An Association Study of C9orf3, a Novel Component of the Renin
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Table of Contents
Table of contents SUMMARY IX ZUSAMMENFASSUNG XI 1. INTRODUCTION 1 1.1 The immune system 1 1.1.1 Function 1 1.1.2 Overview 1 1.1.3 Function of MHC molecules 1 1.1.4 The Major Histocompatibility Complex (MHC) Genes 2 1.1.5 Structure of MHC molecules 2 1.2 Antigen presentation 5 1.2.1 Outline of the conventional MHC class I pathway 5 1.3 The components of the peptide loading complex 7 1.3.1 Transporter associated with antigen presentation (TAP) 7 1.3.2 Calreticulin and calnexin 9 1.3.4 ERp 57/ER60 10 1.3.5 Tapasin (Tpn) 11 1.4 (a) The proteasomes 12 1.4 (b) Cytosolic peptidase 15 1.5 Endoplasmic Reticulum Associated Peptidase/s 16 1.6 Protein sorting 17 1.6.2 Vesicular transport in the Golgi apparatus 20 1.6.3 Endocytic pathway 20 1.6.4 Endocytic proteases 24 1.6.4.1 Biosynthesis and transport of cathepsins 25 1.6.4.2 Activation of cathepsins 26 1.6.4.3 Inhibitors of endosomal proteases. 26 1.7 Antigen processing by alternative pathways 27 1.8 Aim of the thesis 30 i 2. MATERIALS AND METHODS 31 2.1 Materials 31 2.1.1 Bacterial strains 31 2.1.2 Plasmids 31 2.1.2.1 Construction of plasmids 31 2.1.3 Primer list 32 2.1.4 PCR conditions 33 2.1.5 Antibiotics 34 2.1.6 Chemicals 34 2.1.7 Media for bacterial cells 34 2.1.8 Protein Chemicals 35 2.1.9 Animal cell culture media 37 2.1.11 Antibodies 39 2.1.11 siRNA oligonucleotides. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
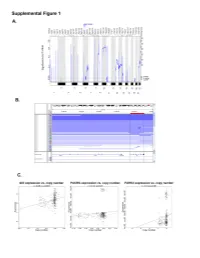
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Serine Proteases with Altered Sensitivity to Activity-Modulating
(19) & (11) EP 2 045 321 A2 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 08.04.2009 Bulletin 2009/15 C12N 9/00 (2006.01) C12N 15/00 (2006.01) C12Q 1/37 (2006.01) (21) Application number: 09150549.5 (22) Date of filing: 26.05.2006 (84) Designated Contracting States: • Haupts, Ulrich AT BE BG CH CY CZ DE DK EE ES FI FR GB GR 51519 Odenthal (DE) HU IE IS IT LI LT LU LV MC NL PL PT RO SE SI • Coco, Wayne SK TR 50737 Köln (DE) •Tebbe, Jan (30) Priority: 27.05.2005 EP 05104543 50733 Köln (DE) • Votsmeier, Christian (62) Document number(s) of the earlier application(s) in 50259 Pulheim (DE) accordance with Art. 76 EPC: • Scheidig, Andreas 06763303.2 / 1 883 696 50823 Köln (DE) (71) Applicant: Direvo Biotech AG (74) Representative: von Kreisler Selting Werner 50829 Köln (DE) Patentanwälte P.O. Box 10 22 41 (72) Inventors: 50462 Köln (DE) • Koltermann, André 82057 Icking (DE) Remarks: • Kettling, Ulrich This application was filed on 14-01-2009 as a 81477 München (DE) divisional application to the application mentioned under INID code 62. (54) Serine proteases with altered sensitivity to activity-modulating substances (57) The present invention provides variants of ser- screening of the library in the presence of one or several ine proteases of the S1 class with altered sensitivity to activity-modulating substances, selection of variants with one or more activity-modulating substances. A method altered sensitivity to one or several activity-modulating for the generation of such proteases is disclosed, com- substances and isolation of those polynucleotide se- prising the provision of a protease library encoding poly- quences that encode for the selected variants. -

Supplementary Table 9. Functional Annotation Clustering Results for the Union (GS3) of the Top Genes from the SNP-Level and Gene-Based Analyses (See ST4)
Supplementary Table 9. Functional Annotation Clustering Results for the union (GS3) of the top genes from the SNP-level and Gene-based analyses (see ST4) Column Header Key Annotation Cluster Name of cluster, sorted by descending Enrichment score Enrichment Score EASE enrichment score for functional annotation cluster Category Pathway Database Term Pathway name/Identifier Count Number of genes in the submitted list in the specified term % Percentage of identified genes in the submitted list associated with the specified term PValue Significance level associated with the EASE enrichment score for the term Genes List of genes present in the term List Total Number of genes from the submitted list present in the category Pop Hits Number of genes involved in the specified term (category-specific) Pop Total Number of genes in the human genome background (category-specific) Fold Enrichment Ratio of the proportion of count to list total and population hits to population total Bonferroni Bonferroni adjustment of p-value Benjamini Benjamini adjustment of p-value FDR False Discovery Rate of p-value (percent form) Annotation Cluster 1 Enrichment Score: 3.8978262119731335 Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_CC_DIRECT GO:0005886~plasma membrane 383 24.33290978 5.74E-05 SLC9A9, XRCC5, HRAS, CHMP3, ATP1B2, EFNA1, OSMR, SLC9A3, EFNA3, UTRN, SYT6, ZNRF2, APP, AT1425 4121 18224 1.18857065 0.038655922 0.038655922 0.086284383 UP_KEYWORDS Membrane 626 39.77128335 1.53E-04 SLC9A9, HRAS, -

1 No. Affymetrix ID Gene Symbol Genedescription Gotermsbp Q Value 1. 209351 at KRT14 Keratin 14 Structural Constituent of Cyto
1 Affymetrix Gene Q No. GeneDescription GOTermsBP ID Symbol value structural constituent of cytoskeleton, intermediate 1. 209351_at KRT14 keratin 14 filament, epidermis development <0.01 biological process unknown, S100 calcium binding calcium ion binding, cellular 2. 204268_at S100A2 protein A2 component unknown <0.01 regulation of progression through cell cycle, extracellular space, cytoplasm, cell proliferation, protein kinase C inhibitor activity, protein domain specific 3. 33323_r_at SFN stratifin/14-3-3σ binding <0.01 regulation of progression through cell cycle, extracellular space, cytoplasm, cell proliferation, protein kinase C inhibitor activity, protein domain specific 4. 33322_i_at SFN stratifin/14-3-3σ binding <0.01 structural constituent of cytoskeleton, intermediate 5. 201820_at KRT5 keratin 5 filament, epidermis development <0.01 structural constituent of cytoskeleton, intermediate 6. 209125_at KRT6A keratin 6A filament, ectoderm development <0.01 regulation of progression through cell cycle, extracellular space, cytoplasm, cell proliferation, protein kinase C inhibitor activity, protein domain specific 7. 209260_at SFN stratifin/14-3-3σ binding <0.01 structural constituent of cytoskeleton, intermediate 8. 213680_at KRT6B keratin 6B filament, ectoderm development <0.01 receptor activity, cytosol, integral to plasma membrane, cell surface receptor linked signal transduction, sensory perception, tumor-associated calcium visual perception, cell 9. 202286_s_at TACSTD2 signal transducer 2 proliferation, membrane <0.01 structural constituent of cytoskeleton, cytoskeleton, intermediate filament, cell-cell adherens junction, epidermis 10. 200606_at DSP desmoplakin development <0.01 lectin, galactoside- sugar binding, extracellular binding, soluble, 7 space, nucleus, apoptosis, 11. 206400_at LGALS7 (galectin 7) heterophilic cell adhesion <0.01 2 S100 calcium binding calcium ion binding, epidermis 12. 205916_at S100A7 protein A7 (psoriasin 1) development <0.01 S100 calcium binding protein A8 (calgranulin calcium ion binding, extracellular 13. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Systematic Evaluation of Genetic Variants for Polycystic Ovary Syndrome in a Chinese Population
RESEARCH ARTICLE Systematic Evaluation of Genetic Variants for Polycystic Ovary Syndrome in a Chinese Population Yuping Xu1,2,3, Zhiqiang Li4, Fenglian Ai1,2,3, Jianhua Chen4, Qiong Xing1,2,3, Ping Zhou1,2,3, Zhaolian Wei1,2,3, Yongyong Shi4*, Xiao-Jin He1,2,3*, Yunxia Cao1,2,3* 1 Reproductive Medicine Center, Department of Obstetrics and Gynecology, The First Affiliated Hospital of Anhui Medical University, Hefei, 230022, China, 2 Institute of Reproductive Genetics, Anhui Medical University, Hefei, 230022, China, 3 Anhui Provincial Engineering Technology Research Center for Biopreservation and Artificial Organs, Hefei, 230022, China, 4 The Bio-X Institutes, Shanghai Jiao Tong University, 1954 Huashan Road, Shanghai, 200030, China * [email protected] (YC); [email protected] (YS); [email protected] (XH) Abstract − To date, eleven genome-wide significant (GWS) loci (P < 5×10 8) for polycystic ovary syn- OPEN ACCESS drome (PCOS) have been identified through genome-wide association studies (GWAS). Citation: Xu Y, Li Z, Ai F, Chen J, Xing Q, Zhou P, et Some of the risk loci have been selected for replications and validated in multiple ethnicities, al. (2015) Systematic Evaluation of Genetic Variants however, few previous studies investigated all loci. Scanning all the GWAS variants would for Polycystic Ovary Syndrome in a Chinese demonstrate a more informative profile of variance they explained. Thus, we analyzed all Population. PLoS ONE 10(10): e0140695. the 17 single nucleotide polymorphisms (SNPs) mapping to the 11 GWAS loci in an inde- doi:10.1371/journal.pone.0140695 pendent sample set of 800 Chinese subjects with PCOS and 1110 healthy controls system- Editor: Stephen L Atkin, Weill Cornell Medical − atically. -

C:\Users\Administrator\Desktop\Array Datasheet\Disease Gene Array
Product Data Sheet ExProfileTM Human Proteases Related Gene qPCR Array For focused group profiling of human proteases genes expression Cat. No. QG096-A (6 x 96-well plate, Format A) Cat. No. QG096-B (6 x 96-well plate, Format B) Cat. No. QG096-C (6 x 96-well plate, Format C) Cat. No. QG096-D (6 x 96-well plate, Format D) Cat. No. QG096-E (6 x 96-well plate, Format E) Plates available individually or as a set of 6. Each set contains 504 unique gene primer pairs deposited in one 96-well plate. Introduction The ExProfile human proteases related gene qPCR array profiles the expression of 504 human genes related to proteases. These genes are carefully chosen for their close correlation based on a thorough literature search of peer-reviewed publications, mainly including genes that encode various proteases. This array allows researchers to study the related genes to gain understanding of their roles in the functioning and characterization of proteases. QG096 plate 01: 84 unique gene PCR primer pairs QG096 plate 02: 84 unique gene PCR primer pairs QG096 plate 03: 84 unique gene PCR primer pairs QG096 plate 04: 84 unique gene PCR primer pairs QG096 plate 05: 84 unique gene PCR primer pairs QG096 plate 06: 84 unique gene PCR primer pairs Shipping and storage condition Shipped at room temperate Stable for at least 6 months when stored at -20°C Array format GeneCopoeia provides five qPCR array formats (A, B, C, D, and E) suitable for use with the following real- time cyclers. -

WO 2019/048500 Al 14 March 2019 (14.03.2019) W 1P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/048500 Al 14 March 2019 (14.03.2019) W 1P O PCT (51) International Patent Classification: C12Q 1/6883 (2018.01) (21) International Application Number: PCT/EP20 18/073 905 (22) International Filing Date: 05 September 2018 (05.09.2018) (25) Filing Language: English (26) Publication Language: English (30) Priority Data: 62/554,427 05 September 2017 (05.09.2017) US (71) Applicant: AMONETA DIAGNOSTICS [FR/FR]; 17, Rue du Fort, 68330 Huningue (FR). (72) Inventors; and (71) Applicants: FIRAT, Hueseyin [FR/FR]; 29a, Rue de Michelfelden, 68330 Huningue (FR). MOUSSAOUI, Sali- ha [FR/FR]; 24, rue des Ecureuils, 68870 Bartenheim (FR). SCHORDAN, Eric [FR/FR]; 4 rue des Paturages, 68220 Wentzwiller (FR). (74) Agent: PHARMA CONCEPTS GMBH; Unterer Rhein- weg 50, 4057 Basel (CH). (81) Designated States (unless otherwise indicated, for every kind of national protection available) : AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. -
QKI Suppl Figs Tables Merged.Pdf
Supplemental Table 1 p<0.05 Human (Hs683) EntrezGene Name shQKI-1/shGFP shQKI-2/shGFP 0 0 26.63 19.00 0 0 11.75 12.75 0 CDNA clone IMAGE:5267346 8.96 4.77 LOC91316: Similar to bK246H3.1 (immunoglobulin lambda-like 91316 polypeptide 1, pre-B-cell specific) 7.94 3.85 RIPK2: receptor-interacting serine- 8767 threonine kinase 2 4.09 3.21 LOC728705: hypothetical protein 728705 LOC728705 3.86 2.44 0 Transcribed locus 3.79 3.95 CCDC57: coiled-coil domain containing 284001 57 3.62 4.73 C1orf101: chromosome 1 open reading 257044 frame 101 3.54 3.33 CDNA FLJ30490 fis, clone 0 BRAWH2000169 3.28 2.76 CHRNE: cholinergic receptor, nicotinic, 1145 epsilon 3.25 2.42 LOC728215: similar to transmembrane 728215 protein 28 3.25 2.86 83849 SYT15: Synaptotagmin XV 3.24 3.30 0 Transcribed locus 3.18 2.64 79838 TMC5: transmembrane channel-like 5 3.16 2.70 144568 A2ML1: alpha-2-macroglobulin-like 1 3.13 2.77 AGGF1: angiogenic factor with G patch 55109 and FHA domains 1 3.09 2.86 91703 ACY3: aspartoacylase (aminocyclase) 3 3.06 2.30 258010 SVIP: small VCP/p97-interacting protein 3.01 2.30 MSR1: macrophage scavenger receptor 4481 1 2.97 2.01 23250 ATP11A: ATPase, class VI, type 11A 2.96 1.71 LOC440934: Hypothetical gene 440934 supported by BC008048 2.89 2.27 0 Transcribed locus 2.87 3.04 Homo sapiens, clone IMAGE:4391558, 0 mRNA 2.85 3.38 C4orf38: chromosome 4 open reading 152641 frame 38 2.79 1.88 54756 IL17RD: interleukin 17 receptor D 2.77 1.98 0 CDNA clone IMAGE:5277449 2.75 2.25 OR51B2: olfactory receptor, family 51, 79345 subfamily B, member 2 2.75 1.89 0 -

Exploring the Regulatory Mechanism of the Notch Ligand Receptor Jagged1 Via the Aryl Hydrocarbon Receptor in Breast Cancer Sean Alan Piwarski
Marshall University Marshall Digital Scholar Theses, Dissertations and Capstones 2018 Exploring the Regulatory Mechanism of the Notch Ligand Receptor Jagged1 Via the Aryl Hydrocarbon Receptor in Breast Cancer Sean Alan Piwarski Follow this and additional works at: https://mds.marshall.edu/etd Part of the Biological Phenomena, Cell Phenomena, and Immunity Commons, Medical Molecular Biology Commons, Medical Toxicology Commons, and the Oncology Commons EXPLORING THE REGULATORY MECHANISM OF THE NOTCH LIGAND RECEPTOR JAGGED1 VIA THE ARYL HYDROCARBON RECEPTOR IN BREAST CANCER A dissertation submitted to the Graduate College of Marshall University In partial fulfillment of the requirements for the degree of Doctor of Philosophy In Biomedical Sciences by Sean Alan Piwarski Approved by Dr. Travis Salisbury, Committee Chairperson Dr. Gary Rankin Dr. Monica Valentovic Dr. Richard Egleton Dr. Todd Green Marshall University July 2018 APPROVAL OF DISSERTATION We, the faculty supervising the work of Sean Alan Piwarski, affirm that the dissertation, Exploring the Regulatory Mechanism of the Notch Ligand Receptor JAGGED1 via the Aryl Hydrocarbon Receptor in Breast Cancer, meets the high academic standards for original scholarship and creative work established by the Biomedical Sciences program and the Graduate College of Marshall University. This work also conforms to the editorial standards of our discipline and the Graduate College of Marshall University. With our signatures, we approve the manuscript for publication. ii © 2018 SEAN ALAN PIWARSKI ALL RIGHTS RESERVED iii DEDICATION To my mom and dad, This dissertation is a product of your hard work and sacrifice as amazing parents. I could never have accomplished something of this magnitude without your love, support, and constant encouragement. -

Genome-Wide Profiling of Druggable Active Tumor Defense Mechanisms to Enhance Cancer Immunotherapy
bioRxiv preprint doi: https://doi.org/10.1101/843185; this version posted November 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Genome-wide profiling of druggable active tumor defense mechanisms to enhance cancer immunotherapy Rigel J. Kishton1,2,*,#, Shashank J. Patel1,2,†,*, Suman K. Vodnala1,2, Amy E. Decker3, Yogin Patel1,2, Madhusudhanan Sukumar1,2, Tori N. Yamamoto1,2,4, Zhiya Yu1,2, Michelle Ji1,2, Amanda N. Henning1,2, Devikala Gurusamy1,2, Douglas C. Palmer1,2, Winifred Lo1, Anna Pasetto1, Parisa Malekzadeh1, Drew C. Deniger1, Kris C. Wood3, Neville E. Sanjana5,6, Nicholas P. Restifo1,2, #, § 1Surgery Branch, Center for Cancer Research, National Cancer Institute, Bethesda, MD 20892, USA 2Center for Cell-Based Therapy, National Cancer Institute, Bethesda, MD 20892, USA 3Department of Pharmacology & Cancer Biology, Duke University School of Medicine, Durham, NC, USA 4Immunology Graduate Group, University of Pennsylvania, Philadelphia, PA 19104, USA 5New York Genome Center, New York, NY 10013 USA 6Department of Biology, New York University, New York, NY 10003, USA *These authors contributed equally to this work. †Present address: NextCure Inc., Beltsville, MD 20705, USA §Present address: Lyell Immunopharma, South San Francisco, CA 94080, USA #Corresponding authors. NPR: [email protected]. RJK: [email protected]. bioRxiv preprint doi: https://doi.org/10.1101/843185; this version posted November 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission.