An Intron Enhancer Recognized by Splicing Factors Activates Polyadenylation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Protein Required for RNA Processing and Splicing in Neurospora
Proc. Nati. Acad. Sci. USA Vol. 89, pp. 1676-1680, March 1992 Genetics A protein required for RNA processing and splicing in Neurospora mitochondria is related to gene products involved in cell cycle protein phosphatase functions BEATRICE TURCQ*, KATHERINE F. DOBINSONt, NOBUFUSA SERIZAWAt, AND ALAN M. LAMBOWITZ§ Departments of Molecular Genetics and Biochemistry, and the Biotechnology Center, The Ohio State University, 484 West Twelfth Avenue, Columbus, OH 43210 Communicated by Thomas R. Cech, November 25, 1991 ABSTRACT The Neurospora crassa cyt4 mutants have specifically affect the splicing of group I introns and have pleiotropic defects in mitochondrial RNA splicing, 5' and 3' end relatively little effect on other mitochondrial RNA processing processing, and RNA turnover. Here, we show that the cyt-4 reactions (1, 4, 7). gene encodes a 120-kDa protein with significant similarity to By contrast, the cyt4 mutants have a complex phenotype the SSD1/SRK1 protein of Saceharomyces cerevisiae and the with defects in a number of different aspects of mitochondrial DIS3 protein of Sclhizosaccharomyces pombe, which have been RNA metabolism in addition to RNA splicing (9, 10). The five implicated in protein phosphatase functions that regulate cell cyt4 mutants are cold sensitive; they grow slowly at 250C and cycle and mitotic chromosome segregation. The CYT-4 protein more rapidly at 370C. Initial studies focusing on the mitochon- is present in mitochondria and is truncated or deficient in two drial large rRNA showed that the cyt4 mutants are defective cyt4 mutants. Assuming that the CYT-4 protein functions in a in both splicing and 3' end synthesis and that the splicing defect manner similar to the SSD1/SRK1 and DIS3 proteins, we infer is more pronounced at lower temperatures (9). -

Differential Analysis of Gene Expression and Alternative Splicing
i bioRxiv preprint doi: https://doi.org/10.1101/2020.08.23.234237; this version posted August 24, 2020. The copyright holder for this preprint i (which was not certified by peer review)“main” is the author/funder, — 2020/8/23 who has — granted 9:38 — bioRxiv page a1 license — #1 to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. i i EmpiReS: Differential Analysis of Gene Expression and Alternative Splicing Gergely Csaba∗, Evi Berchtold*,y, Armin Hadziahmetovic, Markus Gruber, Constantin Ammar and Ralf Zimmer Department of Informatics, Ludwig-Maximilians-Universitat¨ Munchen,¨ 80333, Munich, Germany ABSTRACT to translation. Different transcripts of the same gene are often expressed at the same time, possibly at different abundance, While absolute quantification is challenging in high- yielding a defined mixture of isoforms. Changes to this throughput measurements, changes of features composition are further referred to as differential alternative between conditions can often be determined with splicing (DAS). Various diseases can be linked to DAS high precision. Therefore, analysis of fold changes events (3). Most hallmarks of cancer like tumor progression is the standard method, but often, a doubly and immune escape (4, 5) can be attributed to changes in differential analysis of changes of changes is the transcript composition (6). DAS plays a prominent role required. Differential alternative splicing is an in various types of muscular dystrophy (MD) (7), splicing example of a doubly differential analysis, i.e. fold intervention by targeted exon removal is a therapeutic option changes between conditions for different isoforms in Duchenne MD (8). -

Coupling of Spliceosome Complexity to Intron Diversity
