Engineering an Improved Light-Induced Dimer (Ilid) for Controlling the Localization and Activity of Signaling Proteins
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

291533611.Pdf
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Publications of the IAS Fellows Genome-wide identification, classification, evolutionary expansion and expression analyses of homeobox genes in rice Mukesh Jain, Akhilesh K. Tyagi and Jitendra P. Khurana Interdisciplinary Centre for Plant Genomics and Department of Plant Molecular Biology, University of Delhi South Campus, India Keywords Homeobox genes play a critical role in regulating various aspects of plant abiotic stress; homeobox genes; microarray growth and development. In the present study, we identified a total of 107 analysis; reproductive development; rice homeobox genes in the rice genome and grouped them into ten distinct (Oryza sativa) subfamilies based upon their domain composition and phylogenetic analy- Correspondence sis. A significantly large number of homeobox genes are located in the J. P. Khurana, Department of Plant duplicated segments of the rice genome, which suggests that the expansion Molecular Biology, University of Delhi South of homeobox gene family, in large part, might have occurred due to Campus, Benito Juarez Road, New Delhi segmental duplications in rice. Furthermore, microarray analysis was 110021, India performed to elucidate the expression profiles of these genes in different Fax: +91 011 24115270 tissues and during various stages of vegetative and reproductive develop- Tel: +91 011 24115126 ment. Several genes with predominant expression during various stages of E-mail: [email protected] panicle and seed development were identified. At least 37 homeobox genes (Received 6 November 2007, revised 3 were found to be differentially expressed significantly (more than two-fold; March 2008, accepted 31 March 2008) P < 0.05) under various abiotic stress conditions. -

Supplemental Table S1
Entrez Gene Symbol Gene Name Affymetrix EST Glomchip SAGE Stanford Literature HPA confirmed Gene ID Profiling profiling Profiling Profiling array profiling confirmed 1 2 A2M alpha-2-macroglobulin 0 0 0 1 0 2 10347 ABCA7 ATP-binding cassette, sub-family A (ABC1), member 7 1 0 0 0 0 3 10350 ABCA9 ATP-binding cassette, sub-family A (ABC1), member 9 1 0 0 0 0 4 10057 ABCC5 ATP-binding cassette, sub-family C (CFTR/MRP), member 5 1 0 0 0 0 5 10060 ABCC9 ATP-binding cassette, sub-family C (CFTR/MRP), member 9 1 0 0 0 0 6 79575 ABHD8 abhydrolase domain containing 8 1 0 0 0 0 7 51225 ABI3 ABI gene family, member 3 1 0 1 0 0 8 29 ABR active BCR-related gene 1 0 0 0 0 9 25841 ABTB2 ankyrin repeat and BTB (POZ) domain containing 2 1 0 1 0 0 10 30 ACAA1 acetyl-Coenzyme A acyltransferase 1 (peroxisomal 3-oxoacyl-Coenzyme A thiol 0 1 0 0 0 11 43 ACHE acetylcholinesterase (Yt blood group) 1 0 0 0 0 12 58 ACTA1 actin, alpha 1, skeletal muscle 0 1 0 0 0 13 60 ACTB actin, beta 01000 1 14 71 ACTG1 actin, gamma 1 0 1 0 0 0 15 81 ACTN4 actinin, alpha 4 0 0 1 1 1 10700177 16 10096 ACTR3 ARP3 actin-related protein 3 homolog (yeast) 0 1 0 0 0 17 94 ACVRL1 activin A receptor type II-like 1 1 0 1 0 0 18 8038 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 1 0 0 0 0 19 8751 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 1 0 0 0 0 20 8728 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 1 0 0 0 0 21 81792 ADAMTS12 ADAM metallopeptidase with thrombospondin type 1 motif, 12 1 0 0 0 0 22 9507 ADAMTS4 ADAM metallopeptidase with thrombospondin type 1 -

