A Supergene-Linked Estrogen Receptor Drives Alternative Phenotypes in a Polymorphic Songbird
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genomic Response to the Social Environment: Implications for Health Outcomes
Genomic Response to the Social Environment: Implications for Health Outcomes June 24, 2020 Workshop Summary Workshop Goals Jessica M. Gill, PhD, RN, FAAN Acting Deputy Director, National Institute of Nursing Research (NINR) Dr. Gill welcomed participants and acknowledged funding support from the National Institutes of Health (NIH) Office of Disease Prevention (ODP) and the NIH Office of Behavioral and Social Sciences Research (OBSSR) as well as colleagues across NIH Institutes, Centers, and Offices who participated in planning the workshop. Although it has long been recognized that the social environment can influence the risk, manifestation, and trajectory of disease and associated symptoms, the underlying biological mechanisms remain largely understudied. This transdisciplinary event will address the relationship among genomics (e.g., epigenomics, gene expression, microbiome, telomeres), the social environment, and health outcomes across conditions, populations, and the lifespan. Discussions will include an exchange of ideas to advance a social genomics research strategy that illuminates genetic influences on disease burden and to inform future interventions. Overview of the Day and Introduction of Keynote Speakers Lois Tully, PhD Program Director, NINR Dr. Tully provided an overview of the workshop agenda and introduced the keynote speakers, Drs. Janice Kiecolt-Glaser and Steve Cole. She encouraged participants to engage in thoughtful discussion about potential next steps toward integrating a social genomics approach into personalized strategies that prevent, modify, or buffer adverse social environments and lead to better health outcomes. Lovesick: How Couples’ Relationships Influence Health Janice Kiecolt-Glaser, PhD Director, Institute for Behavioral Medicine Research Distinguished University Professor Institute for Behavioral Medicine Research The Ohio State University College of Medicine Dr. -

Genomic and Transcriptomic Analyses of the Subterranean Termite
bioRxiv preprint doi: https://doi.org/10.1101/2021.07.11.451559; this version posted July 12, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Genomic and transcriptomic analyses of the 2 subterranean termite Reticulitermes speratus: gene 3 duplication facilitates social evolution 4 5 Shuji Shigenobu*+1,2, Yoshinobu Hayashi+3, Dai Watanabe 4,5, Gaku Tokuda6, 6 Masaru Y Hojo6,19, Kouhei Toga5,7, Ryota Saiki5, Hajime Yaguchi5,8, Yudai 7 Masuoka5,9, Ryutaro Suzuki5,10, Shogo Suzuki5, Moe Kimura11, Masatoshi 8 Matsunami4,12, Yasuhiro Sugime4, Kohei Oguchi4,10,13, Teruyuki Niimi2,14, Hiroki 9 Gotoh15,20, Masaru K Hojo8, Satoshi Miyazaki16, Atsushi Toyoda17, Toru Miura*4,13, 10 Kiyoto Maekawa*18 11 12 1) NIBB Research Core Facilities, National Institute for Basic Biology, Okazaki, 13 444-8585 Japan 14 2) Department of Basic Biology, School of Life Science, The Graduate University for 15 Advanced Studies, SOKENDAI, Nishigonaka 38, Myodaiji, Okazaki, Aichi 444- 16 8585, Japan 17 3) Department of Biology, Keio University, Hiyoshi, Yokohama, 223-8521, Japan 18 4) Faculty of Environmental Earth Science, Hokkaido University, Sapporo, 19 Hokkaido, 060-0810, Japan 20 5) Graduate School of Science and Engineering, University of Toyama, Toyama, 21 930-8555, Japan 22 6) Tropical Biosphere Research Center, COMB, University of the Ryukyus, 23 Nishihara, Okinawa 903-0213, Japan 24 7) Department of Biosciences, College of Humanities and Sciences, -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Regular Article
