Tissue-Specific Splicing of a Ubiquitously Expressed Transcription Factor Is Essential for Muscle Differentiation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Core Transcriptional Regulatory Circuitries in Cancer
Oncogene (2020) 39:6633–6646 https://doi.org/10.1038/s41388-020-01459-w REVIEW ARTICLE Core transcriptional regulatory circuitries in cancer 1 1,2,3 1 2 1,4,5 Ye Chen ● Liang Xu ● Ruby Yu-Tong Lin ● Markus Müschen ● H. Phillip Koeffler Received: 14 June 2020 / Revised: 30 August 2020 / Accepted: 4 September 2020 / Published online: 17 September 2020 © The Author(s) 2020. This article is published with open access Abstract Transcription factors (TFs) coordinate the on-and-off states of gene expression typically in a combinatorial fashion. Studies from embryonic stem cells and other cell types have revealed that a clique of self-regulated core TFs control cell identity and cell state. These core TFs form interconnected feed-forward transcriptional loops to establish and reinforce the cell-type- specific gene-expression program; the ensemble of core TFs and their regulatory loops constitutes core transcriptional regulatory circuitry (CRC). Here, we summarize recent progress in computational reconstitution and biologic exploration of CRCs across various human malignancies, and consolidate the strategy and methodology for CRC discovery. We also discuss the genetic basis and therapeutic vulnerability of CRC, and highlight new frontiers and future efforts for the study of CRC in cancer. Knowledge of CRC in cancer is fundamental to understanding cancer-specific transcriptional addiction, and should provide important insight to both pathobiology and therapeutics. 1234567890();,: 1234567890();,: Introduction genes. Till now, one critical goal in biology remains to understand the composition and hierarchy of transcriptional Transcriptional regulation is one of the fundamental mole- regulatory network in each specified cell type/lineage. -

Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin
cells Review Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin Agnieszka Bochy ´nska,Juliane Lüscher-Firzlaff and Bernhard Lüscher * ID Institute of Biochemistry and Molecular Biology, Medical School, RWTH Aachen University, Pauwelsstrasse 30, 52057 Aachen, Germany; [email protected] (A.B.); jluescher-fi[email protected] (J.L.-F.) * Correspondence: [email protected]; Tel.: +49-241-8088850; Fax: +49-241-8082427 Received: 18 January 2018; Accepted: 27 February 2018; Published: 2 March 2018 Abstract: Regulation of gene expression is achieved by sequence-specific transcriptional regulators, which convey the information that is contained in the sequence of DNA into RNA polymerase activity. This is achieved by the recruitment of transcriptional co-factors. One of the consequences of co-factor recruitment is the control of specific properties of nucleosomes, the basic units of chromatin, and their protein components, the core histones. The main principles are to regulate the position and the characteristics of nucleosomes. The latter includes modulating the composition of core histones and their variants that are integrated into nucleosomes, and the post-translational modification of these histones referred to as histone marks. One of these marks is the methylation of lysine 4 of the core histone H3 (H3K4). While mono-methylation of H3K4 (H3K4me1) is located preferentially at active enhancers, tri-methylation (H3K4me3) is a mark found at open and potentially active promoters. Thus, H3K4 methylation is typically associated with gene transcription. The class 2 lysine methyltransferases (KMTs) are the main enzymes that methylate H3K4. KMT2 enzymes function in complexes that contain a necessary core complex composed of WDR5, RBBP5, ASH2L, and DPY30, the so-called WRAD complex. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Dephosphorylation of HDAC4 by PP2A-Bδ Unravels a New Role For
Veloso et et al. al.CellCell Death Death and and Disease Disease (2019) 10:512 (2019) 10:512 Page 1 of 16 https://doi.org/10.1038/s41419-019-1743-6 Cell Death & Disease ARTICLE Open Access Dephosphorylation of HDAC4 by PP2A-Bδ unravels a new role for the HDAC4/MEF2 axis in myoblast fusion Alexandra Veloso1,2, Maud Martin1,2,3, Jonathan Bruyr1,2,TinaO’Grady1,2, Christophe Deroanne1,4, Denis Mottet1,2, Jean-Claude Twizere1,2, Thomas Cherrier1,2,5 and Franck Dequiedt1,2 Abstract Muscle formation is controlled by a number of key myogenic transcriptional regulators that govern stage-specific gene expression programs and act as terminal effectors of intracellular signaling pathways. To date, the role of