ret (Tyr905)/FITC Chinese Name: FITC标记的磷酸化RET原癌基因抗体 Ret(Phospho Y905); Ret Proto-Oncogene; Cadherin-Related Family Member 16; Rearranged During Transfection; RET Receptor Tyrosine Kinase; Cadherin Family Member 12; Proto-Oncogene C-Ret; EC 2.7.10.1; CDHF12; CDHR16; RET51; PTC; Ret Proto-Oncogene (Multiple Endocrine Neoplasia And Medullary Thyroid Carcinoma Alias: 1, Hirschsprung Disease) ; Multiple Endocrine Neoplasia And Medullary Thyroid Carcinoma 1; Proto-Oncogene Tyrosine-Protein Kinase Receptor Ret; Hydroxyaryl- Protein Kinase; RET Transforming Sequence; Receptor Tyrosine Kinase; Hirschsprung Disease 1; Oncogene RET; EC 2.7.10; RET-ELE1; MEN2B; HSCR1; MEN2A; MTC1; RET_HUMAN. Organism Species: Rabbit Clonality: Polyclonal React Species: Human,Mouse,Rat,Chicken,Dog,Cow,Rabbit,Sheep,Guinea Pig, IF=1:50-200 Applications: not yet tested in other applications. optimalwww.sunlongbiotech.com dilutions/concentrations should be determined by the end user. Molecular weight: 34/76/122kDa Form: Lyophilized or Liquid Concentration: 1mg/ml KLH conjugated Synthesised phosphopeptide derived from human Ret around the immunogen: phosphorylation site of Tyr905 Lsotype: IgG Purification: affinity purified by Protein A Storage Buffer: 0.01M TBS(pH7.4) with 1% BSA, 0.03% Proclin300 and 50% Glycerol. Store at -20 ℃ for one year. Avoid repeated freeze/thaw cycles. The lyophilized antibody is stable at room temperature for at least one month and for greater than a year Storage: when kept at -20℃. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 ℃. -

Receptor Protein Tyrosine Phosphatases Control Purkinje Neuron Firing
Receptor protein tyrosine phosphatases control Purkinje neuron firing Alexander S. Brown1, Pratap Meera2, Gabe Quinones1, Jessica Magri1, Thomas S. Otis3, Stefan M. Pulst4, and Anthony E. Oro1,5 1Program in Epithelial Biology Stanford University School of Medicine, Stanford CA, 2Department of Neurobiology University of California Los Angeles, Los Angeles CA 3Sainsbury Wellcome Centre for Neural Circuits and Behavior, University College London, London, United Kingdom 4Department of Neurology, University of Utah Medical Center, Salt Lake City, UT 5To whom correspondence should be addressed: Anthony E.Oro ( [email protected]) . Abstract (173/200 words): Spinocerebellar ataxias (SCA) are a genetically heterogeneous family of cerebellar neurodegenerative diseases characterized by abnormal firing of Purkinje neurons and degeneration. We recently demonstrated the slowed firing rates seen in several SCAs share a common etiology of hyper-activation of the Src family of non-receptor tyrosine kinases (SFKs)1. However, because of the lack of effective neuroactive, clinically available SFK inhibitors, alternative mechanisms to modulate SFK activity are needed. Previous studies demonstrate that SFK activity can be enhanced by the removal of inhibitory phospho-marks by receptor-protein-tyrosine phosphatases (RPTPs)2,3. In this Extra View we show that MTSS1 inhibits SFK activity through the binding and inhibition of a subset of the RPTP family members. RPTP activity normally results in SFK activation in vitro, and lowering RPTP activity in cerebellar slices using recently described RPTP peptide inhibitors increases the suppressed Purkinje neuron basal firing rates seen in two different SCA models. Together these results identify RPTPs as novel effectors of cerebellar activity, extending the MTSS1/SFK regulatory circuit we previously described and expanding the therapeutic targets for SCA patients.