Binding Specificities of Human RNA Binding Proteins Towards Structured and Linear RNA
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Doctoral Thesis Genetics of Male Infertility
DOCTORAL THESIS GENETICS OF MALE INFERTILITY: MOLECULAR STUDY OF NON-SYNDROMIC CRYPTORCHIDISM AND SPERMATOGENIC IMPAIRMENT Deborah Grazia Lo Giacco November 2013 Genetics of male infertility: molecular study of non-syndromic cryptorchidism and spermatogenic impairment Thesis presented by Deborah Grazia Lo Giacco To fulfil the PhD degree at Universitat Autònoma de Barcelona Thesis realized under the direction of Dr. Elisabet Ars Criach and Prof. Csilla Krausz at the laboratory of Molecular Biology of Fundació Puigvert, Barcelona Thesis ascribed to the Department of Cellular Biology, Physiology and Immunology, Medicine School of Universitat Autònoma de Barcelona PhD in Cellular Biology Dr. Elisabet Ars Criach Prof. Csilla Krausz Dr. Carme Nogués Sanmiquel Director of the thesis Director of the thesis Tutor of the thesis Deborah Grazia Lo Giacco Ph.D Candidate A mis padres Agradecimientos Esta tesis es un esfuerzo en el cual, directa o indirectamente, han participado varias personas, leyendo, opinando, corrigiendo, teniéndo paciencia, dando ánimo, acompañando en los momentos de crisis y en los momentos de felicidad. Antes de todo quisiera agradecer a mis directoras de tesis, la Dra Csilla Krausz y la Dra Elisabet Ars, por su dedicación costante y continua a este trabajo de investigación y por sus observaciones y siempre acertados consejos. Gracias por haber sido mentores y amigas, gracias por transmitirme vuestro entusiasmo y por todo lo que he aprendido de vosotras. Mi más profundo agradecimiento a la Sra Esperança Marti por haber creído en el valor de nuestro trabajo y haber hecho que fuera posible. Quisiera agradecer al Dr. Eduard Ruiz-Castañé por su apoyo y ayuda constante, y a todos los médicos del Servicio de Andrología: Dr. -

Whole-Genome Microarray Detects Deletions and Loss of Heterozygosity of Chromosome 3 Occurring Exclusively in Metastasizing Uveal Melanoma
Anatomy and Pathology Whole-Genome Microarray Detects Deletions and Loss of Heterozygosity of Chromosome 3 Occurring Exclusively in Metastasizing Uveal Melanoma Sarah L. Lake,1 Sarah E. Coupland,1 Azzam F. G. Taktak,2 and Bertil E. Damato3 PURPOSE. To detect deletions and loss of heterozygosity of disease is fatal in 92% of patients within 2 years of diagnosis. chromosome 3 in a rare subset of fatal, disomy 3 uveal mela- Clinical and histopathologic risk factors for UM metastasis noma (UM), undetectable by fluorescence in situ hybridization include large basal tumor diameter (LBD), ciliary body involve- (FISH). ment, epithelioid cytomorphology, extracellular matrix peri- ϩ ETHODS odic acid-Schiff-positive (PAS ) loops, and high mitotic M . Multiplex ligation-dependent probe amplification 3,4 5 (MLPA) with the P027 UM assay was performed on formalin- count. Prescher et al. showed that a nonrandom genetic fixed, paraffin-embedded (FFPE) whole tumor sections from 19 change, monosomy 3, correlates strongly with metastatic death, and the correlation has since been confirmed by several disomy 3 metastasizing UMs. Whole-genome microarray analy- 3,6–10 ses using a single-nucleotide polymorphism microarray (aSNP) groups. Consequently, fluorescence in situ hybridization were performed on frozen tissue samples from four fatal dis- (FISH) detection of chromosome 3 using a centromeric probe omy 3 metastasizing UMs and three disomy 3 tumors with Ͼ5 became routine practice for UM prognostication; however, 5% years’ metastasis-free survival. to 20% of disomy 3 UM patients unexpectedly develop metas- tases.11 Attempts have therefore been made to identify the RESULTS. Two metastasizing UMs that had been classified as minimal region(s) of deletion on chromosome 3.12–15 Despite disomy 3 by FISH analysis of a small tumor sample were found these studies, little progress has been made in defining the key on MLPA analysis to show monosomy 3. -

