Role of Heme Oxygenase As a Modulator of Heme-Mediated Pathways
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Regulation of Ribulose-1, 5-Bisphosphate Carboxylase
Plant Physiol. (1993) 102: 21-28 Regulation of Ribulose- 1,5-Bisphosphate Carboxylase/Oxygenase Activity in Response to Reduced Light lntensity in C4 Plants' Rowan F. Sage* and Jeffrey R. Seemann Department of Botany, University of Georgia, Athens, Ceorgia 30602 (R.F.S.); and Department of Biochemistry, University of Nevada, Reno, Nevada 89557 (J.R.S.) tion of Rubisco activity by reversible carbamylation occurs in lhe light-dependent regulation of ribulose-1,5-bisphosphate response to changes in light intensity as well as the concen- carboxylase/oxygenase (Rubisco) activity was studied in 16 species tration of C02and 02,whereas inhibition of Rubisco activity of C, plants representing all three biochemical subtypes and a by CAlP occurs only in response to varying PPFD (Sharkey variety of taxonomic groups. Rubisco regulation was assessed by et al., 1986; Sage et al., 1990; Seemann et al., 1990). At measuring (a) the ratio of initial to total Rubisco activity, which physiological levels of COz in C3 plants (5-10 PM), full reflects primarily the carbamylation state of the enzyme, and (b) carbamylation of Rubisco requires Rubisco activase (Salvucci, total Rubisco activity per mo1 of Rubisco catalytic sites, which 1989; Portis, 1990). In the absence of Rubisco activase, only declines when 2-carboxyarabinitol I-phosphate (CAlP) binds to carbamylated Rubisco. In all species examined, the activity ratio of 20 to 40% of Rubisco catalytic sites are carbamylated under Rubisco declined with a reduction in light intensity, although sub- physiological conditions, leading to a significant inhibition of stantial variation was apparent between species in the degree of photosynthesis (Salvucci, 1989; Portis, 1990). -
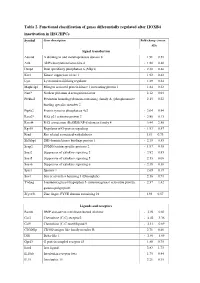
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Chapter 5 High-Pressure Stopped-Flow Kinetic Studies of Nnos
Probing the dynamics and conformational landscape of neuronal nitric oxide synthase A thesis submitted to the University of Manchester for the degree of Doctor of Philosophy in the Faculty of Life Sciences 2013 Anna Sobolewska-Stawiarz Contents Contents ...................................................................................................................... 2 List of Figures ............................................................................................................. 6 List of Tables ............................................................................................................ 10 Abstract ..................................................................................................................... 12 Declaration ................................................................................................................ 13 Copyright Statement ................................................................................................ 13 Acknowledgements ................................................................................................... 14 List of Abbreviations ............................................................................................... 15 List of Amino Acids Abbreviations ........................................................................ 17 CHAPTER 1. INTRODUCTION ........................................................................... 18 1.1 Nitric oxide synthase ......................................................................................... -

Carbon Monoxide Prevents TNF-Α-Induced Enos Downregulation by Inhibiting NF-Κb-Responsive Mir-155-5P Biogenesis
OPEN Experimental & Molecular Medicine (2017) 49, e403; doi:10.1038/emm.2017.193 Official journal of the Korean Society for Biochemistry and Molecular Biology www.nature.com/emm ORIGINAL ARTICLE Carbon monoxide prevents TNF-α-induced eNOS downregulation by inhibiting NF-κB-responsive miR-155-5p biogenesis Seunghwan Choi1, Joohwan Kim1, Ji-Hee Kim1, Dong-Keon Lee1, Wonjin Park1, Minsik Park1, Suji Kim1, Jong Yun Hwang2, Moo-Ho Won3, Yoon Kyung Choi1,4, Sungwoo Ryoo5, Kwon-Soo Ha1, Young-Guen Kwon6 and Young-Myeong Kim1 Heme oxygenase-1-derived carbon monoxide prevents inflammatory vascular disorders. To date, there is no clear evidence that HO-1/CO prevents endothelial dysfunction associated with the downregulation of endothelial NO synthesis in human endothelial cells stimulated with TNF-α. Here, we found that the CO-releasing compound CORM-2 prevented TNF-α-mediated decreases in eNOS expression and NO/cGMP production, without affecting eNOS promoter activity, by maintaining the functional activity of the eNOS mRNA 3′-untranslated region. By contrast, CORM-2 inhibited MIR155HG expression and miR-155-5p biogenesis in TNF-α-stimulated endothelial cells, resulting in recovery of the 3′-UTR activity of eNOS mRNA, a target of miR-155-5p. The beneficial effect of CORM-2 was blocked by an NF-κB inhibitor, a miR-155-5p mimic, a HO-1 inhibitor and siRNA against HO-1, indicating that CO rescues TNF-α-induced eNOS downregulation through NF-κB-responsive miR-155-5p expression via HO-1 induction; similar protective effects of ectopic HO-1 expression and bilirubin were observed in endothelial cells treated with TNF-α. -

Somatic Aging Pathways Regulate Reproductive Plasticity in Caenorhabditis Elegans Maria C Ow, Alexandra M Nichitean, Sarah E Hall*
RESEARCH ARTICLE Somatic aging pathways regulate reproductive plasticity in Caenorhabditis elegans Maria C Ow, Alexandra M Nichitean, Sarah E Hall* Department of Biology, Syracuse University, Syracuse, United States Abstract In animals, early-life stress can result in programmed changes in gene expression that can affect their adult phenotype. In C. elegans nematodes, starvation during the first larval stage promotes entry into a stress-resistant dauer stage until environmental conditions improve. Adults that have experienced dauer (postdauers) retain a memory of early-life starvation that results in gene expression changes and reduced fecundity. Here, we show that the endocrine pathways attributed to the regulation of somatic aging in C. elegans adults lacking a functional germline also regulate the reproductive phenotypes of postdauer adults that experienced early-life starvation. We demonstrate that postdauer adults reallocate fat to benefit progeny at the expense of the parental somatic fat reservoir and exhibit increased longevity compared to controls. Our results also show that the modification of somatic fat stores due to parental starvation memory is inherited in the F1 generation and may be the result of crosstalk between somatic and reproductive tissues mediated by the germline nuclear RNAi pathway. Introduction Evidence indicating that experiences during early development affect behavior and physiology in a stress-specific manner later in life is abundant throughout the animal kingdom (Telang and Wells, *For correspondence: [email protected] 2004; Weaver et al., 2004; Binder et al., 2008; Pellegroms et al., 2009; van Abeelen et al., 2012; Zhao and Zhu, 2014; Canario et al., 2017; Dantzer et al., 2019; Vitikainen et al., 2019). -

A Three-Component Dicamba O-Demethylase from Pseudomonas Maltophilia, Strain DI-6 GENE ISOLATION, CHARACTERIZATION, and HETEROLOGOUS EXPRESSION*
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 280, No. 26, Issue of July 1, pp. 24759–24767, 2005 © 2005 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. A Three-component Dicamba O-Demethylase from Pseudomonas maltophilia, Strain DI-6 GENE ISOLATION, CHARACTERIZATION, AND HETEROLOGOUS EXPRESSION* Received for publication, January 18, 2005, and in revised form, March 16, 2005 Published, JBC Papers in Press, April 26, 2005, DOI 10.1074/jbc.M500597200 Patricia L. Herman‡, Mark Behrens, Sarbani Chakraborty, Brenda M. Chrastil, Joseph Barycki, and Donald P. Weeks§ From the Department of Biochemistry, University of Nebraska-Lincoln, Lincoln, Nebraska 65888-0664 Dicamba O-demethylase is a multicomponent enzyme The herbicide dicamba (2-methoxy-3,6-dichlorobenzoic acid) from Pseudomonas maltophilia, strain DI-6, that cata- has been used to effectively control broadleaf weeds in crops lyzes the conversion of the widely used herbicide di- such as corn and wheat for almost 40 years. Like a number of camba (2-methoxy-3,6-dichlorobenzoic acid) to DCSA other chlorinated organic compounds, dicamba does not persist (3,6-dichlorosalicylic acid). We recently described the in the soil because it is efficiently metabolized by a consortium biochemical characteristics of the three components of of soil bacteria under both aerobic and anaerobic conditions this enzyme (i.e. reductaseDIC, ferredoxinDIC, and oxy- (1–4). Studies with different soil types treated with dicamba genaseDIC) and classified the oxygenase component of have demonstrated that 3,6-dichlorosalicylic acid (DCSA),1 a dicamba O-demethylase as a member of the Rieske non- compound without herbicidal activity, is a major product of the heme iron family of oxygenases. -