Regular Article LYMPHOID NEOPLASIA Opposing regulation of BIM and BCL2 controls glucocorticoid-induced apoptosis of pediatric acute lymphoblastic leukemia cells Duohui Jing,1,2 Vivek A. Bhadri,1,2 Dominik Beck,2,3 Julie A. I. Thoms,2,3 Nurul A. Yakob,1,2 Jason W. H. Wong,2,3 Kathy Knezevic,2,3 John E. Pimanda,2,3,4 and Richard B. Lock1,2 1Children’s Cancer Institute Australia for Medical Research, 2Lowy Cancer Research Centre, and the 3Prince of Wales Clinical School, University of New South Wales, Sydney, Australia; and 4Department of Haematology, Prince of Wales Hospital, Sydney, Australia Key Points Glucocorticoids are critical components of combination chemotherapy regimens in pediatric acute lymphoblastic leukemia (ALL). The proapoptotic BIM protein is an important • The glucocorticoid receptor mediator of glucocorticoid-induced apoptosis in normal and malignant lymphocytes, coordinately regulates the whereas the antiapoptotic BCL2 confers resistance. The signaling pathways regulating BIM antiapoptotic BCL2 and and BCL2 expression in glucocorticoid-treated lymphoid cells remain unclear. In this study, proapoptotic BIM genes in pediatric ALL patient-derived xenografts (PDXs) inherently sensitive or resistant to pediatric ALL cells in vivo. glucocorticoids were exposed to dexamethasone in vivo. Microarray analysis showed KLF13 MYB • GR binding at a novel intronic that and gene expression changes were significantly greater in dexamethasone- sensitive than -resistant PDXs. Chromatin immunoprecipitation (ChIP) analysis detected region is associated with glucocorticoid receptor (GR) binding at the KLF13 promoter to trigger KLF13 expression BIM transcription and only in sensitive PDXs. Next, KLF13 bound to the MYB promoter, deactivating MYB ex- dexamethasone sensitivity in pression only in sensitive PDXs. -

Stance As Nonself Initiates a Complex Cascade of Events Which
JOURNAL OF SURGICAL RESEARCH 39, l72- 18 1 ( 1985) CURRENT RESEARCH REVIEW Major Histocompatibility Complex Regulation of the Immune Response DAVID H. GUTMANN, PH.D., AND JOHN E. NIEDERHUBER, M.D. Departments of Microbiology and Immunology and Surgery, The University of Michigan Medical School, Ann Arbor, Michigan 48109 Submitted for publication July 18, 1984 The ability of an organism to distinguish self from nonself is determined by a cluster of genes located in the major histocompatibility complex. Recent advances in molecular genetics and cellular immunology have begun to elucidate the mechanisms responsible for immune response regulation. In this review article, the genetic organization of the murine and human major histocompatibility complexes and the manner by which their gene products modulate immune responsiveness are discussed. 0 1985 Academic Press. Inc. INTRODUCTION ulate the immune response in several ways. Whether this regulation is their primary The body’s recognition of a foreign sub- function is not clear, as it seemspossible that stance as nonself initiates a complex cascade other as yet undiscovered levels of regulation of events which leads to an immune response. of the immune response may exist. Since Regulation of this immune response is con- much of our understanding of the role of the trolled by a cluster of tightly linked genes MHC in immune regulation has resulted encoded within the major histocompatibility from studies on the mouse MHC, the struc- complex (MHC) [ 1, 21. While the MHC was ture and function of both mouse and, when originally defined as the site for control of available, human MHC molecules will be graft rejection, it is now known that MHC discussed in this review. -

WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT (51) International Patent Classification: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, C12Q 1/68 (2018.01) A61P 31/18 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, C12Q 1/70 (2006.01) HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, (21) International Application Number: MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, PCT/US2018/056167 OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, (22) International Filing Date: SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, 16 October 2018 (16. 10.2018) TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every kind of regional protection available): ARIPO (BW, GH, (26) Publication Language: English GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, TZ, (30) Priority Data: UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, 62/573,025 16 October 2017 (16. 10.2017) US TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, EE, ES, FI, FR, GB, GR, HR, HU, ΓΕ , IS, IT, LT, LU, LV, (71) Applicant: MASSACHUSETTS INSTITUTE OF MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, SM, TECHNOLOGY [US/US]; 77 Massachusetts Avenue, TR), OAPI (BF, BJ, CF, CG, CI, CM, GA, GN, GQ, GW, Cambridge, Massachusetts 02139 (US). -

Chromosomal Rearrangements Maintain a Polymorphic Supergene Controlling Butterfly Mimicry
Published in which should be cited to refer to this work. Chromosomal rearrangements maintain a polymorphic supergene controlling butterfly mimicry Mathieu Joron1,2,3, Lise Frezal1*, Robert T. Jones4*, Nicola L. Chamberlain4, Siu F. Lee5, Christoph R. Haag6, Annabel Whibley1, Michel Becuwe2, Simon W. Baxter7, Laura Ferguson7, Paul A. Wilkinson4, Camilo Salazar8, Claire Davidson9, Richard Clark9, Michael A. Quail9, Helen Beasley9, Rebecca Glithero9, Christine Lloyd9, Sarah Sims9, Matthew C. Jones9, Jane Rogers9, Chris D. Jiggins7 & Richard H. ffrench-Constant4 Supergenes are tight clusters of loci that facilitate the co-segregation be tightly linked from the outset or whether the association between of adaptive variation, providing integrated control of complex elements can be acquired, either gradually or in a single mutational adaptive phenotypes1. Polymorphic supergenes, in which specific step7–10,16–19. Chromosomal rearrangements, which can bring genes into combinations of traits are maintained within a single population, closer physical association and influence local recombination, offer one were first described for ‘pin’ and ‘thrum’ floral types in Primula1 and route through which supergenes may be assembled from more loosely Fagopyrum2, but classic examples are also found in insect mimicry3–5 linked components7,8,17–19. Although there are many cases of poly- and snail morphology6. Understanding the evolutionary mechan- morphic inversions associated with adaptive variation12,20, variation is isms that generate these co-adapted gene sets, as well as the mode in most cases geographical, rather than being maintained within popula- of limiting the production of unfit recombinant forms, remains a tions, and effects on local adaptation are cumulative. In contrast, super- substantial challenge7–10. -

Modulating Antiviral Response Via CRISPR–Cas Systems
viruses Review Immunity and Viral Infections: Modulating Antiviral Response via CRISPR–Cas Systems Sergey Brezgin 1,2,3,† , Anastasiya Kostyusheva 1,†, Ekaterina Bayurova 4 , Elena Volchkova 5, Vladimir Gegechkori 6 , Ilya Gordeychuk 4,7, Dieter Glebe 8 , Dmitry Kostyushev 1,3,*,‡ and Vladimir Chulanov 1,3,5,‡ 1 National Medical Research Center of Tuberculosis and Infectious Diseases, Ministry of Health, 127994 Moscow, Russia; [email protected] (S.B.); [email protected] (A.K.); [email protected] (V.C.) 2 Institute of Immunology, Federal Medical Biological Agency, 115522 