Emx2 Is Required for Growth of the Hippocampus but Not for Hippocampal Field Specification
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Spatially Heterogeneous Choroid Plexus Transcriptomes Encode Positional Identity and Contribute to Regional CSF Production
The Journal of Neuroscience, March 25, 2015 • 35(12):4903–4916 • 4903 Development/Plasticity/Repair Spatially Heterogeneous Choroid Plexus Transcriptomes Encode Positional Identity and Contribute to Regional CSF Production Melody P. Lun,1,3 XMatthew B. Johnson,2 Kevin G. Broadbelt,1 Momoko Watanabe,4 Young-jin Kang,4 Kevin F. Chau,1 Mark W. Springel,1 Alexandra Malesz,1 Andre´ M.M. Sousa,5 XMihovil Pletikos,5 XTais Adelita,1,6 Monica L. Calicchio,1 Yong Zhang,7 Michael J. Holtzman,7 Hart G.W. Lidov,1 XNenad Sestan,5 Hanno Steen,1 XEdwin S. Monuki,4 and Maria K. Lehtinen1 1Department of Pathology, and 2Division of Genetics, Boston Children’s Hospital, Boston, Massachusetts 02115, 3Department of Pathology and Laboratory Medicine, Boston University School of Medicine, Boston, Massachusetts 02118, 4Department of Pathology and Laboratory Medicine, University of California Irvine School of Medicine, Irvine, California 92697, 5Department of Neurobiology and Kavli Institute for Neuroscience, Yale School of Medicine, New Haven, Connecticut 06510, 6Department of Biochemistry, Federal University of Sa˜o Paulo, Sa˜o Paulo 04039, Brazil, and 7Pulmonary and Critical Care Medicine, Department of Medicine, Washington University, St Louis, Missouri 63110 A sheet of choroid plexus epithelial cells extends into each cerebral ventricle and secretes signaling factors into the CSF. To evaluate whether differences in the CSF proteome across ventricles arise, in part, from regional differences in choroid plexus gene expression, we defined the transcriptome of lateral ventricle (telencephalic) versus fourth ventricle (hindbrain) choroid plexus. We find that positional identitiesofmouse,macaque,andhumanchoroidplexiderivefromgeneexpressiondomainsthatparalleltheiraxialtissuesoforigin.We thenshowthatmolecularheterogeneitybetweentelencephalicandhindbrainchoroidplexicontributestoregion-specific,age-dependent protein secretion in vitro. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The Role of Gli3 in Inflammation
University of New Hampshire University of New Hampshire Scholars' Repository Doctoral Dissertations Student Scholarship Winter 2020 THE ROLE OF GLI3 IN INFLAMMATION Stephan Josef Matissek University of New Hampshire, Durham Follow this and additional works at: https://scholars.unh.edu/dissertation Recommended Citation Matissek, Stephan Josef, "THE ROLE OF GLI3 IN INFLAMMATION" (2020). Doctoral Dissertations. 2552. https://scholars.unh.edu/dissertation/2552 This Dissertation is brought to you for free and open access by the Student Scholarship at University of New Hampshire Scholars' Repository. It has been accepted for inclusion in Doctoral Dissertations by an authorized administrator of University of New Hampshire Scholars' Repository. For more information, please contact [email protected]. THE ROLE OF GLI3 IN INFLAMMATION BY STEPHAN JOSEF MATISSEK B.S. in Pharmaceutical Biotechnology, Biberach University of Applied Sciences, Germany, 2014 DISSERTATION Submitted to the University of New Hampshire in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy In Biochemistry December 2020 This dissertation was examined and approved in partial fulfillment of the requirement for the degree of Doctor of Philosophy in Biochemistry by: Dissertation Director, Sherine F. Elsawa, Associate Professor Linda S. Yasui, Associate Professor, Northern Illinois University Paul Tsang, Professor Xuanmao Chen, Assistant Professor Don Wojchowski, Professor On October 14th, 2020 ii ACKNOWLEDGEMENTS First, I want to express my absolute gratitude to my advisor Dr. Sherine Elsawa. Without her help, incredible scientific knowledge and amazing guidance I would not have been able to achieve what I did. It was her encouragement and believe in me that made me overcome any scientific struggles and strengthened my self-esteem as a human being and as a scientist. -

CIC Is a Critical Regulator of Neuronal Differentiation
CIC is a critical regulator of neuronal differentiation Inah Hwang, … , Hongwu Zheng, Jihye Paik JCI Insight. 2020. https://doi.org/10.1172/jci.insight.135826. Research In-Press Preview Oncology Stem cells Capicua (CIC), a member of the high mobility group (HMG)-box superfamily of transcriptional repressors, is frequently mutated in human oligodendrogliomas. But its function in brain development and tumorigenesis remains poorly understood. Here, we report that brain-specific deletion of Cic compromises developmental transition of neuroblast to immature neurons in mouse hippocampus and compromises normal neuronal differentiation. Combined gene expression and ChIP-seq analyses identified VGF as an important CIC-repressed transcriptional surrogate involved in neuronal lineage regulation. Aberrant VGF expression promotes neural progenitor cell proliferation by suppressing their differentiation. Mechanistically, we demonstrated that CIC represses VGF expression by tethering SIN3-HDAC to form a transcriptional corepressor complex. Mass spectrometry analysis of CIC-interacting proteins further identified BRG1 containing mSWI/SNF complex whose function is necessary for transcriptional repression by CIC. Together, this study uncovers a novel regulatory pathway of CIC-dependent neuronal differentiation and may implicate these molecular mechanisms in CIC-dependent brain tumorigenesis. Find the latest version: https://jci.me/135826/pdf CIC is a Critical Regulator of Neuronal Differentiation Inah Hwang1,4, Heng Pan2,3,4, Jun Yao5, Olivier Elemento2,3,4, Hongwu Zheng1,4 and Jihye Paik1,4* 1Department of Pathology and Laboratory medicine. Weil Cornell Medicine, New York, NY10021, USA 2Department of Physiology and Biophysics, Weil Cornell Medicine, New York, NY10021, USA 3Caryl and Israel Englander Institute for Precision Medicine, NewYork-Presbyterian Hospital. -

The Role of Emx1 and Emx2 in the Developing Chick Telencephalon
The Role of Emx1 and Emx2 in the developing chick telencephalon Dissertation Zur Erlangung des Doktorgrades der Naturwissenschaften (Dr.rer.nat.) der Fakultät für Biologie der Ludwig-Maximilian-Universität München Angefertigt am Max-Planck-Institut für Neurobiologie in der Arbeitsgruppe Neuronale Spezifizierung und in der GSF am Institut für Stammzellforschung Julia von Frowein München, Dezember 2004 1. Gutachter: Prof.Dr. Magdalena Götz 2. Gutachter: Prof.Dr. George Boyan eingereicht am 20.12.2004 Tag der mündlichen Prüfung: 26.4.2005 If the brain were so simple we could understand it, we would be so simple we couldn't. Lyall Watson Table of content 1 Table of content 1 Table of content...........................................................................................1 2 Abstract ........................................................................................................5 3 Zusammenfassung......................................................................................6 4 Introduction..................................................................................................8 4.1 General development of the regions of the central nervous system ....................................8 4.2 Patterning and regionalization .............................................................................................8 4.3 Regions of the forebrain.....................................................................................................10 4.4 Migration............................................................................................................................12 -

Cortical Foxp2 Supports Behavioral Flexibility and Developmental Dopamine D1 Receptor Expression
bioRxiv preprint doi: https://doi.org/10.1101/624973; this version posted May 2, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Cortical Foxp2 supports behavioral flexibility and developmental dopamine D1 receptor expression Marissa Co, Stephanie L. Hickey, Ashwinikumar Kulkarni, Matthew Harper, Genevieve Konopka* Department of Neuroscience, University of Texas Southwestern Medical Center, Dallas, TX, USA *Correspondence: [email protected] Contact information Genevieve Konopka, Ph.D. Department of Neuroscience, University of Texas Southwestern Medical Center, 5323 Harry Hines Blvd., ND4.300, Dallas, TX 75390-9111 TEL: 214-648-5135, FAX: 214-648-1801, Email: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/624973; this version posted May 2, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract FoxP2 encodes a forkhead box transcription factor required for the development of neural circuits underlying language, vocalization, and motor-skill learning. Human genetic studies have associated FOXP2 variation with neurodevelopmental disorders (NDDs), and within the cortex, it is coexpressed and interacts with other NDD-associated transcription factors. Cortical Foxp2 is required in mice for proper social interactions, but its role in other NDD-relevant behaviors and molecular pathways is unknown. -

REVIEW Hormones and the Hippocampus
205 REVIEW Hormones and the hippocampus R Lathe Centre for Genome Research and Centre for Neuroscience, University of Edinburgh, West Mains Road, Edinburgh EH9 3JQ, UK (Requests for offprints should be addressed to R Lathe, Centre for Genome Research, King’s Buildings, West Mains Road, Edinburgh EH9 3JQ, UK; Email: [email protected]) Abstract Hippocampal lesions produce memory deficits, but the Functionally, hippocampal enteroception may reflect feed- exact function of the hippocampus remains obscure. back control; evidence is reviewed that the hippocampus Evidence is presented that its role in memory may be modulates body physiology, including the activity of the ancillary to physiological regulation. Molecular studies hypothalamus–pituitary–adrenal axis, blood pressure, demonstrate that the hippocampus is a primary target for immunity, and reproductive function. It is suggested that ligands that reflect body physiology, including ion balance the hippocampus operates, in parallel with the amygdala, and blood pressure, immunity, pain, reproductive status, to modulate body physiology in response to cognitive satiety and stress. Hippocampal receptors are functional, stimuli. Hippocampal outputs are predominantly inhibi- probably accessible to their ligands, and mediate physio- tory on downstream neuroendocrine activity; increased logical and cognitive changes. This argues that an early synaptic efficacy in the hippocampus (e.g. long-term role of the hippocampus may have been in sensing soluble potentiation) could facilitate throughput inhibition. -

A Network of Genetic Repression and Derepression Specifies Projection
A network of genetic repression and derepression INAUGURAL ARTICLE specifies projection fates in the developing neocortex Karpagam Srinivasana,1, Dino P. Leonea, Rosalie K. Batesona, Gergana Dobrevab, Yoshinori Kohwic, Terumi Kohwi-Shigematsuc, Rudolf Grosschedlb, and Susan K. McConnella,2 aDepartment of Biology, Stanford University, Stanford, CA 94305; bMax-Planck Institute of Immunobiology and Epigenetics, 79108 Freiburg, Germany; and cLawrence Berkeley National Laboratory, Berkeley, CA 94720 This contribution is part of the special series of Inaugural Articles by members of the National Academy of Sciences elected in 2011. Contributed by Susan K. McConnell, September 28, 2012 (sent for review August 24, 2012) Neurons within each layer in the mammalian cortex have stereotypic which results in a suppression of corticothalamic and callosal projections. Four genes—Fezf2, Ctip2, Tbr1, and Satb2—regulate fates, respectively, and axon extension along the corticospinal these projection identities. These genes also interact with each tract (CST). Fezf2 mutant neurons fail to repress Satb2 and Tbr1, other, and it is unclear how these interactions shape the final pro- thus their axons cross the CC and/or innervate the thalamus jection identity. Here we show, by generating double mutants of inappropriately. Fezf2, Ctip2, and Satb2, that cortical neurons deploy a complex In callosal projection neurons, Satb2 represses the expression Ctip2 Bhlhb5 genetic switch that uses mutual repression to produce subcortical of and , leading to a repression of subcerebral fates. Satb2 Tbr1 or callosal projections. We discovered that Tbr1, EphA4, and Unc5H3 Interestingly, we found that promotes expression in fi upper layer callosal neurons, and Tbr1 expression in these neu- are critical downstream targets of Satb2 in callosal fate speci ca- fi tion. -
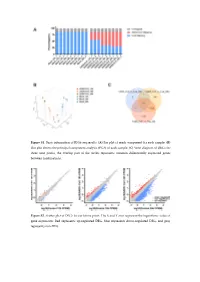
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Conversion of Cerebral Cortex Into Basal Ganglia in Emx2 Pax6 Double
articles Conversion of cerebral cortex into basal ganglia in Emx2 –/– Pax6Sey/Sey double-mutant mice Luca Muzio1, Barbara DiBenedetto1, Anastassia Stoykova2, Edoardo Boncinelli3, Peter Gruss2 and Antonello Mallamaci1 1 Department of Biological and Technological Research (DIBIT), Istituto Scientifico H. San Raffaele, via Olgettina 58, 20132 Milan, Italy 2 Max-Planck Institute of Biophysical Chemistry, Am Fassberg, 37018 Goettingen, Germany 3 International School for Advanced Studies, via Beirut 2/4, 34014 Trieste, Italy Correspondence should be addressed to A.M. ([email protected]) Published online: 15 July 2002, doi:10.1038/nn892 The molecular mechanisms that activate morphogenesis of cerebral cortex are currently the subject of intensive experimental analysis. Transcription factor genes of the homeobox, basic helix-loop- helix (bHLH) and zinc-finger families have recently been shown to have essential roles in this process. However, the actual selector genes activating corticogenesis have not yet been identified. Here we show that high-level expression of at least one functional allele of either of the homeobox genes Emx2 or Pax6 in the dorsal telencephalon is necessary and sufficient to stably activate morphogenesis of cerebral cortex and to repress that of adjacent structures, such as striatum. The molecular mechanisms that trigger morphogenesis of cere- morphologically and molecularly distinguishable from adjacent © http://neurosci.nature.com Group 2002 Nature Publishing bral cortex instead of adjacent telencephalic structures, -

Region-Specific Foxp2 Deletions in Cortex, Striatum Or Cerebellum
www.nature.com/scientificreports OPEN Region‑specifc Foxp2 deletions in cortex, striatum or cerebellum cannot explain vocalization defcits observed in spontaneous global knockouts Bastiaan H. A. Urbanus1*, Saša Peter1, Simon E. Fisher2,3 & Chris I. De Zeeuw1,4* FOXP2 has been identifed as a gene related to speech in humans, based on rare mutations that yield signifcant impairments in speech at the level of both motor performance and language comprehension. Disruptions of the murine orthologue Foxp2 in mouse pups have been shown to interfere with production of ultrasonic vocalizations (USVs). However, it remains unclear which structures are responsible for these defcits. Here, we show that conditional knockout mice with selective Foxp2 deletions targeting the cerebral cortex, striatum or cerebellum, three key sites of motor control with robust neural gene expression, do not recapture the profle of pup USV defcits observed in mice with global disruptions of this gene. Moreover, we observed that global Foxp2 knockout pups show substantive reductions in USV production as well as an overproduction of short broadband noise “clicks”, which was not present in the brain region‑specifc knockouts. These data indicate that defcits of Foxp2 expression in the cortex, striatum or cerebellum cannot solely explain the disrupted vocalization behaviours in global Foxp2 knockouts. Our fndings raise the possibility that the impact of Foxp2 disruption on USV is mediated at least in part by efects of this gene on the anatomical prerequisites for vocalizing. Studies of a large three-generation British family (the KE family) identifed a missense mutation of the human FOXP2 gene causing a severe speech and language disorder1. -

BMC Biology Biomed Central
BMC Biology BioMed Central Research article Open Access Classification and nomenclature of all human homeobox genes PeterWHHolland*†1, H Anne F Booth†1 and Elspeth A Bruford2 Address: 1Department of Zoology, University of Oxford, South Parks Road, Oxford, OX1 3PS, UK and 2HUGO Gene Nomenclature Committee, European Bioinformatics Institute (EMBL-EBI), Wellcome Trust Genome Campus, Hinxton, Cambridgeshire, CB10 1SA, UK Email: Peter WH Holland* - [email protected]; H Anne F Booth - [email protected]; Elspeth A Bruford - [email protected] * Corresponding author †Equal contributors Published: 26 October 2007 Received: 30 March 2007 Accepted: 26 October 2007 BMC Biology 2007, 5:47 doi:10.1186/1741-7007-5-47 This article is available from: http://www.biomedcentral.com/1741-7007/5/47 © 2007 Holland et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: The homeobox genes are a large and diverse group of genes, many of which play important roles in the embryonic development of animals. Increasingly, homeobox genes are being compared between genomes in an attempt to understand the evolution of animal development. Despite their importance, the full diversity of human homeobox genes has not previously been described. Results: We have identified all homeobox genes and pseudogenes in the euchromatic regions of the human genome, finding many unannotated, incorrectly annotated, unnamed, misnamed or misclassified genes and pseudogenes.