Goodness of Fit Tests Multinomial Distribution
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Goodness of Fit: What Do We Really Want to Know? I
PHYSTAT2003, SLAC, Stanford, California, September 8-11, 2003 Goodness of Fit: What Do We Really Want to Know? I. Narsky California Institute of Technology, Pasadena, CA 91125, USA Definitions of the goodness-of-fit problem are discussed. A new method for estimation of the goodness of fit using distance to nearest neighbor is described. Performance of several goodness-of-fit methods is studied for time-dependent CP asymmetry measurements of sin(2β). 1. INTRODUCTION of the test is then defined as the probability of ac- cepting the null hypothesis given it is true, and the The goodness-of-fit problem has recently attracted power of the test, 1 − αII , is defined as the probabil- attention from the particle physics community. In ity of rejecting the null hypothesis given the alterna- modern particle experiments, one often performs an tive is true. Above, αI and αII denote Type I and unbinned likelihood fit to data. The experimenter Type II errors, respectively. An ideal hypothesis test then needs to estimate how accurately the fit function is uniformly most powerful (UMP) because it gives approximates the observed distribution. A number of the highest power among all possible tests at the fixed methods have been used to solve this problem in the confidence level. In most realistic problems, it is not past [1], and a number of methods have been recently possible to find a UMP test and one has to consider proposed [2, 3] in the physics literature. various tests with acceptable power functions. For binned data, one typically applies a χ2 statistic There is a long-standing controversy about the con- to estimate the fit quality. -

Linear Regression: Goodness of Fit and Model Selection
Linear Regression: Goodness of Fit and Model Selection 1 Goodness of Fit I Goodness of fit measures for linear regression are attempts to understand how well a model fits a given set of data. I Models almost never describe the process that generated a dataset exactly I Models approximate reality I However, even models that approximate reality can be used to draw useful inferences or to prediction future observations I ’All Models are wrong, but some are useful’ - George Box 2 Goodness of Fit I We have seen how to check the modelling assumptions of linear regression: I checking the linearity assumption I checking for outliers I checking the normality assumption I checking the distribution of the residuals does not depend on the predictors I These are essential qualitative checks of goodness of fit 3 Sample Size I When making visual checks of data for goodness of fit is important to consider sample size I From a multiple regression model with 2 predictors: I On the left is a histogram of the residuals I On the right is residual vs predictor plot for each of the two predictors 4 Sample Size I The histogram doesn’t look normal but there are only 20 datapoint I We should not expect a better visual fit I Inferences from the linear model should be valid 5 Outliers I Often (particularly when a large dataset is large): I the majority of the residuals will satisfy the model checking assumption I a small number of residuals will violate the normality assumption: they will be very big or very small I Outliers are often generated by a process distinct from those which we are primarily interested in. -

Chapter 2 Simple Linear Regression Analysis the Simple
Chapter 2 Simple Linear Regression Analysis The simple linear regression model We consider the modelling between the dependent and one independent variable. When there is only one independent variable in the linear regression model, the model is generally termed as a simple linear regression model. When there are more than one independent variables in the model, then the linear model is termed as the multiple linear regression model. The linear model Consider a simple linear regression model yX01 where y is termed as the dependent or study variable and X is termed as the independent or explanatory variable. The terms 0 and 1 are the parameters of the model. The parameter 0 is termed as an intercept term, and the parameter 1 is termed as the slope parameter. These parameters are usually called as regression coefficients. The unobservable error component accounts for the failure of data to lie on the straight line and represents the difference between the true and observed realization of y . There can be several reasons for such difference, e.g., the effect of all deleted variables in the model, variables may be qualitative, inherent randomness in the observations etc. We assume that is observed as independent and identically distributed random variable with mean zero and constant variance 2 . Later, we will additionally assume that is normally distributed. The independent variables are viewed as controlled by the experimenter, so it is considered as non-stochastic whereas y is viewed as a random variable with Ey()01 X and Var() y 2 . Sometimes X can also be a random variable. -