phosphatases in the signaling cascades instructing muscle development remains poorly understood. Here, we show that a specific PP2A-B55δ holoenzyme is necessary for skeletal myogenesis. The primary role of PP2A-B55δ is to dephosphorylate histone deacetylase 4 (HDAC4) following myocyte differentiation and ensure repression of Myocyte enhancer factor 2D (MEF2D)-dependent gene expression programs during myogenic fusion. As a crucial HDAC4/MEF2D target gene that governs myocyte fusion, we identify ArgBP2, an upstream inhibitor of Abl, which itself is a repressor of CrkII signaling. Consequently, cells lacking PP2A-B55δ show upregulation of ArgBP2 and hyperactivation of CrkII downstream effectors, including Rac1 and FAK, precluding cytoskeletal and membrane rearrangements associated with myoblast fusion. Both in vitro and in zebrafish, loss-of-function of PP2A-B55δ severely impairs fusion of myocytes and formation of multinucleated muscle fibers, without affecting myoblast differentiation. Taken together, our results establish PP2A-B55δ as the first protein phosphatase to be involved in myoblast fusion and suggest that reversible phosphorylation of HDAC4 may coordinate differentiation and fusion events during myogenesis. -

Supplementary Material and Methods
Supplementary material and methods Generation of cultured human epidermal sheets Normal human epidermal keratinocytes were isolated from human breast skin. Keratinocytes were grown on a feeder layer of irradiated human fibroblasts pre-seeded at 4000 cells /cm² in keratinocyte culture medium (KCM) containing a mix of 3:1 DMEM and HAM’s F12 (Invitrogen, Carlsbad, USA), supplemented with 10% FCS, 10ng/ml epidermal growth factor (EGF; R&D systems, Minneapolis, MN, USA), 0.12 IU/ml insulin (Lilly, Saint- Cloud, France), 0.4 mg/ml hydrocortisone (UpJohn, St Quentin en Yvelelines, France) , 5 mg/ml triiodo-L- thyronine (Sigma, St Quentin Fallavier, France), 24.3 mg/ml adenine (Sigma), isoproterenol (Isuprel, Hospira France, Meudon, France) and antibiotics (20 mg/ml gentamicin (Phanpharma, Fougères, France), 100 IU/ml penicillin (Phanpharma), and 1 mg/ml amphotericin B (Phanpharma)). The medium was changed every two days. NHEK were then cultured over a period of 13 days according to the protocol currently used at the Bank of Tissues and Cells for the generation of clinical grade epidermal sheets used for the treatment of severe extended burns (Ref). When needed, cells were harvested with trypsin-EDTA 0.05% (Thermo Fisher Scientific, Waltham, MA, USA) and collected for analysis. Clonogenic assay Keratinocytes were seeded on a feeder layer of irradiated fibroblasts, at a clonal density of 10-20 cells/cm² and cultivated for 10 to 14 days. Three flasks per tested condition were fixed and colored in a single 30 mns step using rhodamine B (Sigma) diluted at 0.01 g/ml in 4% paraformaldehyde. In each tested condition, cells from 3 other flasks were numerated after detachment by trypsin treatment. -

Precision Medicine in Pediatric Oncology: Translating Genomic Discoveries Into Optimized Therapies Thai Hoa Tran1,2, Avanthi Tayi Shah3,4, and Mignon L
Published OnlineFirst June 9, 2017; DOI: 10.1158/1078-0432.CCR-16-0115 Review Clinical Cancer Research Precision Medicine in Pediatric Oncology: Translating Genomic Discoveries into Optimized Therapies Thai Hoa Tran1,2, Avanthi Tayi Shah3,4, and Mignon L. Loh3,4 Abstract Survival of children with cancers has dramatically improved aberrant activation of signaling pathways, and epigenetic modi- over the past several decades. This success has been achieved fiers that can be targeted by novel agents. Thus, the recently through improvement of combined modalities in treatment described genomic and epigenetic landscapes of many child- approaches, intensification of cytotoxic chemotherapy for hood cancers have expanded the paradigm of precision med- those with high-risk disease, and refinement of risk stratifica- icine in the hopes of improving outcomes while minimizing tion incorporating novel biologic markers in addition to tra- toxicities. In this review, we will discuss the biologic rationale ditional clinical and histologic features. Advances in cancer for molecularly targeted therapies in genomically defined sub- genomics have shed important mechanistic insights on disease sets of pediatric leukemias, solid tumors, and brain tumors. biology and have identified "driver" genomic alterations, Clin Cancer Res; 23(18); 5329–38. Ó2017 AACR. Introduction which may represent actionable therapeutic targets. In this review, we will discuss how genomic discoveries are being translated from Survival rates for children diagnosed with cancer have the bench into the clinic, resulting in the development of precision improved substantially over the past five decades. Today, long- medicine trials for specific subtypes of pediatric hematologic term survival is expected for approximately 80% of children malignancies, solid tumors, and brain tumors. -