The PAS Domain Confers Target Gene Specificity of Drosophila Bhlh/PAS Proteins
Downloaded from genesdev.cshlp.org on October 3, 2021 - Published by Cold Spring Harbor Laboratory Press The PAS domain confers target gene specificity of Drosophila bHLH/PAS proteins Elazar Zelzer, Pablo Wappner, and Ben-Zion Shilo1 Department of Molecular Genetics, Weizmann Institute of Science, Rehovot 76100, Israel Trachealess (Trh) and Single-minded (Sim) are highly similar Drosophila bHLH/PAS transcription factors. They activate nonoverlapping target genes and induce diverse cell fates. A single Drosophila gene encoding a bHLH/PAS protein homologous to the vertebrate ARNT protein was isolated and may serve as a partner for both Trh and Sim. We show that Trh and Sim complexes recognize similar DNA-binding sites in the embryo. To examine the basis for their distinct target gene specificity, the activity of Trh–Sim chimeric proteins was monitored in embryos. Replacement of the Trh PAS domain by the analogous region of Sim was sufficient to convert it into a functional Sim protein. The PAS domain thus mediates all the features conferring specificity and the distinct recognition of target genes. The normal expression pattern of additional proteins essential for the activity of the Trh or Sim complexes can be inferred from the induction pattern of target genes and binding-site reporters, triggered by ubiquitous expression of Trh or Sim. We postulate that the capacity of bHLH/PAS heterodimers to associate, through the PAS domain, with additional distinct proteins that bind target-gene DNA, is essential to confer specificity. [Key Words: Gene expression; bHLH/PAS; Trachealess; Single minded; HIF1a; ARNT; trachea; midline] Received February 20, 1997; revised version accepted July 1, 1997. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Of Antigen Receptor-Driven T Cells Hypoxia-Inducible Factor Regulates Survival
Hypoxia-Inducible Factor Regulates Survival of Antigen Receptor-Driven T Cells Yuichi Makino, Hiroshi Nakamura, Eiji Ikeda, Kei Ohnuma, Kenji Yamauchi, Yutaka Yabe, Lorenz Poellinger, Yasunori This information is current as Okada, Chikao Morimoto and Hirotoshi Tanaka of September 28, 2021. J Immunol 2003; 171:6534-6540; ; doi: 10.4049/jimmunol.171.12.6534 http://www.jimmunol.org/content/171/12/6534 Downloaded from References This article cites 46 articles, 17 of which you can access for free at: http://www.jimmunol.org/content/171/12/6534.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 28, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2003 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Hypoxia-Inducible Factor Regulates Survival of Antigen Receptor-Driven T Cells1 ‡,Kenji Yamauchi ء,Eiji Ikeda,† Kei Ohnuma ء,Hiroshi Nakamura ء,Yuichi Makino and ء,Yutaka Yabe,‡ Lorenz Poellinger,§ Yasunori Okada,† Chikao Morimoto ءHirotoshi Tanaka2 Peripheral T lymphocytes undergo activation by antigenic stimulation and function in hypoxic areas of inflammation. -

Molecular Evolution of PAS Domain-Containing Proteins of Filamentous Cyanobacteria Through Domain Shuffling and Domain Duplication
DNA Research 11, 69–81 (2004) Molecular Evolution of PAS Domain-Containing Proteins of Filamentous Cyanobacteria Through Domain Shuffling and Domain Duplication Rei Narikawa, Shinobu Okamoto, Masahiko Ikeuchi,∗ and Masayuki Ohmori Department of Life Sciences (Biology), Graduate School of Arts and Sciences, University of Tokyo, Komaba, Meguro-ku, Tokyo 153-8902, Japan (Received 15 December 2003; revised 11 March 2004) Downloaded from https://academic.oup.com/dnaresearch/article/11/2/69/534432 by guest on 28 September 2021 Abstract When the entire genome of a filamentous heterocyst-forming N2-fixing cyanobacterium, Anabaena sp. PCC 7120 (Anabaena) was determined in 2001, a large number of PAS domains were detected in signal-transducing proteins. The draft genome sequence is also available for the cyanobacterium, Nostoc punctiforme strain ATCC 29133 (Nostoc), that is closely related to Anabaena. In this study, we extracted all PAS domains from the Nostoc genome sequence and analyzed them together with those of Anabaena. Clustering analysis of all the PAS domains gave many specific pairings, indicative of evolutionary conser- vations. Ortholog analysis of PAS-containing proteins showed composite multidomain architecture in some cases of conserved domains and domains of disagreement between the two species. Further inspection of the domains of disagreement allowed us to trace them back in evolution. Thus, multidomain proteins could have been generated by duplication or shuffling in these cyanobacteria. The conserved PAS domains in the orthologous proteins were analyzed by structural fitting to the known PAS domains. We detected several subclasses with unique sequence features, which will be the target of experimental analysis. Key words: cyanobacterium; PAS domain; domain shuffling; ortholog pair; molecular evolution 12,13 1. -
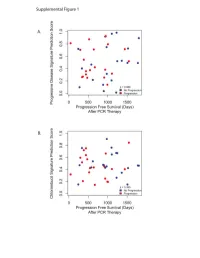
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