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.19.436190; this version posted March 20, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Coupling of spliceosome complexity to intron diversity Jade Sales-Lee1, Daniela S. Perry1, Bradley A. Bowser2, Jolene K. Diedrich3, Beiduo Rao1, Irene Beusch1, John R. Yates III3, Scott W. Roy4,6, and Hiten D. Madhani1,6,7 1Dept. of Biochemistry and Biophysics University of California – San Francisco San Francisco, CA 94158 2Dept. of Molecular and Cellular Biology University of California - Merced Merced, CA 95343 3Department of Molecular Medicine The Scripps Research Institute, La Jolla, CA 92037 4Dept. of Biology San Francisco State University San Francisco, CA 94132 5Chan-Zuckerberg Biohub San Francisco, CA 94158 6Corresponding authors: [email protected], [email protected] 7Lead Contact 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.03.19.436190; this version posted March 20, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. SUMMARY We determined that over 40 spliceosomal proteins are conserved between many fungal species and humans but were lost during the evolution of S. cerevisiae, an intron-poor yeast with unusually rigid splicing signals. We analyzed null mutations in a subset of these factors, most of which had not been investigated previously, in the intron-rich yeast Cryptococcus neoformans. -

Transcription to RNA from Gene to Phenotype
11/8/11 Transcription to RNA From Gene to Phenotype DNA Gene 2 • The central dogma: molecule – DNA->RNA->protein Gene 1 – One Gene, One Enzyme Gene 3 • Beadle & Tatum expts. • Transcription DNA strand 3! 5! – Initiation (template) A C C A A A C C G A G T – Elongation – Termination TRANSCRIPTION U G G U U U G G C U C A • mRNA processing mRNA 5! 3! – Introns and exons Codon TRANSLATION • Other types of RNA 11/9/2011 Protein Trp Phe Gly Ser Amino acid In Prokaryotes transcription and translation occur simultaneously In Eukaryotes Nuclear – Transcription and envelope translation occur in separate compartments TRANSCRIPTION DNA DNA TRANSCRIPTION of the cell Pre-mRNA RNA PROCESSING mRNA Ribosome – RNA transcripts are mRNA TRANSLATION modified before becoming true mRNA Ribosome Polypeptide TRANSLATION Polypeptide Figure 17.3a Figure 17.3b One Gene -> One Enzyme “One Gene -> One Enzyme” EXPERIMENT Class I Class II Class III Wild type Mutants Mutants Mutants Minimal Normal bread-mold cells can medium synthesize arginine from precursors (MM) (control) in the minimal medium MM + Ornithine Mutant 2 could MM + grow if either Citrulline Precursor Ornithine Citrulline Arginine citruline or MM + arginine was Specific enzymes (arrows) Arginine is an essential Arginine amino acid, required for (control) supplied. catalyze each step growth Therefore it must lack the enzyme to make Citruline Precursor Ornithine Citrulline Arginine 1 11/8/11 From Gene to Phenotype RNA Polymerase Non-template strand of DNA DNA Gene 2 RNA nucleotides molecule Gene 1 Gene 3 T C C A A A T 3 C T U ! 3 end DNA strand 3! 5! ! G (template) A C C A A A C C G A G T T A U G G A 5 C A U C C A C TRANSCRIPTION ! A T A A G G T T U G G U U U G G C U C A mRNA 5! 3! Direction of transcription 5! Codon (“downstream) Template TRANSLATION strand of DNA Protein Trp Phe Gly Ser New RNA Amino acid Synthesis of an RNA Transcript DNA is copied to make messenger RNA Promoter Transcription unit 5! 3! 3! 5! RNA polymerase DNA This is the “non-template” Start point binds to a promoter RNA polymerase strand. -

Nuclear Expression of a Group II Intron Is Consistent with Spliceosomal Intron Ancestry
Downloaded from genesdev.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press Nuclear expression of a group II intron is consistent with spliceosomal intron ancestry Venkata R. Chalamcharla, M. Joan Curcio, and Marlene Belfort1 Center for Medical Sciences, Wadsworth Center, New York State Department of Health, Albany, New York 12208, USA; and School of Public Health, State University of New York at Albany, Albany, New York 12201, USA Group II introns are self-splicing RNAs found in eubacteria, archaea, and eukaryotic organelles. They are mechanistically similar to the metazoan nuclear spliceosomal introns; therefore, group II introns have been invoked as the progenitors of the eukaryotic pre-mRNA introns. However, the ability of group II introns to function outside of the bacteria-derived organelles is debatable, since they are not found in the nuclear genomes of eukaryotes. Here, we show that the Lactococcus lactis Ll.LtrB group II intron splices accurately and efficiently from different pre-mRNAs in a eukaryote, Saccharomyces cerevisiae. However, a pre-mRNA harboring a group II intron is spliced predominantly in the cytoplasm and is subject to nonsense-mediated mRNA decay (NMD), and the mature mRNA from which the group II intron is spliced is poorly translated. In contrast, a pre-mRNA bearing the Tetrahymena group I intron or the yeast spliceosomal ACT1 intron at the same location is not subject to NMD, and the mature mRNA is translated efficiently. Thus, a group II intron can splice from a nuclear transcript, but RNA instability and translation defects would have favored intron loss or evolution into protein-dependent spliceosomal introns, consistent with the bacterial group II intron ancestry hypothesis. -
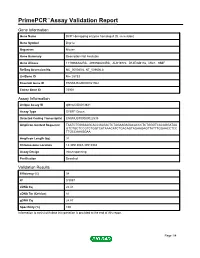
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name DCP1 decapping enzyme homolog A (S. cerevisiae) Gene Symbol Dcp1a Organism Mouse Gene Summary Description Not Available Gene Aliases 1110066A22Rik, 4930568L04Rik, AU019772, D14Ertd817e, Mitc1, SMIF RefSeq Accession No. NC_000080.6, NT_039606.8 UniGene ID Mm.28733 Ensembl Gene ID ENSMUSG00000021962 Entrez Gene ID 75901 Assay Information Unique Assay ID qMmuCID0013841 Assay Type SYBR® Green Detected Coding Transcript(s) ENSMUST00000022535 Amplicon Context Sequence TAATCTGGGAAGCACCGAGACTCTAGAAGAGACACCCTCTGGGTCACAGGATAA GTCTGCTCCGTCTGGTCATAAACATCTGACAGTAGAAGAGTTATTTGGAACCTCC TTGCCAAAGGAA Amplicon Length (bp) 91 Chromosome Location 14:30513043-30518984 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 98 R2 0.9997 cDNA Cq 22.41 cDNA Tm (Celsius) 81 gDNA Cq 24.87 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report Dcp1a, Mouse Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Mouse Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. Detected Coding Transcript(s) This is a list of the Ensembl transcript ID(s) that this assay will detect. -

Exon Amplification: a Strategy to Isolate Mammalian Genes Based on RNA Splicing (Gene Cloning/Polymerase Chain Reaction) ALAN J
Proc. Natl. Acad. Sci. USA Vol. 88, pp. 4005-4009, May 1991 Genetics Exon amplification: A strategy to isolate mammalian genes based on RNA splicing (gene cloning/polymerase chain reaction) ALAN J. BUCKLER*, DAVID D. CHANG, SHARON L. GRAw, J. DAVID BROOK, DANIEL A. HABER, PHILLIP A. SHARP, AND DAVID E. HOUSMAN Center for Cancer Research, Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02139 Contributed by Phillip A. Sharp, January 25, 1991 ABSTRACT We have developed a method, exon amplifi- (7, 8). Thus, this method may be generally applicable for the cation, for fast and efficient isolation of coding sequences from selection of exon sequences from any gene. The method is complex mammalian genomic DNA. This method is based on also both rapid and easily adapted to large scale experiments. the selection of RNA sequences, exons, which are flanked by A series of cloned genomic DNA fragments can be screened functional 5' and 3' splice sites. Fragments of cloned genomic within 1-2 weeks. The sensitivity of this method is high. DNA are inserted into an intron, which is flanked by 5' and 3' Genomic DNA segments of 20 kilobases (kb) or more can be splice sites of the human immunodeficiency virus 1 tat gene successfully screened in a single transfection by using a set contained within the plasmid pSPL1. COS-7 cells are trans- of pooled subclones. This method thus allows the rapid fected with these constructs, and the resulting RNA transcripts identification of exons in mammalian genomic DNA and are processed in vivo. Splice sites of exons contained within the should facilitate the isolation of a wide spectrum of genes of inserted genomic fragment are paired with splice sites of the significance in physiology and development. -

Mrna Turnover Philip Mitchell* and David Tollervey†
320 mRNA turnover Philip Mitchell* and David Tollervey† Nuclear RNA-binding proteins can record pre-mRNA are cotransported to the cytoplasm with the mRNP. These processing events in the structure of messenger proteins may preserve a record of the nuclear history of the ribonucleoprotein particles (mRNPs). During initial rounds of pre-mRNA in the cytoplasmic mRNP structure. This infor- translation, the mature mRNP structure is established and is mation can strongly influence the cytoplasmic fate of the monitored by mRNA surveillance systems. Competition for the mRNA and is used by mRNA surveillance systems that act cap structure links translation and subsequent mRNA as a checkpoint of mRNP integrity, particularly in the identi- degradation, which may also involve multiple deadenylases. fication of premature translation termination codons (PTCs). Addresses Cotransport of nuclear mRNA-binding proteins with mRNA Wellcome Trust Centre for Cell Biology, ICMB, University of Edinburgh, from the nucleus to the cytoplasm (nucleocytoplasmic shut- Kings’ Buildings, Edinburgh EH9 3JR, UK tling) was first observed for the heterogeneous nuclear *e-mail: [email protected] ribonucleoprotein (hnRNP) proteins. Some hnRNP proteins †e-mail: [email protected] are stripped from the mRNA at export [1], but hnRNP A1, Current Opinion in Cell Biology 2001, 13:320–325 A2, E, I and K are all exported (see [2]). Although roles for 0955-0674/01/$ — see front matter these hnRNP proteins in transport and translation have been © 2001 Elsevier Science Ltd. All rights reserved. reported [3•,4•], their affects on mRNA stability have been little studied. More is known about hnRNP D/AUF1 and Abbreviations AREs AU-rich sequence elements another nuclear RNA-binding protein, HuR, which act CBC cap-binding complex antagonistically to modulate the stability of a range of DAN deadenylating nuclease mRNAs containing AU-rich sequence elements (AREs) DSEs downstream sequence elements (reviewed in [2]). -

RNA Components of the Spliceosome Regulate Tissue- and Cancer-Specific Alternative Splicing
Downloaded from genome.cshlp.org on September 29, 2021 - Published by Cold Spring Harbor Laboratory Press Research RNA components of the spliceosome regulate tissue- and cancer-specific alternative splicing Heidi Dvinge,1,2,4 Jamie Guenthoer,3 Peggy L. Porter,3 and Robert K. Bradley1,2 1Computational Biology Program, Public Health Sciences Division, Fred Hutchinson Cancer Research Center, Seattle, Washington 98109, USA; 2Basic Sciences Division, Fred Hutchinson Cancer Research Center, Seattle, Washington 98109, USA; 3Human Biology Division, Fred Hutchinson Cancer Research Center, Seattle, Washington 98109, USA Alternative splicing of pre-mRNAs plays a pivotal role during the establishment and maintenance of human cell types. Characterizing the trans-acting regulatory proteins that control alternative splicing has therefore been the focus of much research. Recent work has established that even core protein components of the spliceosome, which are required for splicing to proceed, can nonetheless contribute to splicing regulation by modulating splice site choice. We here show that the RNA components of the spliceosome likewise influence alternative splicing decisions. Although these small nuclear RNAs (snRNAs), termed U1, U2, U4, U5, and U6 snRNA, are present in equal stoichiometry within the spliceosome, we found that their relative levels vary by an order of magnitude during development, across tissues, and across cancer samples. Physiologically relevant perturbation of individual snRNAs drove widespread gene-specific differences in alternative splic- ing but not transcriptome-wide splicing failure. Genes that were particularly sensitive to variations in snRNA abundance in a breast cancer cell line model were likewise preferentially misspliced within a clinically diverse cohort of invasive breast ductal carcinomas. -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Sequences at the Exon-Intron Boundaries* (Split Gene/Mrna Splicing/Eukaryotic Gene Structure) R
Proc. Nati. Acad. Sci. USA Vol. 75, No. 10, pp. 4853-4857, October 1978 Biochemistry Ovalbumin gene: Evidence for a leader sequence in mRNA and DNA sequences at the exon-intron boundaries* (split gene/mRNA splicing/eukaryotic gene structure) R. BREATHNACH, C. BENOIST, K. O'HARE, F. GANNON, AND P. CHAMBON Laboratoire de Genetique Mol6culaire des Eucaryotes du Centre National de la Recherche Scientifique, Unite 44 de l'Institut National de la Sant6 et de la Recherche MWdicale, Institut de Chimie Biologique, Facult6 de Melecine, Strasbourg 67085, France Communicated by A. Frey-Wyssling, July 31, 1978 ABSTRACT Selected regions of cloned EcoRI fragments the 5' end of ov-mRNA and have revealed some interesting of the chicken ovalbumin gene have been sequenced. The po- features in the DNA sequences at exon-intron boundaries. sitions where the sequences coding for ovalbumin mRNA (ov- mRNA) are interrupted in the genome have been determined, and a previously unreported interruption in the DNA sequences MATERIALS AND METHODS coding for the 5' nontranslated region of the messenger has been discovered. Because directly repeated sequences are found at Plasmid pCR1 ov 2.1 containing the ov-ds-cDNA insert was exon-intron boundaries, the nucleotide sequence alone cannot prepared as described (9). EcoRI fragments "b," "c," and "d" define unique excision-ligation points for the processing of a previously cloned in X vectors (3) were transferred to the plas- possible ov-mRNA precursor. However, the sequences in these mid pBR 322. An EcoRI/HindIII of the EcoRI boundary regions share common features; this leads to the fragment proposal that there are, in fact, unique excision-ligation points fragment "a" containing the entirety of exon 7 (Fig. -

Splicing Graphs and RNA-Seq Data
Splicing graphs and RNA-seq data Herv´ePag`es Daniel Bindreither Marc Carlson Martin Morgan Last modified: January 2019; Compiled: May 19, 2021 Contents 1 Introduction 1 2 Splicing graphs 2 2.1 Definitions and example . .2 2.2 Details about nodes and edges . .2 2.3 Uninformative nodes . .5 3 Computing splicing graphs from annotations 6 3.1 Choosing and loading a gene model . .6 3.2 Generating a SplicingGraphs object . .6 3.3 Basic manipulation of a SplicingGraphs object . .8 3.4 Extracting and plotting graphs from a SplicingGraphs object . 12 4 Splicing graph bubbles 15 4.1 Some definitions . 15 4.2 Computing the bubbles of a splicing graph . 17 4.3 AScodes ............................................... 18 4.4 Tabulating all the AS codes for Human chromosome 14 . 19 5 Counting reads 20 5.1 Assigning reads to the edges of a SplicingGraphs object . 22 5.2 How does read assignment work? . 23 5.3 Counting and summarizing the assigned reads . 25 6 Session Information 26 1 Introduction The SplicingGraphs package allows the user to create and manipulate splicing graphs [1] based on annotations for a given organism. Annotations must describe a gene model, that is, they need to contain the following information: The exact exon/intron structure (i.e., genomic coordinates) for the known transcripts. The grouping of transcripts by gene. Annotations need to be provided as a TxDb or GRangesList object, which is how a gene model is typically represented in Bioconductor. 1 The SplicingGraphs package defines the SplicingGraphs container for storing the splicing graphs together with the gene model that they are based on.