Gene Networks Activated by Specific Patterns of Action Potentials in Dorsal Root Ganglia Neurons Received: 10 August 2016 Philip R
www.nature.com/scientificreports OPEN Gene networks activated by specific patterns of action potentials in dorsal root ganglia neurons Received: 10 August 2016 Philip R. Lee1,*, Jonathan E. Cohen1,*, Dumitru A. Iacobas2,3, Sanda Iacobas2 & Accepted: 23 January 2017 R. Douglas Fields1 Published: 03 March 2017 Gene regulatory networks underlie the long-term changes in cell specification, growth of synaptic connections, and adaptation that occur throughout neonatal and postnatal life. Here we show that the transcriptional response in neurons is exquisitely sensitive to the temporal nature of action potential firing patterns. Neurons were electrically stimulated with the same number of action potentials, but with different inter-burst intervals. We found that these subtle alterations in the timing of action potential firing differentially regulates hundreds of genes, across many functional categories, through the activation or repression of distinct transcriptional networks. Our results demonstrate that the transcriptional response in neurons to environmental stimuli, coded in the pattern of action potential firing, can be very sensitive to the temporal nature of action potential delivery rather than the intensity of stimulation or the total number of action potentials delivered. These data identify temporal kinetics of action potential firing as critical components regulating intracellular signalling pathways and gene expression in neurons to extracellular cues during early development and throughout life. Adaptation in the nervous system in response to external stimuli requires synthesis of new gene products in order to elicit long lasting changes in processes such as development, response to injury, learning, and memory1. Information in the environment is coded in the pattern of action-potential firing, therefore gene transcription must be regulated by the pattern of neuronal firing. -
Drosophila and Human Transcriptomic Data Mining Provides Evidence for Therapeutic
Drosophila and human transcriptomic data mining provides evidence for therapeutic mechanism of pentylenetetrazole in Down syndrome Author Abhay Sharma Institute of Genomics and Integrative Biology Council of Scientific and Industrial Research Delhi University Campus, Mall Road Delhi 110007, India Tel: +91-11-27666156, Fax: +91-11-27662407 Email: [email protected] Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 Running head: Pentylenetetrazole mechanism in Down syndrome 1 Abstract Pentylenetetrazole (PTZ) has recently been found to ameliorate cognitive impairment in rodent models of Down syndrome (DS). The mechanism underlying PTZ’s therapeutic effect is however not clear. Microarray profiling has previously reported differential expression of genes in DS. No mammalian transcriptomic data on PTZ treatment however exists. Nevertheless, a Drosophila model inspired by rodent models of PTZ induced kindling plasticity has recently been described. Microarray profiling has shown PTZ’s downregulatory effect on gene expression in fly heads. In a comparative transcriptomics approach, I have analyzed the available microarray data in order to identify potential mechanism of PTZ action in DS. I find that transcriptomic correlates of chronic PTZ in Drosophila and DS counteract each other. A significant enrichment is observed between PTZ downregulated and DS upregulated genes, and a significant depletion between PTZ downregulated and DS dowwnregulated genes. Further, the common genes in PTZ Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 downregulated and DS upregulated sets show enrichment for MAP kinase pathway. My analysis suggests that downregulation of MAP kinase pathway may mediate therapeutic effect of PTZ in DS. Existing evidence implicating MAP kinase pathway in DS supports this observation. -
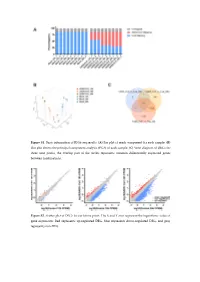
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Enhancer Λ Ig Differentiation, Bind an Essential Site in the Mef2 Proteins