Moscow, Russia 3 Scientific Center for Genetics and Life Sciences, Division of Biotechnology, Sirius University of Science and Technology, 354340 Sochi, Russia 4 Chumakov Federal Scientific Center for Research and Development of Immune-and-Biological Products of Russian Academy of Sciences, 108819 Moscow, Russia; [email protected] (E.B.); [email protected] (I.G.) 5 Department of Infectious Diseases, Sechenov University, 119991 Moscow, Russia; [email protected] 6 Department of Pharmaceutical and Toxicological Chemistry, Sechenov University, 119991 Moscow, Russia; [email protected] 7 Department of Organization and Technology of Immunobiological Drugs, Sechenov University, 119991 Moscow, Russia 8 National Reference Center for Hepatitis B Viruses and Hepatitis D Viruses, Institute of Medical Virology, Justus Liebig University of Giessen, 35392 Giessen, Germany; [email protected] * Correspondence: [email protected] † Co-first authors. Citation: Brezgin, S.; Kostyusheva, ‡ Co-senior authors. A.; Bayurova, E.; Volchkova, E.; Gegechkori, V.; Gordeychuk, I.; Glebe, Abstract: Viral infections cause a variety of acute and chronic human diseases, sometimes resulting D.; Kostyushev, D.; Chulanov, V. Immunity and Viral Infections: in small local outbreaks, or in some cases spreading across the globe and leading to global pandemics. -

Identification of Potential Key Genes and Pathway Linked with Sporadic Creutzfeldt-Jakob Disease Based on Integrated Bioinformatics Analyses
medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Identification of potential key genes and pathway linked with sporadic Creutzfeldt-Jakob disease based on integrated bioinformatics analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Abstract Sporadic Creutzfeldt-Jakob disease (sCJD) is neurodegenerative disease also called prion disease linked with poor prognosis. The aim of the current study was to illuminate the underlying molecular mechanisms of sCJD. The mRNA microarray dataset GSE124571 was downloaded from the Gene Expression Omnibus database. Differentially expressed genes (DEGs) were screened. -
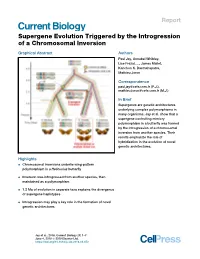
Supergene Evolution Triggered by the Introgression of a Chromosomal Inversion
Report Supergene Evolution Triggered by the Introgression of a Chromosomal Inversion Graphical Abstract Authors Paul Jay, Annabel Whibley, Lise Frezal, ..., James Mallet, Kanchon K. Dasmahapatra, Mathieu Joron Correspondence [email protected] (P.J.), [email protected] (M.J.) In Brief Supergenes are genetic architectures underlying complex polymorphisms in many organisms. Jay et al. show that a supergene controlling mimicry polymorphism in a butterfly was formed by the introgression of a chromosomal inversion from another species. Their results emphasize the role of hybridization in the evolution of novel genetic architectures. Highlights d Chromosomal inversions underlie wing-pattern polymorphism in a Heliconius butterfly d Inversion was introgressed from another species, then maintained as a polymorphism d 1.3 Ma of evolution in separate taxa explains the divergence of supergene haplotypes d Introgression may play a key role in the formation of novel genetic architectures Jay et al., 2018, Current Biology 28, 1–7 June 4, 2018 ª 2018 Elsevier Ltd. https://doi.org/10.1016/j.cub.2018.04.072 Please cite this article in press as: Jay et al., Supergene Evolution Triggered by the Introgression of a Chromosomal Inversion, Current Biology (2018), https://doi.org/10.1016/j.cub.2018.04.072 Current Biology Report Supergene Evolution Triggered by the Introgression of a Chromosomal Inversion Paul Jay,1,* Annabel Whibley,2 Lise Frezal, 3 Marı´aA´ ngeles Rodrı´guez de Cara,1 Reuben W. Nowell,4 James Mallet,5 Kanchon K. Dasmahapatra,6 and -

"Description"" ""Generatio"
TABLE S4 "ID ""Description"" ""GeneRatio"" ""BgRatio"" ""pvalue"" ""p.adjust"" ""qvalue"" ""geneID"" ""Count""" "GO:0003712 ""GO:0003712"" ""transcription coregulator activity"" ""84/1859"" ""454/22744"" 9.49597175224444e-13 9.80933882006851e-10 8.20651874588704e-10 ""Ncoa2/Zfp451/Dhx9/Hnrnpu/Cited2/Ncoa7/Ccar1/ Sirt1/Arid5b/Sirt6/Med1/Rara/Atxn7l3/Ddx5/Wbp2/Hdac9/Zmynd11/Cdyl/ Mier3/Sfmbt1/Gata4/Med4/Basp1/Zfpm2/Zhx2/Ddx17/Mkl2/Hes1/Nrip1/Usp16/ Elob/Rrp1b/Rxrb/Kat2b/Mta3/Hsbp1l1/Tle4/Sfr1/Eid1/Cops2/Sox12/Raly/ Ncoa6/Rbm39/Lpin3/Skil/Jade1/Maml3/Supt20/Med12l/Hdgf/Glmp/Nfib/Jun/ Pex14/Rere/Psmd9/Ncor2/Trim24/Ruvbl1/Rybp/Bhlhe40/Atf7ip/Ube3a/Mef2a/ Nrg1/Rbpms/Cnot7/Sin3b/Pou4f2/Pkn1/Cdyl2/Taf5l/Irf2bp2/Birc2/Yap1/ Skor1/Tfdp2/Rad54l2/Ctnnb1/Limd1/Med14/Rap2c/Tbl1x"" 84" "GO:0003779 ""GO:0003779"" ""actin binding"" ""65/1859"" ""414/22744"" 2.57546466939442e-07 8.86818334494813e-05 7.41914559148359e-05 ""Actr3/Cxcr4/Hnrnpu/Enah/Utrn/Epb41l2/Marcks/Ctnna3/Eef2/Pawr/ Ccdc88a/Anxa6/Gas7/Lasp1/Tns4/Syne2/Sipa1l1/Syne3/Phactr1/Enc1/Pxk/ Vcl/Ang/Myo10/Mtss1/Triobp/Mkl2/Afdn/Daam2/Svil/Ctnna1/Synpo/Myo5b/ Nrap/Ablim1/Shtn1/Fmnl2/Itprid2/Ino80/Pfn2/Myoz2/Pdlim5/Cap1/Macf1/ Epb41/Wasf2/Myom3/Ywhah/Coro1c/Ssh1/Hip1/Ppp1r9a/Wasl/Ctnna2/Mical3/ Eps8/Tlnrd1/Myom2/Klhl2/Sntb2/Spire2/Coro2b/Clasp2/Hdac6/Diaph2"" 65" "GO:0046332 ""GO:0046332"" ""SMAD binding"" ""22/1859"" ""84/22744"" 6.87941766027971e-07 0.000177660961076723 0.000148631628923412 ""Bmpr2/Cited2/Usp15/Ddx5/Axin2/Ppm1a/Yy1/Gata4/Tgif1/Ldlrad4/Smad7/ Acvr2a/Pmepa1/Skil/Trim33/Jun/Mef2a/Ipo7/Skor1/Rnf111/Tcf12/Ctnnb1"" -

How Genome-Wide Association Studies on Educational Attainment Reify Eugenic Ideologies
How Genome-Wide Association Studies on Educational Attainment Reify Eugenic Ideologies Dawn Warfield* Mentor: Dr. Emily Merchant Science and Technology Studies, UC Davis Abstract Genomic science and data are becoming ubiquitously entwined with how we describe the human condition in its physical, psychological, and social expression. Behavioral heredity research delves into genetic data science to reveal possible conditions, correlates, and determinants for particular behaviors and aptitudes that shape our individual and collective human experience. This article surveys the history, research culture, and methods of behavioral heredity science to investigate the techno-social interaction and motivations for recent genome-wide association studies (GWAS) performed to identify genomic biomarkers associated with educational attainment. Introduction The knowledge regime of behavioral heredity began in the late 19th century with Mendelian inference and extrapolated findings from animal breeding. As the technology of molecular biology enabled new understandings about the relationship between the heritability of traits and medical conditions through genes, social scientists began to explore genetics as a means to biologically explain human behavior and socioeconomic potential. In the most recent iteration of behavioral heredity, sociogenomic researchers now have access to both genomic and empirical survey data derived from longitudinal studies with broadening population samples, engendering new possibilities for finding associations. A Historical Perspective on Sociogenomics The longer history of sociogenomics begins with Francis Galton, who coined the word “eugenics,” literally meaning “well borne” in Greek, in 1883 (Müller-Wille and Rheinberger). Galton’s belief that human society could be perfected through selective breeding and social divestment of those considered to be a threat to the gene pool proved popular among scientists at the turn of the twentieth century.