Entrez Symbols Name Termid Termdesc 117553 Uba3,Ube1c
Entrez Symbols Name TermID TermDesc 117553 Uba3,Ube1c ubiquitin-like modifier activating enzyme 3 GO:0016881 acid-amino acid ligase activity 299002 G2e3,RGD1310263 G2/M-phase specific E3 ubiquitin ligase GO:0016881 acid-amino acid ligase activity 303614 RGD1310067,Smurf2 SMAD specific E3 ubiquitin protein ligase 2 GO:0016881 acid-amino acid ligase activity 308669 Herc2 hect domain and RLD 2 GO:0016881 acid-amino acid ligase activity 309331 Uhrf2 ubiquitin-like with PHD and ring finger domains 2 GO:0016881 acid-amino acid ligase activity 316395 Hecw2 HECT, C2 and WW domain containing E3 ubiquitin protein ligase 2 GO:0016881 acid-amino acid ligase activity 361866 Hace1 HECT domain and ankyrin repeat containing, E3 ubiquitin protein ligase 1 GO:0016881 acid-amino acid ligase activity 117029 Ccr5,Ckr5,Cmkbr5 chemokine (C-C motif) receptor 5 GO:0003779 actin binding 117538 Waspip,Wip,Wipf1 WAS/WASL interacting protein family, member 1 GO:0003779 actin binding 117557 TM30nm,Tpm3,Tpm5 tropomyosin 3, gamma GO:0003779 actin binding 24779 MGC93554,Slc4a1 solute carrier family 4 (anion exchanger), member 1 GO:0003779 actin binding 24851 Alpha-tm,Tma2,Tmsa,Tpm1 tropomyosin 1, alpha GO:0003779 actin binding 25132 Myo5b,Myr6 myosin Vb GO:0003779 actin binding 25152 Map1a,Mtap1a microtubule-associated protein 1A GO:0003779 actin binding 25230 Add3 adducin 3 (gamma) GO:0003779 actin binding 25386 AQP-2,Aqp2,MGC156502,aquaporin-2aquaporin 2 (collecting duct) GO:0003779 actin binding 25484 MYR5,Myo1e,Myr3 myosin IE GO:0003779 actin binding 25576 14-3-3e1,MGC93547,Ywhah -

Molecular Mechanisms Underlying Noncoding Risk Variations in Psychiatric Genetic Studies
OPEN Molecular Psychiatry (2017) 22, 497–511 www.nature.com/mp REVIEW Molecular mechanisms underlying noncoding risk variations in psychiatric genetic studies X Xiao1,2, H Chang1,2 and M Li1 Recent large-scale genetic approaches such as genome-wide association studies have allowed the identification of common genetic variations that contribute to risk architectures of psychiatric disorders. However, most of these susceptibility variants are located in noncoding genomic regions that usually span multiple genes. As a result, pinpointing the precise variant(s) and biological mechanisms accounting for the risk remains challenging. By reviewing recent progresses in genetics, functional genomics and neurobiology of psychiatric disorders, as well as gene expression analyses of brain tissues, here we propose a roadmap to characterize the roles of noncoding risk loci in the pathogenesis of psychiatric illnesses (that is, identifying the underlying molecular mechanisms explaining the genetic risk conferred by those genomic loci, and recognizing putative functional causative variants). This roadmap involves integration of transcriptomic data, epidemiological and bioinformatic methods, as well as in vitro and in vivo experimental approaches. These tools will promote the translation of genetic discoveries to physiological mechanisms, and ultimately guide the development of preventive, therapeutic and prognostic measures for psychiatric disorders. Molecular Psychiatry (2017) 22, 497–511; doi:10.1038/mp.2016.241; published online 3 January 2017 RECENT GENETIC ANALYSES OF NEUROPSYCHIATRIC neurodevelopment and brain function. For example, GRM3, DISORDERS GRIN2A, SRR and GRIA1 were known to involve in the neuro- Schizophrenia, bipolar disorder, major depressive disorder and transmission mediated by glutamate signaling and synaptic autism are highly prevalent complex neuropsychiatric diseases plasticity. -