Mef2 Proteins, Required for Muscle Differentiation, Bind an Essential Site in the Ig λ Enhancer This information is current as Ebenezer Satyaraj and Ursula Storb of September 24, 2021. J Immunol 1998; 161:4795-4802; ; http://www.jimmunol.org/content/161/9/4795 Downloaded from References This article cites 52 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/161/9/4795.full#ref-list-1 Why The JI? Submit online. http://www.jimmunol.org/ • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average by guest on September 24, 2021 Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 1998 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Mef2 Proteins, Required for Muscle Differentiation, Bind an Essential Site in the Ig l Enhancer1 Ebenezer Satyaraj2* and Ursula Storb3† The Ig l light chain gene enhancer has two unique essential motifs, lA and lB. The transcription factors that bind the lB motif have been identified as Pu.1 and Pu.1-interacting partner (Pip). -

Biologic and Therapeutic Implications of Genomic Alterations in Acute Lymphoblastic Leukemia
Journal of Clinical Medicine Review Biologic and Therapeutic Implications of Genomic Alterations in Acute Lymphoblastic Leukemia Ilaria Iacobucci 1,*, Shunsuke Kimura 1 and Charles G. Mullighan 1,2,* 1 Department of Pathology, St. Jude Children’s Research Hospital, 262 Danny Thomas Place, Memphis, TN 38105, USA; [email protected] 2 Comprehensive Cancer Center, Hematological Malignancies Program, St. Jude Children’s Research Hospital, 262 Danny Thomas Place, Memphis, TN 38105, USA * Correspondence: [email protected] (I.I.); [email protected] (C.G.M.) Abstract: Acute lymphoblastic leukemia (ALL) is the most successful paradigm of how risk-adapted therapy and detailed understanding of the genetic alterations driving leukemogenesis and therapeutic response may dramatically improve treatment outcomes, with cure rates now exceeding 90% in children. However, ALL still represents a leading cause of cancer-related death in the young, and the outcome for older adolescents and young adults with ALL remains poor. In the past decade, next generation sequencing has enabled critical advances in our understanding of leukemogenesis. These include the identification of risk-associated ALL subtypes (e.g., those with rearrangements of MEF2D, DUX4, NUTM1, ZNF384 and BCL11B; the PAX5 P80R and IKZF1 N159Y mutations; and genomic phenocopies such as Ph-like ALL) and the genomic basis of disease evolution. These advances have been complemented by the development of novel therapeutic approaches, including those that are of mutation-specific, such as tyrosine kinase inhibitors, and those that are mutation- Citation: Iacobucci, I.; Kimura, S.; agnostic, including antibody and cellular immunotherapies, and protein degradation strategies such Mullighan, C.G. Biologic and as proteolysis-targeting chimeras. -

Mutational Landscape and Patterns of Clonal Evolution in Relapsed Pediatric Acute Lymphoblastic Leukemia
RESEARCH ARTICLE Mutational Landscape and Patterns of Clonal Evolution in Relapsed Pediatric Acute Lymphoblastic Leukemia Esmé Waanders1,2,3, Zhaohui Gu1, Stephanie M. Dobson4,5, Željko Antić2, Jeremy Chase Crawford6, Xiaotu Ma7, Michael N. Edmonson7, Debbie Payne-Turner1, Maartje van de Vorst8, Marjolijn C.J. Jongmans2,3, Irina McGuire9, Xin Zhou7, Jian Wang7, Lei Shi10, Stanley Pounds10, Deqing Pei10, Cheng Cheng10, Guangchun Song1, Yiping Fan7, Ying Shao7, Michael Rusch7, Kelly McCastlain1, Jiangyan Yu2, Ruben van Boxtel2, Francis Blokzijl11,12, Ilaria Iacobucci1, Kathryn G. Roberts1, Ji Wen1, Gang Wu7, Jing Ma1, John Easton7, Geoffrey Neale13, Scott R. Olsen13, Kim E. Nichols14, Ching-Hon Pui14, Jinghui Zhang7, William E. Evans15, Mary V. Relling15, Jun J. Yang15, Paul G. Thomas6, John E. Dick4,5, Roland P. Kuiper2, and Charles G. Mullighan1 Downloaded from https://bloodcancerdiscov.aacrjournals.org by guest on October 2, 2021. Copyright 2020 American Copyright 2020 by AssociationAmerican for Association Cancer Research. for Cancer Research. ABSTRACT Relapse of acute lymphoblastic leukemia (ALL) remains a leading cause of child- hood cancer-related death. Prior studies have shown clonal mutations at relapse often arise from relapse-fated subclones that exist at diagnosis. However, the genomic landscape, evolutionary trajectories, and mutational mechanisms driving relapse are incompletely understood. In an analysis of 92 cases of relapsed childhood ALL incorporating multimodal DNA and RNA sequencing, deep digital mutational tracking, and xenografting to formally define clonal structure, we identied 50 significant targets of mutation with distinct patterns of mutational acquisition or enrichment.CREBBP, NOTCH1, and RAS signaling mutations arose from diagnosis subclones, whereas variants in NCOR2, USH2A, and NT5C2 were exclusively observed at relapse.