Flavin-Containing Monooxygenases: Mutations, Disease and Drug Response Phillips, IR; Shephard, EA
Flavin-containing monooxygenases: mutations, disease and drug response Phillips, IR; Shephard, EA For additional information about this publication click this link. http://qmro.qmul.ac.uk/jspui/handle/123456789/1015 Information about this research object was correct at the time of download; we occasionally make corrections to records, please therefore check the published record when citing. For more information contact [email protected] Flavin-containing monooxygenases: mutations, disease and drug response Ian R. Phillips1 and Elizabeth A. Shephard2 1School of Biological and Chemical Sciences, Queen Mary, University of London, Mile End Road, London E1 4NS, UK 2Department of Biochemistry and Molecular Biology, University College London, Gower Street, London WC1E 6BT, UK Corresponding author: Shephard, E.A. ([email protected]). and, thus, contribute to drug development. This review Flavin-containing monooxygenases (FMOs) metabolize considers the role of FMOs and their genetic variants in numerous foreign chemicals, including drugs, pesticides disease and drug response. and dietary components and, thus, mediate interactions between humans and their chemical environment. We Mechanism and structure describe the mechanism of action of FMOs and insights For catalysis FMOs require flavin adenine dinucleotide gained from the structure of yeast FMO. We then (FAD) as a prosthetic group, NADPH as a cofactor and concentrate on the three FMOs (FMOs 1, 2 and 3) that are molecular oxygen as a cosubstrate [5,6]. In contrast to most important for metabolism of foreign chemicals in CYPs FMOs accept reducing equivalents directly from humans, focusing on the role of the FMOs and their genetic NADPH and, thus, do not require accessory proteins. -

Targeting Heme Oxygenase-1 in Cardiovascular and Kidney Disease
antioxidants Review Targeting Heme Oxygenase-1 in Cardiovascular and Kidney Disease Heather A. Drummond 1, Zachary L. Mitchell 1, Nader G. Abraham 2,3 and David E. Stec 1,* 1 Department of Physiology and Biophysics, Mississippi Center for Obesity Research, University of Mississippi Medical Center, Jackson, MI 39216, USA; [email protected] (H.A.D.); [email protected] (Z.L.M.) 2 Departments of Medicine and Pharmacology, New York Medical College, Vahalla, NY 10595, USA; [email protected] 3 Joan C. Edwards School of Medicine, Marshall University, Huntington, VA 25701, USA * Correspondence: [email protected]; Tel.: +1-601-815-1859; Fax: +1-601-984-1817 Received: 3 June 2019; Accepted: 15 June 2019; Published: 18 June 2019 Abstract: Heme oxygenase (HO) plays an important role in the cardiovascular system. It is involved in many physiological and pathophysiological processes in all organs of the cardiovascular system. From the regulation of blood pressure and blood flow to the adaptive response to end-organ injury, HO plays a critical role in the ability of the cardiovascular system to respond and adapt to changes in homeostasis. There have been great advances in our understanding of the role of HO in the regulation of blood pressure and target organ injury in the last decade. Results from these studies demonstrate that targeting of the HO system could provide novel therapeutic opportunities for the treatment of several cardiovascular and renal diseases. The goal of this review is to highlight the important role of HO in the regulation of cardiovascular and renal function and protection from disease and to highlight areas in which targeting of the HO system needs to be translated to help benefit patient populations. -

Nobel Lecture by Roger Y. Tsien
CONSTRUCTING AND EXPLOITING THE FLUORESCENT PROTEIN PAINTBOX Nobel Lecture, December 8, 2008 by Roger Y. Tsien Howard Hughes Medical Institute, University of California San Diego, 9500 Gilman Drive, La Jolla, CA 92093-0647, USA. MOTIVATION My first exposure to visibly fluorescent proteins (FPs) was near the end of my time as a faculty member at the University of California, Berkeley. Prof. Alexander Glazer, a friend and colleague there, was the world’s expert on phycobiliproteins, the brilliantly colored and intensely fluorescent proteins that serve as light-harvesting antennae for the photosynthetic apparatus of blue-green algae or cyanobacteria. One day, probably around 1987–88, Glazer told me that his lab had cloned the gene for one of the phycobilipro- teins. Furthermore, he said, the apoprotein produced from this gene became fluorescent when mixed with its chromophore, a small molecule cofactor that could be extracted from dried cyanobacteria under conditions that cleaved its bond to the phycobiliprotein. I remember becoming very excited about the prospect that an arbitrary protein could be fluorescently tagged in situ by genetically fusing it to the phycobiliprotein, then administering the chromophore, which I hoped would be able to cross membranes and get inside cells. Unfortunately, Glazer’s lab then found out that the spontane- ous reaction between the apoprotein and the chromophore produced the “wrong” product, whose fluorescence was red-shifted and five-fold lower than that of the native phycobiliprotein1–3. An enzyme from the cyanobacteria was required to insert the chromophore correctly into the apoprotein. This en- zyme was a heterodimer of two gene products, so at least three cyanobacterial genes would have to be introduced into any other organism, not counting any gene products needed to synthesize the chromophore4. -

Relating Metatranscriptomic Profiles to the Micropollutant
1 Relating Metatranscriptomic Profiles to the 2 Micropollutant Biotransformation Potential of 3 Complex Microbial Communities 4 5 Supporting Information 6 7 Stefan Achermann,1,2 Cresten B. Mansfeldt,1 Marcel Müller,1,3 David R. Johnson,1 Kathrin 8 Fenner*,1,2,4 9 1Eawag, Swiss Federal Institute of Aquatic Science and Technology, 8600 Dübendorf, 10 Switzerland. 2Institute of Biogeochemistry and Pollutant Dynamics, ETH Zürich, 8092 11 Zürich, Switzerland. 3Institute of Atmospheric and Climate Science, ETH Zürich, 8092 12 Zürich, Switzerland. 4Department of Chemistry, University of Zürich, 8057 Zürich, 13 Switzerland. 14 *Corresponding author (email: [email protected] ) 15 S.A and C.B.M contributed equally to this work. 16 17 18 19 20 21 This supporting information (SI) is organized in 4 sections (S1-S4) with a total of 10 pages and 22 comprises 7 figures (Figure S1-S7) and 4 tables (Table S1-S4). 23 24 25 S1 26 S1 Data normalization 27 28 29 30 Figure S1. Relative fractions of gene transcripts originating from eukaryotes and bacteria. 31 32 33 Table S1. Relative standard deviation (RSD) for commonly used reference genes across all 34 samples (n=12). EC number mean fraction bacteria (%) RSD (%) RSD bacteria (%) RSD eukaryotes (%) 2.7.7.6 (RNAP) 80 16 6 nda 5.99.1.2 (DNA topoisomerase) 90 11 9 nda 5.99.1.3 (DNA gyrase) 92 16 10 nda 1.2.1.12 (GAPDH) 37 39 6 32 35 and indicates not determined. 36 37 38 39 S2 40 S2 Nitrile hydration 41 42 43 44 Figure S2: Pearson correlation coefficients r for rate constants of bromoxynil and acetamiprid with 45 gene transcripts of ECs describing nucleophilic reactions of water with nitriles. -

Surprising Roles for Bilins in a Green Alga Jean-David Rochaix1 Departments of Molecular Biology and Plant Biology, University of Geneva,1211 Geneva, Switzerland
COMMENTARY COMMENTARY Surprising roles for bilins in a green alga Jean-David Rochaix1 Departments of Molecular Biology and Plant Biology, University of Geneva,1211 Geneva, Switzerland It is well established that the origin of plastids which serves as chromophore of phyto- can be traced to an endosymbiotic event in chromes (Fig. 1). An intriguing feature of which a free-living photosynthetic prokaryote all sequenced chlorophyte genomes is that, invaded a eukaryotic cell more than 1 billion although they lack phytochromes, their years ago. Most genes from the intruder genomes encode two HMOXs, HMOX1 were gradually transferred to the host nu- andHMOX2,andPCYA.InPNAS,Duanmu cleus whereas a small number of these genes et al. (6) investigate the role of these genes in were maintained in the plastid and gave the green alga Chlamydomonas reinhardtii rise to the plastid genome with its associated and made unexpected findings. protein synthesizing system. The products of Duanmu et al. first show that HMOX1, many of the genes transferred to the nucleus HMOX2, and PCYA are catalytically active were then retargeted to the plastid to keep it and produce bilins in vitro (6). They also functional. Altogether, approximately 3,000 demonstrate in a very elegant way that these nuclear genes in plants and algae encode proteins are functional in vivo by expressing plastid proteins, whereas chloroplast ge- a cyanobacteriochrome in the chloroplast Fig. 1. Tetrapyrrole biosynthetic pathways. The heme nomes contain between 100 and 120 genes of C. reinhardtii, where, remarkably, the and chlorophyll biosynthetic pathways diverge at pro- (1). A major challenge for eukaryotic pho- photoreceptor is assembled with bound toporphyrin IX (ProtoIX). -

Investigating the Role of Heme Oxygenase and Oxidative Stress in Oesophagogastric Cancer
Investigating the Role of Heme Oxygenase and Oxidative Stress in Oesophagogastric Cancer Oliver Hartwell Priest Department of Surgery and Cancer Imperial College London St Mary’s Campus London, UK A thesis submitted to Imperial College London for the degree of Doctorate of Philosophy Supervisors Professor George Hanna Department of Surgery and Cancer, Imperial College London St Mary’s Campus, London, UK Dr Nandor Marczin Department of Anaesthetics, Pain Medicine and Intensive Care Imperial College London, Chelsea and Westminster Campus, London, UK 2 ABSTRACT Background: Molecular mechanisms underlying gastric and oesophageal cancer include alterations in growth factors, cytokines and cell adhesion molecules. Heme oxygenase (HO) enzyme catalyses the degradation of heme and generates bilirubin and carbon monoxide that have antioxidant, anti-inflammatory and anti-apoptotic activities. HO enzyme is implicated in the biology of cancer by its effects on cell growth and resistance to apoptosis. The roles of HO-1 and HO-2 enzymes in cancer cell growth are poorly understood, with reports suggesting both anti-inflammatory, anti-tumour effects and tumour-protective mechanisms mediated by HO activity. The role of HO-2 in inflammation and cancer is largely unexplored. Further understanding the influence of the HO enzyme system may provide improved novel targets for oesophagogastric cancer therapy. Materials and Methods: The primary objective of the thesis was to characterise the role of the heme oxygenase pathway and modulation of HO activity in upper gastrointestinal cancer cell growth in vitro. Cell culture techniques included MTT growth assay, Western blotting protein analysis, pharmacological modulation of HO activity and targeted knockdown of HO mRNA.