Comprehensive Computational Analysis of Leucine-Rich Repeat (LRR) Proteins Encoded in the Genome of the Diatom Phaeodactylum Tricornutum
Erschienen in: Marine Genomics ; 21 (2015). - S. 43-51 https://dx.doi.org/10.1016/j.margen.2015.02.007 Comprehensive computational analysis of leucine-rich repeat (LRR) proteins encoded in the genome of the diatom Phaeodactylum tricornutum Birgit Schulze ⁎, Matthias T. Buhmann 1, Carolina Río Bártulos, Peter G. Kroth Department of Biological Sciences, Universität Konstanz, 78457 Konstanz, Germany abstract We have screened the genome of the marine diatom Phaeodactylum tricornutum for gene models encoding proteins exhibiting leucine rich repeat (LRR) structures. In order to reveal the functionality of these proteins, their amino acid sequences were scanned for known domains and for homologies to other proteins. Additionally, proteins were categorized into different LRR families according to the variable sequence part of their LRR. This approach enabled us to group proteins with potentially similar functionality and to classify also LRR proteins Keywords: where no characterized homologues in other organisms exist. Most interestingly, we were able to indentify LRR several transmembrane LRR proteins, which are likely to function as receptor like molecules. However, none Receptor-like protein of them carry additional domains that are typical for mammalian or plant like receptors. Thus, the respective Adhesion protein signal recognition pathways seem to be substantially different in diatoms. Moreover, P. tricornutum encodes a RanGAP family of secreted LRR proteins likely to function as adhesion or binding proteins as part of the extracellular matrix. Additionally, intracellular LRR only proteins were divided into proteins similar to RasGTPase activators, regulators of nuclear transport, and mitotic regulation. Our approach allowed us to draw a detailed picture of the conservation and diversification of LRR proteins in the marine diatom P. -

Supplementary Table 1: Significantly
Supplementary Table 1: Significantly upregulated genes in the “malignant” cluster of PNETs (p<0.05) Fold change Gene (WDEC/WDET) FEV protein 11.61 adenylate cyclase 2 (brain) 4.47 nuclear receptor subfamily 4, group A, member 2 4.45 growth arrest and DNA-damage-inducible, beta 3.28 plexin B1 3.10 neuronal PAS domain protein 2 2.92 caldesmon 1 2.81 drebrin 1 2.43 hypothetical protein BC013764 2.41 TGFB-induced factor (TALE family homeobox) 2.26 Homo sapiens cDNA FLJ12843 fis, clone NT2RP2003293, mRNA sequence 2.17 nuclear prelamin A recognition factor 2.15 chromobox homolog 4 (Pc class homolog, Drosophila) 2.14 RuvB (E coli homolog)-like 1 [Homo sapiens], mRNA sequence 2.03 latent transforming growth factor beta binding protein 4 2.02 putative glialblastoma cell differentiation-related 1.92 hypothetical protein FLJ10298 1.92 protein tyrosine phosphatase type IVA, member 1 1.90 ADP-ribosylation factor-like 4 1.88 NIMA (never in mitosis gene a)-related kinase 7 1.87 RAB11B, member RAS oncogene family 1.87 myelin transcription factor 1 1.86 centaurin, beta 2 1.83 Homo sapiens cDNA FLJ38575 fis, clone HCHON2007046, mRNA sequence 1.83 SH3 domain binding glutamic acid-rich protein like 3 1.81 immunoglobulin heavy constant gamma 3 (G3m marker) 1.77 chromobox homolog 3 (HP1 gamma homolog, Drosophila) 1.74 basic transcription element binding protein 1 1.73 H1 histone family, member X 1.73 protein phosphatase 2A, regulatory subunit B' (PR 53) 1.73 Homo sapiens cDNA FLJ30096 fis, clone BNGH41000045, mRNA sequence 1.73 KIAA0397 gene product 1.71 OK/SW-CL.4 [Homo sapiens], mRNA sequence 1.69 high-mobility group 20B 1.69 phosphoserine phosphatase 1.68 Homo sapiens mRNA; cDNA DKFZp434B142 (from clone DKFZp434B142), mRNA sequence 1.67 nuclear mitotic apparatus protein 1 1.67 hypothetical protein 628 1.66 PRKC, apoptosis, WT1, regulator 1.64 E3 ubiquitin ligase SMURF1 1.64 chromodomain protein, Y chromosome-like 1.63 hypothetical protein FLJ21839 1.62 hypothetical protein MGC2550 1.62 Suppl. -

NPAS1-ARNT and NPAS3-ARNT Crystal Structures Implicate The
RESEARCH ARTICLE NPAS1-ARNT and NPAS3-ARNT crystal structures implicate the bHLH-PAS family as multi-ligand binding transcription factors Dalei Wu1†, Xiaoyu Su1, Nalini Potluri1, Youngchang Kim2, Fraydoon Rastinejad1* 1Integrative Metabolism Program, Sanford Burnham Prebys Medical Discovery Institute, Orlando, United States; 2Structural Biology Center, Biosciences Division, Argonne National Laboratory, Argonne, United States Abstract The neuronal PAS domain proteins NPAS1 and NPAS3 are members of the basic helix- loop-helix-PER-ARNT-SIM (bHLH-PAS) family, and their genetic deficiencies are linked to a variety of human psychiatric disorders including schizophrenia, autism spectrum disorders and bipolar disease. NPAS1 and NPAS3 must each heterodimerize with the aryl hydrocarbon receptor nuclear translocator (ARNT), to form functional transcription complexes capable of DNA binding and gene regulation. Here we examined the crystal structures of multi-domain NPAS1-ARNT and NPAS3- ARNT-DNA complexes, discovering each to contain four putative ligand-binding pockets. Through expanded architectural comparisons between these complexes and HIF-1a-ARNT, HIF-2a-ARNT *For correspondence: and CLOCK-BMAL1, we show the wider mammalian bHLH-PAS family is capable of multi-ligand- [email protected] binding and presents as an ideal class of transcription factors for direct targeting by small-molecule Present address: †State Key drugs. Laboratory of Microbial DOI: 10.7554/eLife.18790.001 Technology, School of Life Sciences, Shandong University, Qingdao, China Competing interests: The Introduction authors declare that no The mammalian bHLH-PAS transcription factors share a common protein architecture consisting of a competing interests exist. conserved bHLH DNA-binding domain, tandem PAS domains (PAS-A and PAS-B), and a variable activation or repression domain. -

(PAS) Kinase in the Stimulation of Preproinsulin and Pancreatic Duodenum Homeobox 1 Gene Expression by Glucose
Involvement of Per–Arnt–Sim (PAS) kinase in the stimulation of preproinsulin and pancreatic duodenum homeobox 1 gene expression by glucose Gabriela da Silva Xavier*, Jared Rutter†, and Guy A. Rutter*‡ *Henry Wellcome Signalling Laboratories and Department of Biochemistry, University of Bristol, Bristol BS8 ITD, United Kingdom; and †Department of Biochemistry, University of Utah, Salt Lake City, UT 83132 Edited by Donald F. Steiner, University of Chicago, Chicago, IL, and approved April 15, 2004 (received for review November 21, 2003) Per–Arnt–Sim (PAS) domain-containing kinases are common in PASK is also regulated by glucose in  cells and may play a prokaryotes, but a mammalian counterpart has only recently been complementary role in the regulation of gene expression. described. Although the PAS domain of the mammalian PAS kinase (PASK) is closely related to the bacterial oxygen sensor FixL, it is Materials and Methods unclear whether PASK activity is changed in mammalian cells in Materials. The silencer small interfering RNA (siRNA) construc- response to nutrients and might therefore contribute to signal tion kit was from Ambion (Austin, TX). siRNA oligonucleotides transduction by these or other stimuli. Here, we show that ele- were from Cruachem (Herndon, VA). TransIT-TKO transfec- vated glucose concentrations rapidly increase PASK activity in tion reagent was from Mirus (Madison, WI), human growth pancreatic islet  cells, an event followed by the accumulation of hormone (hGH) ELISA kit was from Roche Diagnostics, and rat both PASK mRNA and protein. Demonstrating a physiological role insulin radioimmunoassay kit was from Linco Research Immu- for PASK activation, comicroinjection into clonal  cells of cDNA noassay (St. -

Identifying and Characterizing Yeast PAS Kinase 1 Substrates Reveals Regulation of Mitochondrial and Cell Growth Pathways
Brigham Young University BYU ScholarsArchive Theses and Dissertations 2015-06-01 Identifying and Characterizing Yeast PAS Kinase 1 Substrates Reveals Regulation of Mitochondrial and Cell Growth Pathways Desiree DeMille Brigham Young University - Provo Follow this and additional works at: https://scholarsarchive.byu.edu/etd Part of the Microbiology Commons BYU ScholarsArchive Citation DeMille, Desiree, "Identifying and Characterizing Yeast PAS Kinase 1 Substrates Reveals Regulation of Mitochondrial and Cell Growth Pathways" (2015). Theses and Dissertations. 5930. https://scholarsarchive.byu.edu/etd/5930 This Dissertation is brought to you for free and open access by BYU ScholarsArchive. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of BYU ScholarsArchive. For more information, please contact [email protected], [email protected]. Identifying and Characterizing Yeast PAS Kinase 1 Substrates Reveals Regulation of Mitochondrial and Cell Growth Pathways Desiree DeMille A dissertation submitted to the faculty of Brigham Young University in partial fulfillment of the requirements for the degree of Doctor of Philosophy Julianne H. Grose, Chair William R. McCleary Laura C. Bridgewater David M. Thomson Steven M. Johnson Department of Microbiology & Molecular Biology Brigham Young University June 2015 Copyright © 2015 Desiree DeMille All Rights Reserved ABSTRACT Identifying and Characterizing Yeast PAS Kinase 1 Substrates Reveals Regulation of Mitochondrial and Cell Growth Pathways Desiree