Role of CCCH-Type Zinc Finger Proteins in Human Adenovirus Infections
viruses Review Role of CCCH-Type Zinc Finger Proteins in Human Adenovirus Infections Zamaneh Hajikhezri 1, Mahmoud Darweesh 1,2, Göran Akusjärvi 1 and Tanel Punga 1,* 1 Department of Medical Biochemistry and Microbiology, Uppsala University, 75123 Uppsala, Sweden; [email protected] (Z.H.); [email protected] (M.D.); [email protected] (G.A.) 2 Department of Microbiology and Immunology, Al-Azhr University, Assiut 11651, Egypt * Correspondence: [email protected]; Tel.: +46-733-203-095 Received: 28 October 2020; Accepted: 16 November 2020; Published: 18 November 2020 Abstract: The zinc finger proteins make up a significant part of the proteome and perform a huge variety of functions in the cell. The CCCH-type zinc finger proteins have gained attention due to their unusual ability to interact with RNA and thereby control different steps of RNA metabolism. Since virus infections interfere with RNA metabolism, dynamic changes in the CCCH-type zinc finger proteins and virus replication are expected to happen. In the present review, we will discuss how three CCCH-type zinc finger proteins, ZC3H11A, MKRN1, and U2AF1, interfere with human adenovirus replication. We will summarize the functions of these three cellular proteins and focus on their potential pro- or anti-viral activities during a lytic human adenovirus infection. Keywords: human adenovirus; zinc finger protein; CCCH-type; ZC3H11A; MKRN1; U2AF1 1. Zinc Finger Proteins Zinc finger proteins are a big family of proteins with characteristic zinc finger (ZnF) domains present in the protein sequence. The ZnF domains consists of various ZnF motifs, which are short 30–100 amino acid sequences, coordinating zinc ions (Zn2+). -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

RNA-Binding Proteins in Human Oogenesis: Balancing Differentiation and Self-Renewal in the Female Fetal Germline
Stem Cell Research 21 (2017) 193–201 Contents lists available at ScienceDirect Stem Cell Research journal homepage: www.elsevier.com/locate/scr RNA-binding proteins in human oogenesis: Balancing differentiation and self-renewal in the female fetal germline Roseanne Rosario a, Andrew J. Childs b, Richard A. Anderson a,⁎ a MRC Centre for Reproductive Health, Queen's Medical Research Institute, University of Edinburgh, 47 Little France Crescent, Edinburgh EH16 4TJ, UK b Department of Comparative Biomedical Sciences, The Royal Veterinary College, London NW1 0TU, UK article info abstract Article history: Primordial germ cells undergo three significant processes on their path to becoming primary oocytes: the initia- Received 7 October 2016 tion of meiosis, the formation and breakdown of germ cell nests, and the assembly of single oocytes into primor- Received in revised form 29 March 2017 dial follicles. However at the onset of meiosis, the germ cell becomes transcriptionally silenced. Consequently Accepted 13 April 2017 translational control of pre-stored mRNAs plays a central role in coordinating gene expression throughout the re- Available online 18 April 2017 mainder of oogenesis; RNA binding proteins are key to this regulation. In this review we examine the role of ex- Keywords: emplars of such proteins, namely LIN28, DAZL, BOLL and FMRP, and highlight how their roles during germ cell Germ cell differentiation development are critical to oogenesis and the establishment of the primordial follicle pool. RNA binding proteins © 2017 The Authors. -

A Role for Dazl in Commitment to Gametogenic Fate in Embryonic Germ Cells of C57BL/6 Mice
A Role for Dazl in Commitment to Gametogenic Fate in Embryonic Germ Cells of C57BL/6 Mice by Yanfeng Lin (Yen-Hong Lim) B.S. Biochemistry and Molecular and Cell Biology University of Wisconsin-Madison, 1998 SUBMITTED TO THE DEPARTMENT OF BIOLOGY IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY IN BIOLOGY AT THE MASSACHUSETTS INSTITUTE OF TECHNOLOGY SEPT 2005 C 2005 Yanfeng Lin. All rights reserved. The author hereby grants to MIT permission to reproduce and distribute publicly paper and electronic copies of this thesis document in whole or in part. Signature of Author __ _ __ Department of Biology August, 2005 V /-' ~J-2 Certified by David C. Page Professor of Biology Howard Hughes Medical Institute Thesis Supervisor Accepted b) Stephen P. Bell Co-chair, Biology Graduate Student Committee 'MACHUS ETT S NS1 I OF TECHNOLOGY ARCHIVES' OCT 0 2005 . ,. -~ I LIBRARIES .i. __ A Role for Dazl in Commitment to Gametogenic Fate in Embryonic Germ Cells of C57BL/6 Mice by Yanfeng Lin (Yen-Hong Lim) Submitted to the Department of Biology on June, 2005 in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy in Biology Abstract Germ cells can be defined as the cells that undergo the terminal differentiating process of meiosis. In mice, as XX germ cells enter meiosis around Embryonic days 13.5-14.5 (E13.5-E14.5), they form meiotic figures and down-regulate pluripotency markers. XY germ cells enter proliferation arrest between E13.5 and E16.5, which is accompanied by a distinct morphological change as well. -
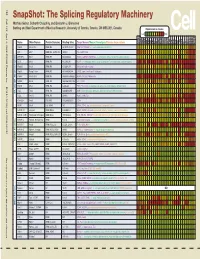
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

Mir‑128‑3P Serves As an Oncogenic Microrna in Osteosarcoma Cells by Downregulating ZC3H12D
ONCOLOGY LETTERS 21: 152, 2021 miR‑128‑3p serves as an oncogenic microRNA in osteosarcoma cells by downregulating ZC3H12D MAOSHU ZHU1*, YULONG WU2*, ZHAOWEI WANG3, MINGHUA LIN4, BIN SU5, CHUNYANG LI6, FULONG LIANG7 and XINJIANG CHEN6 Departments of 1Central Laboratory, 2Urinary Surgery, 3Gynecology, 4Pathology, 5Pharmacy, 6Orthopedics and 7Neurology, The Fifth Hospital of Xiamen, Xiang'an, Xiamen, Fujian 361000, P.R. China Received August 30, 2019; Accepted November 24, 2020 DOI: 10.3892/ol.2020.12413 Abstract. Osteosarcoma is the second leading cause of miR‑128‑3p‑mediated molecular pathway and how it is associ‑ cancer‑associated mortality worldwide in children and ated with osteosarcoma development. adolescents. ZC3H12D has been shown to negatively regulate Toll‑like receptor signaling and serves as a possible tumor Introduction suppressor gene. MicroRNAs (miRNAs/miRs) are known to play an important role in the proliferation of human osteo‑ As the most commonly occurring primary malignant bone sarcoma cells. However, whether miRNAs can affect tumor tumor, osteosarcoma accounts for >10% of all solid tumors development by regulating the expression of ZC3H12D has worldwide (1‑3). Osteosarcoma is the second leading cause of not yet been investigated. The aim of the present study was cancer‑associated mortality worldwide and primarily affects to investigate the role of miR128‑3p in regulating ZC3H12D young individuals, including children and adolescents (4,5). expression, as well as its function in tumor cell proliferation, Owing to advancements in surgical technology and combined apoptosis, and metastasis. Reverse transcription‑quantitative therapeutic strategies, the 5‑year overall survival rate of osteo‑ PCR, western blotting and dual luciferase reporter assays were sarcoma has increased to 60‑70% (6). -

Expression of DAZL Gene in Selected Tissues and Association of Its Polymorphisms with Testicular Size in Hu Sheep
animals Article Expression of DAZL Gene in Selected Tissues and Association of Its Polymorphisms with Testicular Size in Hu Sheep 1, 1, 1 1,2 1 1, Zehu Yuan y, Jing Luo y, Li Wang , Fadi Li , Wanhong Li and Xiangpeng Yue * 1 State Key Laboratory of Grassland Agro-Ecosystems, Key Laboratory of Grassland Livestock Industry Innovation, Ministry of Agriculture and Rural Affairs, Engineering Research Center of Grassland Industry, Ministry of Education, College of Pastoral Agriculture Science and Technology, Lanzhou University, Lanzhou 730020, China; [email protected] (Z.Y.); [email protected] (J.L.); [email protected] (L.W.); [email protected] (F.L.); [email protected] (W.L.) 2 Engineering Laboratory of Sheep Breeding and Reproduction Biotechnology in Gansu Province, Minqin 733300, China * Correspondence: [email protected] These authors contributed equally to this work. y Received: 9 March 2020; Accepted: 20 April 2020; Published: 23 April 2020 Simple Summary: The deleted in azoospermia-like (DAZL) is an RNA binding protein coding gene in autosomal, playing important roles in testicular development and gametogenesis. In this paper, we found that DAZL is extremely highly expressed in testis compared with other organs and reaches to a peak at sex maturity (6-month old) in testis. Two single nucleotide polymorphisms (SNPs) within DAZL were found to have significant effect on the variation coefficient between left and right epididymis weight. Abstract: The deleted in azoospermia-like (DAZL) gene encoding an RNA binding protein is pivotal in gametogenesis in lots of species and also acts as a pre-meiosis marker. -

Binding Specificities of Human RNA Binding Proteins Towards Structured
bioRxiv preprint doi: https://doi.org/10.1101/317909; this version posted March 1, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Binding specificities of human RNA binding proteins towards structured and linear 2 RNA sequences 3 4 Arttu Jolma1,#, Jilin Zhang1,#, Estefania Mondragón4,#, Teemu Kivioja2, Yimeng Yin1, 5 Fangjie Zhu1, Quaid Morris5,6,7,8, Timothy R. Hughes5,6, Louis James Maher III4 and Jussi 6 Taipale1,2,3,* 7 8 9 AUTHOR AFFILIATIONS 10 11 1Department of Medical Biochemistry and Biophysics, Karolinska Institutet, Solna, Sweden 12 2Genome-Scale Biology Program, University of Helsinki, Helsinki, Finland 13 3Department of Biochemistry, University of Cambridge, Cambridge, United Kingdom 14 4Department of Biochemistry and Molecular Biology and Mayo Clinic Graduate School of 15 Biomedical Sciences, Mayo Clinic College of Medicine and Science, Rochester, USA 16 5Department of Molecular Genetics, University of Toronto, Toronto, Canada 17 6Donnelly Centre, University of Toronto, Toronto, Canada 18 7Edward S Rogers Sr Department of Electrical and Computer Engineering, University of 19 Toronto, Toronto, Canada 20 8Department of Computer Science, University of Toronto, Toronto, Canada 21 #Authors contributed equally 22 *Correspondence: [email protected] 23 24 25 SUMMARY 26 27 Sequence specific RNA-binding proteins (RBPs) control many important 28 processes affecting gene expression. They regulate RNA metabolism at multiple 29 levels, by affecting splicing of nascent transcripts, RNA folding, base modification, 30 transport, localization, translation and stability. Despite their central role in most 31 aspects of RNA metabolism and function, most RBP binding specificities remain 32 unknown or incompletely defined.