Proteolytic Removal of Core Histone Amino Termini And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Involvement of Ubiquitination Machinery in Cell Cycle Regulation and Cancer Progression
International Journal of Molecular Sciences Review The Involvement of Ubiquitination Machinery in Cell Cycle Regulation and Cancer Progression Tingting Zou and Zhenghong Lin * School of Life Sciences, Chongqing University, Chongqing 401331, China; [email protected] * Correspondence: [email protected] Abstract: The cell cycle is a collection of events by which cellular components such as genetic materials and cytoplasmic components are accurately divided into two daughter cells. The cell cycle transition is primarily driven by the activation of cyclin-dependent kinases (CDKs), which activities are regulated by the ubiquitin-mediated proteolysis of key regulators such as cyclins, CDK inhibitors (CKIs), other kinases and phosphatases. Thus, the ubiquitin-proteasome system (UPS) plays a pivotal role in the regulation of the cell cycle progression via recognition, interaction, and ubiquitination or deubiquitination of key proteins. The illegitimate degradation of tumor suppressor or abnormally high accumulation of oncoproteins often results in deregulation of cell proliferation, genomic instability, and cancer occurrence. In this review, we demonstrate the diversity and complexity of the regulation of UPS machinery of the cell cycle. A profound understanding of the ubiquitination machinery will provide new insights into the regulation of the cell cycle transition, cancer treatment, and the development of anti-cancer drugs. Keywords: cell cycle regulation; CDKs; cyclins; CKIs; UPS; E3 ubiquitin ligases; Deubiquitinases (DUBs) Citation: Zou, T.; Lin, Z. The Involvement of Ubiquitination Machinery in Cell Cycle Regulation and Cancer Progression. 1. Introduction Int. J. Mol. Sci. 2021, 22, 5754. https://doi.org/10.3390/ijms22115754 The cell cycle is a ubiquitous, complex, and highly regulated process that is involved in the sequential events during which a cell duplicates its genetic materials, grows, and di- Academic Editors: Kwang-Hyun Bae vides into two daughter cells. -

PTEN-L Is a Novel Protein Phosphatase for Ubiquitin Dephosphorylation to Inhibit PINK1–Parkin-Mediated Mitophagy
www.nature.com/cr www.cell-research.com ARTICLE OPEN PTEN-L is a novel protein phosphatase for ubiquitin dephosphorylation to inhibit PINK1–Parkin-mediated mitophagy Liming Wang1, Yik-Lam Cho1, Yancheng Tang2, Jigang Wang1, Jung-Eun Park3, Yajun Wu4, Chunxin Wang 5, Yan Tong6, Ritu Chawla1, Jianbin Zhang1,7, Yin Shi1, Shuo Deng1, Guang Lu1, Yihua Wu8, Hayden Weng-Siong Tan1,9, Pornteera Pawijit1, Grace Gui-Yin Lim10, Hui-Ying Chan1,9, Jingzi Zhang11, Lei Fang11, Hanry Yu1,12,13, Yih-Cherng Liou6, Mallilankaraman Karthik1, Boon-Huat Bay4, Kah-Leong Lim1,10, Siu-Kwan Sze 3, Celestial T. Yap1 and Han-Ming Shen1,9 Mitophagy is an important type of selective autophagy for specific elimination of damaged mitochondria. PTEN-induced putative kinase protein 1 (PINK1)-catalyzed phosphorylation of ubiquitin (Ub) plays a critical role in the onset of PINK1–Parkin-mediated mitophagy. Phosphatase and tensin homolog (PTEN)-long (PTEN-L) is a newly identified isoform of PTEN, with addition of 173 amino acids to its N-terminus. Here we report that PTEN-L is a novel negative regulator of mitophagy via its protein phosphatase activity against phosphorylated ubiquitin. We found that PTEN-L localizes at the outer mitochondrial membrane (OMM) and overexpression of PTEN-L inhibits, whereas deletion of PTEN-L promotes, mitophagy induced by various mitochondria-damaging agents. Mechanistically, PTEN-L is capable of effectively preventing Parkin mitochondrial translocation, reducing Parkin phosphorylation, maintaining its closed inactive conformation, and inhibiting its E3 ligase activity. More importantly, PTEN-L reduces the level of phosphorylated ubiquitin (pSer65-Ub) in vivo, and in vitro phosphatase assay confirms that PTEN-L dephosphorylates pSer65-Ub via its protein phosphatase activity, independently of its lipid phosphatase function. -
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
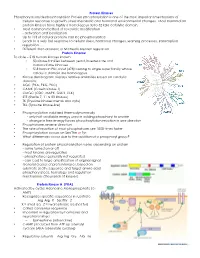
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

Computational Modeling of Lysine Post-Translational Modification: an Overview Md
c and S eti ys h te nt m y s S B Hasan MM et al., Curr Synthetic Sys Biol 2018, 6:1 t i n o e l Current Synthetic and o r r g DOI: 10.4172/2332-0737.1000137 u y C ISSN: 2332-0737 Systems Biology CommentaryResearch Article OpenOpen Access Access Computational Modeling of Lysine Post-Translational Modification: An Overview Md. Mehedi Hasan 1*, Mst. Shamima Khatun2, and Hiroyuki Kurata1,3 1Department of Bioscience and Bioinformatics, Kyushu Institute of Technology, 680-4 Kawazu, Iizuka, Fukuoka 820-8502, Japan 2Department of Statistics, Laboratory of Bioinformatics, Rajshahi University-6205, Bangladesh 3Biomedical Informatics R&D Center, Kyushu Institute of Technology, 680-4 Kawazu, Iizuka, Fukuoka 820-8502, Japan Commentary hot spot for PTMs, and a number of protein lysine modifications could occur in both histone and non-histone proteins [11,12]. For instance, Living organisms have a magnificent ordered and complex lysine methylation in non-histone proteins can regulate the protein structure. In regulating the cellular functions, post-translational activity and protein structure stability [13]. In 2004, the Nobel Prize in modifications (PTMs) are critical molecular measures. They alter Chemistry was awarded jointly to Aaron Ciechanover, Avram Hershko protein conformation, modulating their activity, stability and and Irwin Rose for the discovery of lysine ubiquitin-mediated protein localization. Up to date, more than 300 types of PTMs are experimentally degradation [14]. discovered in vivo and in vitro pathways [1,2]. Major and common PTMs are methylation, ubiquitination, succinylation, phosphorylation, Moreover, in biological process, lysine can be modified by the glycosylation, acetylation, and sumoylation. -

Protein Phosphatase 1-Α Regulates AS160 Ser588 and Thr642
2606 Diabetes Volume 65, September 2016 Pragya Sharma,1 Edward B. Arias,1 and Gregory D. Cartee1,2,3 Protein Phosphatase 1-a Regulates AS160 Ser588 and Thr642 Dephosphorylation in Skeletal Muscle Diabetes 2016;65:2606–2617 | DOI: 10.2337/db15-0867 Akt substrate of 160 kDa (AS160) phosphorylation on insulin resistance is crucial for whole-body insulin re- Thr642 and Ser588 by Akt is essential for insulin’s full ef- sistance and type 2 diabetes (1). Muscle insulin resistance fect on glucose transport. However, protein phosphory- is secondary, in large part, to defective GLUT4 transloca- lation is determined by the balance of actions by kinases tion and glucose transport (2). Insulin’s stimulation of glu- fi and phosphatases, and the speci c phosphatase(s) cose transport is triggered by a complex insulin-signaling controlling AS160 dephosphorylation is (are) unknown. pathway that begins with insulin’s binding to its receptor, Accordingly, we assessed roles of highly expressed leading to receptor autophosphorylation and activation of skeletal muscle serine/threonine phosphatases (PP1, receptor tyrosine kinase (2). The insulin receptor kinase PP2A, PP2B, and PP2C) on AS160 dephosphorylation. Preliminary screening of candidate phosphatases used phosphorylates insulin receptor substrate (IRS) proteins an AS160 dephosphorylation assay. Lysates from insu- on multiple tyrosine residues, resulting in IRS protein en- lin-stimulated skeletal muscle were treated with phar- gagement with phosphatidylinositol (PI) 3-kinase (PI3K), macological phosphatase inhibitors and assessed for that in turn, phosphorylates PI 4,5-bisphosphate to create AS160 Ser588 and Thr642 dephosphorylation. AS160 de- 3,4,5-trisphosphate (PIP3). The serine/threonine kinase phosphorylation on both phosphorylation sites was un- Akt is recruited to bind PIP3 and become activated second- altered by PP2B or PP2C inhibitors. -

A Protein Phosphatase 2Cα–Ca
1914 • The Journal of Neuroscience, February 23, 2005 • 25(8):1914–1923 Cellular/Molecular A Protein Phosphatase 2c␣–Ca2ϩ Channel Complex for Dephosphorylation of Neuronal Ca2ϩ Channels Phosphorylated by Protein Kinase C Dongjun Li, Fushun Wang, Meizan Lai, Yuan Chen, and Ji-fang Zhang Department of Physiology, Jefferson Medical College, Philadelphia, Pennsylvania 19107 Phosphorylation and dephosphorylation are primary means for rapid regulation of a variety of neuronal functions, such as membrane excitability, neurotransmitter release, and gene expression. Voltage-gated Ca 2ϩ channels are targets for phosphorylation by a variety of second messengers through activation of different types of protein kinases (PKs). Protein phosphatases (PPs), like PKs, are equally important in regulating Ca 2ϩ channels in neurons. However, much less is understood about whether and how a particular type of PP contributes to regulating neuronal Ca 2ϩ channel activities. This is primarily because of the lack of specific inhibitors/activators for different types of PPs, particularly the PP2c family. The functional roles of PP2c and its substrates in the brain remain virtually unknown. During our yeast two-hybrid screening, PP2c␣ was pulled out by both N- and P/Q-type Ca 2ϩ channel C termini. This raised the possibility that PP2c␣ might be associated with voltage-gated Ca 2ϩ channels for regulation of the Ca 2ϩ channel activity. Biochemical studies show ϩ that PP2c␣ binds directly to neuronal Ca 2 channels forming a functional protein complex in vivo. PP2c␣, unlike PP1, PP2a and PP2b, is more effective in dephosphorylation of neuronal Ca 2ϩ channels after their phosphorylation by PKC. In hippocampal neurons, disruption of the PP2c␣–Ca 2ϩ channel interaction significantly enhances the response of Ca 2ϩ channels to modulation by PKC. -

Faqs for DNA End Modification: Dephosphorylation
FAQs for DNA End Modification: Dephosphorylation 1. Does the DNA need to be purified after a restriction digest and prior to the dephosphorylation step? If the enzymes used in the restriction digest are heat inactivatable, then a purification step is not needed. If the enzymes used are not heat-inactivatable, then a clean-up step between the restriction digest and the dephosphorylation step is recommended. 2. Does the DNA need to be purified after the dephosphorylation step and prior to the ligation step? If you use rSAP or Antarctic Phosphatase, which are heat-inactivatable, then a clean- up step is not necessary. If the alkaline phosphatase used is CIP, which is not heat- inactivatable, then a clean-up step is necessary. 3. Which alkaline phosphatase, CIP, Antarctic or rSAP, works best? rSAP (NEB #M0371) is a superior choice because it combines the advantages of both CIP (NEB #M0290) and Antarctic Phosphatase (NEB #M0289). rSAP has a very high specific activity comparable to CIP, but CIP cannot be completely heat-inactivated. rSAP is heat-inactivated in 5 minutes, as is Antarctic Phosphatase, but rSAP is preferred over Antarctic Phosphatase because it does not require added zinc or any other co-factors. As a result, rSAP can be added directly to restriction enzyme digests. Additionally, after heat-inactivation of rSAP, it is not necessary to purify vector DNA prior to the ligation reaction. 4. Are the alkaline phosphatases active in NEBuffers? rSAP: rSAP is active in all restriction enzyme NEBuffers 1.1, 2.1, 3.1 and CutSmart™ Buffer, and can be added directly to digested DNA. -

Dephosphorylation of Threonine-14 and Tyrosine-15
Proc. Natl. Acad. Sci. USA Vol. 90, pp. 3521-3524, April 1993 Biochemistry Cdc25M2 activation of cyclin-dependent kinases by dephosphorylation of threonine-14 and tyrosine-15 BYRON SEBASTIAN*t, AKIRA KAKIZUKAt, AND TONY HUNTER* *Molecular Biology and Virology Laboratory and tGene Expression Laboratory, The Salk Institute for Biological Studies, P.O. Box 85800, San Diego, CA 92186; and tDepartment of Biology, University of California at San Diego, La Jolla, CA 92093 Communicated by Renato Dulbecco, January 4, 1993 (receivedfor review December 15, 1992) ABSTRACT Recent evidence has suggested that human in regulating entry into S phase is suggested by reports that cyclin-dependent kinase 2 (CDK2) is an essential regulator of cycin E-associated kinase activity peaks in G1 (23, 24) and cell cycle progression through S phase. CDK2 is known to that overexpression ofcyclin E decreases the length ofG1 and complex with at least two distinct human cyclins, E and A. The diminishes the dependency of proliferating human cells on kinase activity of these complexes peaks in G1 and S phase, growth factors (23). Although both CDK2 and CDC2 asso- respectively. The vertebrate CDC2/cyclin Bi complex is an ciate with cyclin E, the predominant complex appears to be essential regulator of the onset of mitosis and is inhibited by CDK2/cyclin E. A role for CDK2/cyclin A in regulating phosphorylation of CDC2 on Thr-14 and Tyr-15. In vitro, progression through S phase is suggested by observations CDC2/cyclin Bi is activated by treatment with the members of that microinjection of either anti-cyclin A antibodies or the Cdc25 family of phosphatases. -

P53 Acetylation: Regulation and Consequences
Cancers 2015, 7, 30-69; doi:10.3390/cancers7010030 OPEN ACCESS cancers ISSN 2072-6694 www.mdpi.com/journal/cancers Review p53 Acetylation: Regulation and Consequences Sara M. Reed 1,2 and Dawn E. Quelle 1,2,3,* 1 Department of Pharmacology, The University of Iowa Carver College of Medicine, Iowa City, IA 52242, USA; E-Mail: [email protected] 2 Medical Scientist Training Program, The University of Iowa Carver College of Medicine, Iowa City, IA 52242, USA 3 Department of Pathology, The University of Iowa Carver College of Medicine, Iowa City, IA 52242, USA * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +1-319-353-5749; Fax: +1-319-335-8930. Academic Editor: Rebecca S. Hartley Received: 18 March 2014 / Accepted: 12 December 2014 / Published: 23 December 2014 Abstract: Post-translational modifications of p53 are critical in modulating its tumor suppressive functions. Ubiquitylation, for example, plays a major role in dictating p53 stability, subcellular localization and transcriptional vs. non-transcriptional activities. Less is known about p53 acetylation. It has been shown to govern p53 transcriptional activity, selection of growth inhibitory vs. apoptotic gene targets, and biological outcomes in response to diverse cellular insults. Yet recent in vivo evidence from mouse models questions the importance of p53 acetylation (at least at certain sites) as well as canonical p53 functions (cell cycle arrest, senescence and apoptosis) to tumor suppression. This review discusses the cumulative findings regarding p53 acetylation, with a focus on the acetyltransferases that modify p53 and the mechanisms regulating their activity. We also evaluate what is known regarding the influence of other post-translational modifications of p53 on its acetylation, and conclude with the current outlook on how p53 acetylation affects tumor suppression. -

AMP-Activated Protein Kinase: the Current Landscape for Drug Development
REVIEWS AMP-activated protein kinase: the current landscape for drug development Gregory R. Steinberg 1* and David Carling2 Abstract | Since the discovery of AMP-activated protein kinase (AMPK) as a central regulator of energy homeostasis, many exciting insights into its structure, regulation and physiological roles have been revealed. While exercise, caloric restriction, metformin and many natural products increase AMPK activity and exert a multitude of health benefits, developing direct activators of AMPK to elicit beneficial effects has been challenging. However, in recent years, direct AMPK activators have been identified and tested in preclinical models, and a small number have entered clinical trials. Despite these advances, which disease(s) represent the best indications for therapeutic AMPK activation and the long-term safety of such approaches remain to be established. Cardiovascular disease Dramatic improvements in health care coupled with identifying a unifying mechanism that could link these (CVD). A term encompassing an increased standard of living, including better nutri- changes to multiple branches of metabolism followed diseases affecting the heart tion and education, have led to a remarkable increase in the discovery that the AMP-activated protein kinase or circulatory system. human lifespan1. Importantly, the number of years spent (AMPK) provided a common regulatory mechanism in good health is also increasing2. Despite these positive for inhibiting both cholesterol (through phosphoryla- Non-alcoholic fatty liver disease developments, there are substantial risks that challenge tion of HMG-CoA reductase (HMGR)) and fatty acid (NAFLD). A very common continued improvements in human health. Perhaps the (through phosphorylation of acetyl-CoA carboxylase disease in humans in which greatest threat to future health is a chronic energy imbal- (ACC)) synthesis8 (BOx 1). -

Protein Acetylation at the Interface of Genetics, Epigenetics and Environment in Cancer
H OH metabolites OH Review Protein Acetylation at the Interface of Genetics, Epigenetics and Environment in Cancer Mio Harachi 1, Kenta Masui 1,* , Webster K. Cavenee 2, Paul S. Mischel 3 and Noriyuki Shibata 1 1 Department of Pathology, Division of Pathological Neuroscience, Tokyo Women’s Medical University, Tokyo 162-8666, Japan; [email protected] (M.H.); [email protected] (N.S.) 2 Ludwig Institute for Cancer Research, University of California San Diego, La Jolla, CA 92093, USA; [email protected] 3 Department of Pathology, Stanford University School of Medicine, Stanford, CA 94305, USA; [email protected] * Correspondence: [email protected]; Tel.: +81-3-3353-8111; Fax: +81-3-5269-7408 Abstract: Metabolic reprogramming is an emerging hallmark of cancer and is driven by abnormalities of oncogenes and tumor suppressors. Accelerated metabolism causes cancer cell aggression through the dysregulation of rate-limiting metabolic enzymes as well as by facilitating the production of intermediary metabolites. However, the mechanisms by which a shift in the metabolic landscape reshapes the intracellular signaling to promote the survival of cancer cells remain to be clarified. Recent high-resolution mass spectrometry-based proteomic analyses have spotlighted that, unex- pectedly, lysine residues of numerous cytosolic as well as nuclear proteins are acetylated and that this modification modulates protein activity, sublocalization and stability, with profound impact on cellular function. More importantly, cancer cells exploit acetylation as a post-translational protein for microenvironmental adaptation, nominating it as a means for dynamic modulation of the phenotypes of cancer cells at the interface between genetics and environments. -

Control of Smad7 Stability by Competition Between Acetylation and Ubiquitination
Molecular Cell, Vol. 10, 483–493, September, 2002, Copyright 2002 by Cell Press Control of Smad7 Stability by Competition between Acetylation and Ubiquitination Eva Gro¨ nroos, Ulf Hellman, Carl-Henrik Heldin, 1998), where it binds the receptors and inhibits further and Johan Ericsson1 signaling by at least two different mechanisms. First, Ludwig Institute for Cancer Research Smad7 is able to interfere with TGF signaling by Box 595 blocking the interactions between the R-Smads and the Husargatan 3 activated receptors (Hayashi et al., 1997; Nakao et al., S-751 24 Uppsala 1997). Second, Smad7 interacts with the E3-ubiquitin Sweden ligases Smurf1 or Smurf2 in the nucleus; after TGF stimulation, the Smad7-Smurf complex translocates from the nucleus to the plasma membrane, where Smurf Summary induces ubiquitination and degradation of the TGF re- ceptors (Ebisawa et al., 2001; Kavsak et al., 2000). Smad proteins regulate gene expression in response Acetylation is a dynamic posttranslational modifica- to TGF signaling. Here we present evidence that tion of lysine residues. Proteins with intrinsic histone Smad7 interacts with the transcriptional coactivator acetyltransferase (HAT) activity act as transcriptional p300, resulting in acetylation of Smad7 on two lysine coactivators by acetylating histones and thereby induce residues in its N terminus. Acetylation or mutation of an open chromatin conformation, which allows the tran- these lysine residues stabilizes Smad7 and protects scriptional machinery access to promoters (Roth et al., it from TGF-induced degradation. Furthermore, we 2001). The best characterized HATs are p300, CBP, and demonstrate that the acetylated residues in Smad7 P/CAF (Roth et al., 2001).