A Study of Some Issues of Goodness-Of-Fit Tests for Logistic Regression Wei Ma
Florida State University Libraries Electronic Theses, Treatises and Dissertations The Graduate School 2018 A Study of Some Issues of Goodness-of-Fit Tests for Logistic Regression Wei Ma Follow this and additional works at the DigiNole: FSU's Digital Repository. For more information, please contact [email protected] FLORIDA STATE UNIVERSITY COLLEGE OF ARTS AND SCIENCES A STUDY OF SOME ISSUES OF GOODNESS-OF-FIT TESTS FOR LOGISTIC REGRESSION By WEI MA A Dissertation submitted to the Department of Statistics in partial fulfillment of the requirements for the degree of Doctor of Philosophy 2018 Copyright c 2018 Wei Ma. All Rights Reserved. Wei Ma defended this dissertation on July 17, 2018. The members of the supervisory committee were: Dan McGee Professor Co-Directing Dissertation Qing Mai Professor Co-Directing Dissertation Cathy Levenson University Representative Xufeng Niu Committee Member The Graduate School has verified and approved the above-named committee members, and certifies that the dissertation has been approved in accordance with university requirements. ii ACKNOWLEDGMENTS First of all, I would like to express my sincere gratitude to my advisors, Dr. Dan McGee and Dr. Qing Mai, for their encouragement, continuous support of my PhD study, patient guidance. I could not have completed this dissertation without their help and immense knowledge. I have been extremely lucky to have them as my advisors. I would also like to thank the rest of my committee members: Dr. Cathy Levenson and Dr. Xufeng Niu for their support, comments and help for my thesis. I would like to thank all the staffs and graduate students in my department. -

Testing Goodness Of
Dr. Wolfgang Rolke University of Puerto Rico - Mayaguez CERN Phystat Seminar 1 Problem statement Hypothesis testing Chi-square Methods based on empirical distribution function Other tests Power studies Running several tests Tests for multi-dimensional data 2 ➢ We have a probability model ➢ We have data from an experiment ➢ Does the data agree with the probability model? 3 Good Model? Or maybe needs more? 4 F: cumulative distribution function 퐻0: 퐹 = 퐹0 Usually more useful: 퐻0: 퐹 ∊ ℱ0 ℱ0 a family of distributions, indexed by parameters. 5 Type I error: reject true null hypothesis Type II error: fail to reject false null hypothesis A: HT has to have a true type I error probability no higher than the nominal one (α) B: probability of committing the type II error (β) should be as low as possible (subject to A) Historically A was achieved either by finding an exact test or having a large enough sample. p value = probability to reject true null hypothesis when repeating the experiment and observing value of test statistic or something even less likely. If method works p-value has uniform distribution. 6 Note above: no alternative hypothesis 퐻1 Different problem: 퐻0: 퐹 = 푓푙푎푡 vs 퐻0: 퐹 = 푙푖푛푒푎푟 → model selection Usually better tests: likelihood ratio test, F tests, BIC etc. Easy to confuse: all GOF papers do power studies, those need specific alternative. Our question: is F a good enough model for data? We want to guard against any alternative. 7 Not again … Actually no, GOF equally important to both (everybody has a likelihood) Maybe more so for Bayesians, no non- parametric methods. -

Measures of Fit for Logistic Regression Paul D
Paper 1485-2014 SAS Global Forum Measures of Fit for Logistic Regression Paul D. Allison, Statistical Horizons LLC and the University of Pennsylvania ABSTRACT One of the most common questions about logistic regression is “How do I know if my model fits the data?” There are many approaches to answering this question, but they generally fall into two categories: measures of predictive power (like R-square) and goodness of fit tests (like the Pearson chi-square). This presentation looks first at R-square measures, arguing that the optional R-squares reported by PROC LOGISTIC might not be optimal. Measures proposed by McFadden and Tjur appear to be more attractive. As for goodness of fit, the popular Hosmer and Lemeshow test is shown to have some serious problems. Several alternatives are considered. INTRODUCTION One of the most frequent questions I get about logistic regression is “How can I tell if my model fits the data?” Often the questioner is expressing a genuine interest in knowing whether a model is a good model or a not-so-good model. But a more common motivation is to convince someone else--a boss, an editor, or a regulator--that the model is OK. There are two very different approaches to answering this question. One is to get a statistic that measures how well you can predict the dependent variable based on the independent variables. I’ll refer to these kinds of statistics as measures of predictive power. Typically, they vary between 0 and 1, with 0 meaning no predictive power whatsoever and 1 meaning perfect predictions. -

Goodness of Fit of a Straight Line to Data
5. A linear trend. 10.4 The Least Squares Regression Line LEARNING OBJECTIVES 1. To learn how to measure how well a straight line fits a collection of data. 2. To learn how to construct the least squares regression line, the straight line that best fits a collection of data. 3. To learn the meaning of the slope of the least squares regression line. 4. To learn how to use the least squares regression line to estimate the response variable y in terms of the predictor variablex. Goodness of Fit of a Straight Line to Data Once the scatter diagram of the data has been drawn and the model assumptions described in the previous sections at least visually verified (and perhaps the correlation coefficient r computed to quantitatively verify the linear trend), the next step in the analysis is to find the straight line that best fits the data. We will explain how to measure how well a straight line fits a collection of points by examining how well the line y=12x−1 fits the data set Saylor URL: http://www.saylor.org/books Saylor.org 503 To each point in the data set there is associated an “error,” the positive or negative vertical distance from the point to the line: positive if the point is above the line and negative if it is below theline. The error can be computed as the actual y-value of the point minus the y-value yˆ that is “predicted” by inserting the x-value of the data point into the formula for the line: error at data point (x,y)=(true y)−(predicted y)=y−yˆ The computation of the error for each of the five points in the data set is shown in Table 10.1 "The Errors in Fitting Data with a Straight Line". -
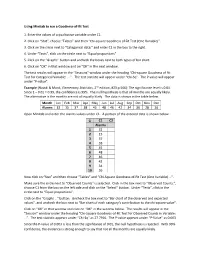
Using Minitab to Run a Goodness‐Of‐Fit Test 1. Enter the Values of A
Using Minitab to run a Goodness‐of‐fit Test 1. Enter the values of a qualitative variable under C1. 2. Click on “Stat”, choose “Tables” and then “Chi‐square Goodness of Fit Test (One Variable)”. 3. Click on the circle next to “Categorical data:” and enter C1 in the box to the right. 4. Under “Tests”, click on the circle next to “Equal proportions”. 5. Click on the “Graphs” button and uncheck the boxes next to both types of bar chart. 6. Click on “OK” in that window and on “OK” in the next window. The test results will appear in the “Sessions” window under the heading “Chi‐square Goodness of Fit Test for Categorical Variable: … ”. The test statistic will appear under “Chi‐Sq”. The P‐value will appear under “P‐value”. Example (Navidi & Monk, Elementary Statistics, 2nd edition, #23 p.566): The significance level is 0.01. Since 1 – 0.01 = 0.99, the confidence is 99%. The null hypothesis is that all months are equally likely. The alternative is the months are not all equally likely. The data is shown in the table below. Month Jan Feb Mar Apr May Jun Jul Aug Sep Oct Nov Dec Alarms 32 15 37 38 45 48 46 42 34 36 28 26 Open Minitab and enter the alarms values under C1. A portion of the entered data is shown below. ↓ C1 C2 Alarms 1 32 2 15 3 37 4 38 5 45 6 48 7 46 8 42 9 34 10 36 Now click on “Stat” and then choose “Tables” and “Chi‐Square Goodness‐of‐Fit Test (One Variable) …”. -

Goodness-Of-Fit Tests for Parametric Regression Models
Goodness-of-Fit Tests for Parametric Regression Models Jianqing Fan and Li-Shan Huang Several new tests are proposed for examining the adequacy of a family of parametric models against large nonparametric alternatives. These tests formally check if the bias vector of residuals from parametric ts is negligible by using the adaptive Neyman test and other methods. The testing procedures formalize the traditional model diagnostic tools based on residual plots. We examine the rates of contiguous alternatives that can be detected consistently by the adaptive Neyman test. Applications of the procedures to the partially linear models are thoroughly discussed. Our simulation studies show that the new testing procedures are indeed powerful and omnibus. The power of the proposed tests is comparable to the F -test statistic even in the situations where the F test is known to be suitable and can be far more powerful than the F -test statistic in other situations. An application to testing linear models versus additive models is also discussed. KEY WORDS: Adaptive Neyman test; Contiguous alternatives; Partial linear model; Power; Wavelet thresholding. 1. INTRODUCTION independent observations from a population, Parametric linear models are frequently used to describe Y m4x5 ˜1 ˜ N 401‘ 251 (1.1) the association between a response variable and its predic- D C tors. The adequacy of such parametric ts often arises. Con- ventional methods rely on residual plots against tted values where x is a p-dimensional vector and m4 5 is a smooth regres- sion surface. Let f 4 1 ˆ5 be a given parametri¢ c family. The or a covariate variable to detect if there are any system- ¢ atic departures from zero in the residuals. -

Measures of Fit for Logistic Regression Paul D
Paper 1485-2014 Measures of Fit for Logistic Regression Paul D. Allison, Statistical Horizons LLC and the University of Pennsylvania ABSTRACT One of the most common questions about logistic regression is “How do I know if my model fits the data?” There are many approaches to answering this question, but they generally fall into two categories: measures of predictive power (like R-square) and goodness of fit tests (like the Pearson chi-square). This presentation looks first at R-square measures, arguing that the optional R-squares reported by PROC LOGISTIC might not be optimal. Measures proposed by McFadden and Tjur appear to be more attractive. As for goodness of fit, the popular Hosmer and Lemeshow test is shown to have some serious problems. Several alternatives are considered. INTRODUCTION One of the most frequent questions I get about logistic regression is “How can I tell if my model fits the data?” Often the questioner is expressing a genuine interest in knowing whether a model is a good model or a not-so-good model. But a more common motivation is to convince someone else--a boss, an editor, or an audience--that the model is OK. There are two very different approaches to answering this question. One is to get a statistic that measures how well you can predict the dependent variable based on the independent variables. I’ll refer to these kinds of statistics as measures of predictive power. Typically, they vary between 0 and 1, with 0 meaning no predictive power whatsoever and 1 meaning perfect predictions. Predictive power statistics available in PROC LOGISTIC include R-square, the area under the ROC curve, and several rank-order correlations. -

Lecture 10: F-Tests, R2, and Other Distractions
00:35 Friday 16th October, 2015 See updates and corrections at http://www.stat.cmu.edu/~cshalizi/mreg/ Lecture 10: F -Tests, R2, and Other Distractions 36-401, Section B, Fall 2015 1 October 2015 Contents 1 The F Test 1 1.1 F test of β1 = 0 vs. β1 6=0 .....................2 1.2 The Likelihood Ratio Test . .7 2 What the F Test Really Tests 11 2.1 The Essential Thing to Remember . 12 3 R2 16 3.1 Theoretical R2 ............................ 17 3.2 Distraction or Nuisance? . 17 4 The Correlation Coefficient 19 5 More on the Likelihood Ratio Test 20 6 Concluding Comment 21 7 Further Reading 22 1 The F Test The F distribution with a; b degrees of freedom is defined to be the distribution of the ratio 2 χa=a 2 χb =b 2 2 when χa and χb are independent. Since χ2 distributions arise from sums of Gaussians, F -distributed random variables tend to arise when we are dealing with ratios of sums of Gaussians. The outstanding examples of this are ratios of variances. 1 2 1.1 F test of β1 = 0 vs. β1 6= 0 1.1 F test of β1 = 0 vs. β1 6= 0 Let's consider testing the null hypothesis β1 = 0 against the alternative β1 6= 0, in the context of the Gaussian-noise simple linear regression model. That is, we won't question, in our mathematics, whether or not the assumptions of that model hold, we'll presume that they all do, and just ask how we can tell whether β1 = 0. -

Goodness of Fit Tests for High-Dimensional Linear Models
Goodness of fit tests for high-dimensional linear models Rajen D. Shah∗ Peter B¨uhlmann University of Cambridge ETH Z¨urich April 8, 2017 Abstract In this work we propose a framework for constructing goodness of fit tests in both low and high-dimensional linear models. We advocate applying regression methods to the scaled residuals following either an ordinary least squares or Lasso fit to the data, and using some proxy for prediction error as the final test statistic. We call this family Residual Prediction (RP) tests. We show that simulation can be used to obtain the critical values for such tests in the low-dimensional setting, and demonstrate using both theoretical results and extensive numerical studies that some form of the parametric bootstrap can do the same when the high- dimensional linear model is under consideration. We show that RP tests can be used to test for significance of groups or individual variables as special cases, and here they compare favourably with state of the art methods, but we also argue that they can be designed to test for as diverse model misspecifications as heteroscedasticity and nonlinearity. 1 Introduction High-dimensional data, where the number of variables may greatly exceed the number of obser- vations, has become increasingly more prevalent across a variety of disciplines. While such data pose many challenges to statisticians, we now have a variety of methods for fitting models to high- dimensional data, many of which are based on the Lasso [Tibshirani, 1996]; see B¨uhlmannand van de Geer [2011] for a review of some of the developments.