Genome-Wide Meta-Analysis Unravels Interactions Between Magnesium Homeostasis and Metabolic Phenotypes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Data
Figure 2S 4 7 A - C 080125 CSCs 080418 CSCs - + IFN-a 48 h + IFN-a 48 h + IFN-a 72 h 6 + IFN-a 72 h 3 5 MRFI 4 2 3 2 1 1 0 0 MHC I MHC II MICA MICB ULBP-1 ULBP-2 ULBP-3 ULBP-4 MHC I MHC II MICA MICB ULBP-1 ULBP-2 ULBP-3 ULBP-4 7 B 13 080125 FBS - D 080418 FBS - + IFN-a 48 h 12 + IFN-a 48 h + IFN-a 72 h + IFN-a 72 h 6 080125 FBS 11 10 5 9 8 4 7 6 3 MRFI 5 4 2 3 2 1 1 0 0 MHC I MHC II MICA MICB ULBP-1 ULBP-2 ULBP-3 ULBP-4 MHC I MHC II MICA MICB ULBP-1 ULBP-2 ULBP-3 ULBP-4 Molecule Molecule FIGURE 4S FIGURE 5S Panel A Panel B FIGURE 6S A B C D Supplemental Results Table 1S. Modulation by IFN-α of APM in GBM CSC and FBS tumor cell lines. Molecule * Cell line IFN-α‡ HLA β2-m# HLA LMP TAP1 TAP2 class II A A HC§ 2 7 10 080125 CSCs - 1∞ (1) 3 (65) 2 (91) 1 (2) 6 (47) 2 (61) 1 (3) 1 (2) 1 (3) + 2 (81) 11 (80) 13 (99) 1 (3) 8 (88) 4 (91) 1 (2) 1 (3) 2 (68) 080125 FBS - 2 (81) 4 (63) 4 (83) 1 (3) 6 (80) 3 (67) 2 (86) 1 (3) 2 (75) + 2 (99) 14 (90) 7 (97) 5 (75) 7 (100) 6 (98) 2 (90) 1 (4) 3 (87) 080418 CSCs - 2 (51) 1 (1) 1 (3) 2 (47) 2 (83) 2 (54) 1 (4) 1 (2) 1 (3) + 2 (81) 3 (76) 5 (75) 2 (50) 2 (83) 3 (71) 1 (3) 2 (87) 1 (2) 080418 FBS - 1 (3) 3 (70) 2 (88) 1 (4) 3 (87) 2 (76) 1 (3) 1 (3) 1 (2) + 2 (78) 7 (98) 5 (99) 2 (94) 5 (100) 3 (100) 1 (4) 2 (100) 1 (2) 070104 CSCs - 1 (2) 1 (3) 1 (3) 2 (78) 1 (3) 1 (2) 1 (3) 1 (3) 1 (2) + 2 (98) 8 (100) 10 (88) 4 (89) 3 (98) 3 (94) 1 (4) 2 (86) 2 (79) * expression of APM molecules was evaluated by intracellular staining and cytofluorimetric analysis; ‡ cells were treatead or not (+/-) for 72 h with 1000 IU/ml of IFN-α; # β-2 microglobulin; § β-2 microglobulin-free HLA-A heavy chain; ∞ values are indicated as ratio between the mean of fluorescence intensity of cells stained with the selected mAb and that of the negative control; bold values indicate significant MRFI (≥ 2). -

In Silico Prediction of High-Resolution Hi-C Interaction Matrices
ARTICLE https://doi.org/10.1038/s41467-019-13423-8 OPEN In silico prediction of high-resolution Hi-C interaction matrices Shilu Zhang1, Deborah Chasman 1, Sara Knaack1 & Sushmita Roy1,2* The three-dimensional (3D) organization of the genome plays an important role in gene regulation bringing distal sequence elements in 3D proximity to genes hundreds of kilobases away. Hi-C is a powerful genome-wide technique to study 3D genome organization. Owing to 1234567890():,; experimental costs, high resolution Hi-C datasets are limited to a few cell lines. Computa- tional prediction of Hi-C counts can offer a scalable and inexpensive approach to examine 3D genome organization across multiple cellular contexts. Here we present HiC-Reg, an approach to predict contact counts from one-dimensional regulatory signals. HiC-Reg pre- dictions identify topologically associating domains and significant interactions that are enri- ched for CCCTC-binding factor (CTCF) bidirectional motifs and interactions identified from complementary sources. CTCF and chromatin marks, especially repressive and elongation marks, are most important for HiC-Reg’s predictive performance. Taken together, HiC-Reg provides a powerful framework to generate high-resolution profiles of contact counts that can be used to study individual locus level interactions and higher-order organizational units of the genome. 1 Wisconsin Institute for Discovery, 330 North Orchard Street, Madison, WI 53715, USA. 2 Department of Biostatistics and Medical Informatics, University of Wisconsin-Madison, Madison, WI 53715, USA. *email: [email protected] NATURE COMMUNICATIONS | (2019) 10:5449 | https://doi.org/10.1038/s41467-019-13423-8 | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | https://doi.org/10.1038/s41467-019-13423-8 he three-dimensional (3D) organization of the genome has Results Temerged as an important component of the gene regulation HiC-Reg for predicting contact count using Random Forests. -

Multiple Activities of Arl1 Gtpase in the Trans-Golgi Network Chia-Jung Yu1,2 and Fang-Jen S
© 2017. Published by The Company of Biologists Ltd | Journal of Cell Science (2017) 130, 1691-1699 doi:10.1242/jcs.201319 COMMENTARY Multiple activities of Arl1 GTPase in the trans-Golgi network Chia-Jung Yu1,2 and Fang-Jen S. Lee3,4,* ABSTRACT typical features of an Arf-family GTPase, including an amphipathic ADP-ribosylation factors (Arfs) and ADP-ribosylation factor-like N-terminal helix and a consensus site for N-myristoylation (Lu et al., proteins (Arls) are highly conserved small GTPases that function 2001; Price et al., 2005). In yeast, recruitment of Arl1 to the Golgi as main regulators of vesicular trafficking and cytoskeletal complex requires a second Arf-like GTPase, Arl3 (Behnia et al., reorganization. Arl1, the first identified member of the large Arl family, 2004; Setty et al., 2003). Yeast Arl3 lacks a myristoylation site and is an important regulator of Golgi complex structure and function in is, instead, N-terminally acetylated; this modification is required for organisms ranging from yeast to mammals. Together with its effectors, its recruitment to the Golgi complex by Sys1. In mammalian cells, Arl1 has been shown to be involved in several cellular processes, ADP-ribosylation-factor-related protein 1 (Arfrp1), a mammalian including endosomal trans-Golgi network and secretory trafficking, lipid ortholog of yeast Arl3, plays a pivotal role in the recruitment of Arl1 droplet and salivary granule formation, innate immunity and neuronal to the trans-Golgi network (TGN) (Behnia et al., 2004; Panic et al., development, stress tolerance, as well as the response of the unfolded 2003b; Setty et al., 2003; Zahn et al., 2006). -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Structural Basis for Disruption of Claudin Assembly in Tight Junctions
www.nature.com/scientificreports OPEN Structural basis for disruption of claudin assembly in tight junctions by an enterotoxin Received: 11 May 2016 Takehiro Shinoda1,2, Naoko Shinya1,2, Kaori Ito1,2, Noboru Ohsawa1,2, Takaho Terada1,3, Accepted: 01 September 2016 Kunio Hirata4, Yoshiaki Kawano4, Masaki Yamamoto4, Tomomi Kimura-Someya1,2, Published: 20 September 2016 Shigeyuki Yokoyama1,3 & Mikako Shirouzu1,2 The food-poisoning bacterium Clostridium perfringens produces an enterotoxin (~35 kDa) that specifically targets human claudin-4, among the 26 human claudin proteins, and causes diarrhea by fluid accumulation in the intestinal cavity. The C-terminal domain of the Clostridium perfringens enterotoxin (C-CPE, ~15 kDa) binds tightly to claudin-4, and disrupts the intestinal tight junction barriers. In this study, we determined the 3.5-Å resolution crystal structure of the cell-free synthesized human claudin- 4•C-CPE complex, which is significantly different from the structure of the off-target complex of an engineered C-CPE with mouse claudin-19. The claudin-4•C-CPE complex structure demonstrated the mechanism underlying claudin assembly disruption. A comparison of the present C-CPE-bound structure of claudin-4 with the enterotoxin-free claudin-15 structure revealed sophisticated C-CPE- induced conformation changes of the extracellular segments, induced on the foundation of the rigid four-transmembrane-helix bundle structure. These conformation changes provide a mechanistic model for the disruption of the lateral assembly of claudin molecules. Furthermore, the present novel structural mechanism for selecting a specific member of the claudin family can be used as the foundation to develop novel medically important technologies to selectively regulate the tight junctions formed by claudin family members in different organs. -

Extrinsic Regulators of Mrna Translation in Developing Brain: Story of Wnts
cells Article Extrinsic Regulators of mRNA Translation in Developing Brain: Story of WNTs Yongkyu Park * , Midori Lofton , Diana Li and Mladen-Roko Rasin * Department of Neuroscience and Cell Biology, Robert Wood Johnson Medical School, Rutgers University, Piscataway, NJ 08854, USA; [email protected] (M.L.); [email protected] (D.L.) * Correspondence: [email protected] (Y.P.); [email protected] (M.-R.R.) Abstract: Extrinsic molecules such as morphogens can regulate timed mRNA translation events in developing neurons. In particular, Wingless-type MMTV integration site family, member 3 (Wnt3), was shown to regulate the translation of Foxp2 mRNA encoding a Forkhead transcription factor P2 in the neocortex. However, the Wnt receptor that possibly mediates these translation events remains unknown. Here, we report Frizzled member 7 (Fzd7) as the Wnt3 receptor that lays downstream in Wnt3-regulated mRNA translation. Fzd7 proteins co-localize with Wnt3 ligands in developing neo- cortices. In addition, the Fzd7 proteins overlap in layer-specific neuronal subpopulations expressing different transcription factors, Foxp1 and Foxp2. When Fzd7 was silenced, we found decreased Foxp2 protein expression and increased Foxp1 protein expression, respectively. The Fzd7 silencing also dis- rupted the migration of neocortical glutamatergic neurons. In contrast, Fzd7 overexpression reversed the pattern of migratory defects and Foxp protein expression that we found in the Fzd7 silencing. We further discovered that Fzd7 is required for Wnt3-induced Foxp2 mRNA translation. Surprisingly, we also determined that the Fzd7 suppression of Foxp1 protein expression is not Wnt3 dependent. In conclusion, it is exhibited that the interaction between Wnt3 and Fzd7 regulates neuronal identity and the Fzd7 receptor functions as a downstream factor in ligand Wnt3 signaling for mRNA translation. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Oral Presentations
Journal of Inherited Metabolic Disease (2018) 41 (Suppl 1):S37–S219 https://doi.org/10.1007/s10545-018-0233-9 ABSTRACTS Oral Presentations PARALLEL SESSION 1A: Clycosylation and cardohydrate disorders O-002 Link between glycemia and hyperlipidemia in Glycogen Storage O-001 Disease type Ia Hoogerland J A1, Hijmans B S1, Peeks F1, Kooijman S3, 4, Bos T2, Fertility in classical galactosaemia, N-glycan, hormonal and inflam- Bleeker A1, Van Dijk T H2, Wolters H1, Havinga R1,PronkACM3, 4, matory gene expression interactions Rensen P C N3, 4,MithieuxG5, 6, Rajas F5, 6, Kuipers F1, 2,DerksTGJ1, Reijngoud D1,OosterveerMH1 Colhoun H O1,Rubio-GozalboME2,BoschAM3, Knerr I4,DawsonC5, Brady J J6,GalliganM8,StepienKM9, O'Flaherty R O7,MossC10, 1Dep Pediatrics, CLDM, Univ of Groningen, Groningen, Barker P11, Fitzgibbon M C6, Doran P8,TreacyEP1, 4, 9 Netherlands, 2Lab Med, CLDM, Univ of Groningen, Groningen, Netherlands, 3Dep of Med, Div of Endocrinology, LUMC, Leiden, 1Dept Paediatrics, Trinity College Dublin, Dublin, Ireland, 2Dept Paeds and Netherlands, 4Einthoven Lab Exp Vasc Med, LUMC, Leiden, Clin Genetics, UMC, Maastricht, Netherlands, 3Dept Paediatrics, AMC, Netherlands, 5Institut Nat Sante et Recherche Med, Lyon, Amsterdam, Netherlands, 4NCIMD, TSCUH, Dublin, Ireland, 5Dept France, 6Univ Lyon 1, Villeurbanne, France Endocrinology, NHS Foundation Trust, Birmingham, United Kingdom, 6Dept Clin Biochem, MMUH, Dublin, Ireland, 7NIBRT Glycoscience, Background: Glycogen Storage Disease type Ia (GSD Ia) is an NIBRT, Dublin, Ireland, 8UCDCRC,UCD,Dublin,Ireland,9NCIMD, inborn error of glucose metabolism characterized by fasting hypo- MMUH, Dublin, Ireland, 10Conway Institute, UCD, Dublin, Ireland, glycemia, hyperlipidemia and fatty liver disease. We have previ- 11CBAL, NHS Foundation, Cambridge, United Kingdom ously reported considerable heterogeneity in circulating triglycer- ide levels between individual GSD Ia patients, a phenomenon that Background: Classical Galactosaemia (CG) is caused by deficiency of is poorly understood. -

(Numbl) Downregulation Increases Tumorigenicity, Cancer Stem Cell-Like Properties and Resistance to Chemotherapy
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 7, No. 39 Research Paper Numb-like (NumbL) downregulation increases tumorigenicity, cancer stem cell-like properties and resistance to chemotherapy José M. García-Heredia1,2, Eva M. Verdugo Sivianes1, Antonio Lucena-Cacace1, Sonia Molina-Pinelo1,3, Amancio Carnero1 1Instituto de Biomedicina de Sevilla (IBIS), Hospital Universitario Virgen del Rocio, Universidad de Sevilla, Consejo Superior de Investigaciones Cientificas, Seville, Spain 2Department of Vegetal Biochemistry and Molecular Biology, University of Seville, Seville, Spain 3Present address: Instituto de Investigación Hospital 12 de Octubre, Madrid, Spain Correspondence to: Amancio Carnero, email: [email protected] Keywords: NumbL, Notch, cancer stem cells, tumor suppressor, tumorigenicity Received: April 22, 2016 Accepted: August 12, 2016 Published: August 23, 2016 ABSTRACT NumbL, or Numb-like, is a close homologue of Numb, and is part of an evolutionary conserved protein family implicated in some important cellular processes. Numb is a protein involved in cell development, in cell adhesion and migration, in asymmetric cell division, and in targeting proteins for endocytosis and ubiquitination. NumbL exhibits some overlapping functions with Numb, but its role in tumorigenesis is not fully known. Here we showed that the downregulation of NumbL alone is sufficient to increase NICD nuclear translocation and induce Notch pathway activation. Furthermore, NumbL downregulation increases epithelial-mesenchymal transition (EMT) and cancer stem cell (CSC)-related gene transcripts and CSC-like phenotypes, including an increase in the CSC-like pool. These data suggest that NumbL can act independently as a tumor suppressor gene. Furthermore, an absence of NumbL induces chemoresistance in tumor cells. An analysis of human tumors indicates that NumbL is downregulated in a variable percentage of human tumors, with lower levels of this gene correlated with worse prognosis in colon, breast and lung tumors. -

OSTF1 (AT9G4): Sc-517418
SAN TA C RUZ BI OTEC HNOL OG Y, INC . OSTF1 (AT9G4): sc-517418 BACKGROUND PRODUCT OSTF1 (osteoclast-stimulating factor 1) is a 214 amino acid cytoplasmic pro - Each vial contains 100 µg IgG 2a kappa light chain in 1.0 ml of PBS with tein that contains 3 ANK repeats and one SH3 domain. The SH3 domain < 0.1% sodium azide and 0.1% gelatin. mediates interaction of OSTF1 with SMN. In addition to SMN, OSTF1 inter - acts with Src and FAS-L. In osteoclasts, OSTF1 is thought to induce bone APPLICATIONS resorption most likely through a signaling cascade that results in the secre - OSTF1 (AT9G4) is recommended for detection of OSTF1 of mouse, rat and tion of factors that enhance osteoclast formation and activity. The gene that human origin by Western Blotting (starting dilution 1:200, dilution range encdoes OSTF1 consists of almost 59,000 bases and maps to human chro mo - 1:100-1:1000), immunoprecipitation [1-2 µg per 100-500 µg of total protein some 9q21.13. Housing over 900 genes, chromosome 9 comprises nearly 4% (1 ml of cell lysate)], immunofluorescence (starting dilution 1:50, dilution of the human genome. Hereditary hemorrhagic telangiectasia, which is char - range 1:50-1:500), flow cytometry (1 µg per 1 x 10 6 cells) and solid phase acterized by harmful vascular defects, and familial dysautonomia, are both ELISA (starting dilution 1:30, dilution range 1:30-1:3000). associated with chromosome 9. Notably, chromosome 9 encompasses the largest interferon family gene cluster. Suitable for use as control antibody for OSTF1 siRNA (h): sc-92800, OSTF1 siRNA (m): sc-151336, OSTF1 shRNA Plasmid (h): sc-92800-SH, OSTF1 REFERENCES shRNA Plasmid (m): sc-151336-SH, OSTF1 shRNA (h) Lentiviral Particles: sc-92800-V and OSTF1 shRNA (m) Lentiviral Particles: sc-151336-V. -

Massive Parallel Sequencing As a New Diagnostic Approach For
Chaiyasap et al. BMC Medical Genetics (2017) 18:102 DOI 10.1186/s12881-017-0464-x RESEARCH ARTICLE Open Access Massive parallel sequencing as a new diagnostic approach for phenylketonuria and tetrahydrobiopterin-deficiency in Thailand Pongsathorn Chaiyasap1, Chupong Ittiwut1,2, Chalurmpon Srichomthong1,2, Apiruk Sangsin3, Kanya Suphapeetiporn1,2,4* and Vorasuk Shotelersuk1,2 Abstract Background: Hyperphenylalaninemia (HPA) can be classified into phenylketonuria (PKU) which is caused by mutations in the phenylalanine hydroxylase (PAH) gene, and BH4 deficiency caused by alterations in genes involved in tetrahydrobiopterin (BH4) biosynthesis pathway. Dietary restriction of phenylalanine is considered to be the main treatment of PKU to prevent irreversible intellectual disability. However, the same dietary intervention in BH4 deficiency patients is not as effective, as BH4 is also a cofactor in many neurotransmitter syntheses. Method: We utilized next generation sequencing (NGS) technique to investigate four unrelated Thai patients with hyperphenylalaninemia. Result: We successfully identified all eight mutant alleles in PKU or BH4-deficiency associated genes including three novel mutations, one in PAH and two in PTS, thus giving a definite diagnosis to these patients. Appropriate management can then be provided. Conclusion: This study identified three novel mutations in either the PAH or PTS gene and supported the use of NGS as an alternative molecular genetic approach for definite diagnosis of hyperphenylalaninemia, thus leading to proper management of these patients in Thailand. Keywords: Next generation sequencing, Exome, Hyperphenylalaninemia, Phenylketonuria, Tetrahydrobiopterin deficiency, Newborn screening Background 120 μmol/l (2 mg/dl), the individual is considered to be Phenylketonuria (PKU) is an autosomal recessive metabolic hyperphenylalaninemia (HPA) and needs further diagnosis disorder, characterized by progressive intellectual disability, [4]. -
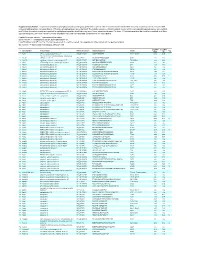
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding