Structure and Function of Per-ARNT-Sim Domains and Their Possible Role in the Life-Cycle Biology of Trypanosoma Cruzi
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

291533611.Pdf
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Publications of the IAS Fellows Genome-wide identification, classification, evolutionary expansion and expression analyses of homeobox genes in rice Mukesh Jain, Akhilesh K. Tyagi and Jitendra P. Khurana Interdisciplinary Centre for Plant Genomics and Department of Plant Molecular Biology, University of Delhi South Campus, India Keywords Homeobox genes play a critical role in regulating various aspects of plant abiotic stress; homeobox genes; microarray growth and development. In the present study, we identified a total of 107 analysis; reproductive development; rice homeobox genes in the rice genome and grouped them into ten distinct (Oryza sativa) subfamilies based upon their domain composition and phylogenetic analy- Correspondence sis. A significantly large number of homeobox genes are located in the J. P. Khurana, Department of Plant duplicated segments of the rice genome, which suggests that the expansion Molecular Biology, University of Delhi South of homeobox gene family, in large part, might have occurred due to Campus, Benito Juarez Road, New Delhi segmental duplications in rice. Furthermore, microarray analysis was 110021, India performed to elucidate the expression profiles of these genes in different Fax: +91 011 24115270 tissues and during various stages of vegetative and reproductive develop- Tel: +91 011 24115126 ment. Several genes with predominant expression during various stages of E-mail: [email protected] panicle and seed development were identified. At least 37 homeobox genes (Received 6 November 2007, revised 3 were found to be differentially expressed significantly (more than two-fold; March 2008, accepted 31 March 2008) P < 0.05) under various abiotic stress conditions. -

Supplemental Table S1
Entrez Gene Symbol Gene Name Affymetrix EST Glomchip SAGE Stanford Literature HPA confirmed Gene ID Profiling profiling Profiling Profiling array profiling confirmed 1 2 A2M alpha-2-macroglobulin 0 0 0 1 0 2 10347 ABCA7 ATP-binding cassette, sub-family A (ABC1), member 7 1 0 0 0 0 3 10350 ABCA9 ATP-binding cassette, sub-family A (ABC1), member 9 1 0 0 0 0 4 10057 ABCC5 ATP-binding cassette, sub-family C (CFTR/MRP), member 5 1 0 0 0 0 5 10060 ABCC9 ATP-binding cassette, sub-family C (CFTR/MRP), member 9 1 0 0 0 0 6 79575 ABHD8 abhydrolase domain containing 8 1 0 0 0 0 7 51225 ABI3 ABI gene family, member 3 1 0 1 0 0 8 29 ABR active BCR-related gene 1 0 0 0 0 9 25841 ABTB2 ankyrin repeat and BTB (POZ) domain containing 2 1 0 1 0 0 10 30 ACAA1 acetyl-Coenzyme A acyltransferase 1 (peroxisomal 3-oxoacyl-Coenzyme A thiol 0 1 0 0 0 11 43 ACHE acetylcholinesterase (Yt blood group) 1 0 0 0 0 12 58 ACTA1 actin, alpha 1, skeletal muscle 0 1 0 0 0 13 60 ACTB actin, beta 01000 1 14 71 ACTG1 actin, gamma 1 0 1 0 0 0 15 81 ACTN4 actinin, alpha 4 0 0 1 1 1 10700177 16 10096 ACTR3 ARP3 actin-related protein 3 homolog (yeast) 0 1 0 0 0 17 94 ACVRL1 activin A receptor type II-like 1 1 0 1 0 0 18 8038 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 1 0 0 0 0 19 8751 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 1 0 0 0 0 20 8728 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 1 0 0 0 0 21 81792 ADAMTS12 ADAM metallopeptidase with thrombospondin type 1 motif, 12 1 0 0 0 0 22 9507 ADAMTS4 ADAM metallopeptidase with thrombospondin type 1 -

Pin1 Inhibitors: Towards Understanding the Enzymatic
Pin1 Inhibitors: Towards Understanding the Enzymatic Mechanism Guoyan Xu Dissertation submitted to the faculty of the Virginia Polytechnic Institute and State University In the partial fulfillment of the requirement for the degree of Doctor of Philosophy In Chemistry Felicia A Etzkorn, Chair David G. I. Kingston Neal Castagnoli Paul R. Carlier Brian E. Hanson May 6, 2010 Blacksburg, Virginia Keywords: Pin1, anti-cancer drug target, transition-state analogues, ketoamides, ketones, reduced amides, PPIase assay, inhibition Pin1 Inhibitors: Towards Understanding the Enzymatic Mechanism Guoyan Xu Abstract An important role of Pin1 is to catalyze the cis-trans isomerization of pSer/Thr- Pro bonds; as such, it plays an important role in many cellular events through the effects of conformational change on the function of its biological substrates, including Cdc25, c- Jun, and p53. The expression of Pin1 correlates with cyclin D1 levels, which contributes to cancer cell transformation. Overexpression of Pin1 promotes tumor growth, while its inhibition causes tumor cell apoptosis. Because Pin1 is overexpressed in many human cancer tissues, including breast, prostate, and lung cancer tissues, it plays an important role in oncogenesis, making its study vital for the development of anti-cancer agents. Many inhibitors have been discovered for Pin1, including 1) several classes of designed inhibitors such as alkene isosteres, non-peptidic, small molecular Pin1 inhibitors, and indanyl ketones, and 2) several natural products such as juglone, pepticinnamin E analogues, PiB and its derivatives obtained from a library screen. These Pin1 inhibitors show promise in the development of novel diagnostic and therapeutic anticancer drugs due to their ability to block cell cycle progression. -

The PAS Domain Confers Target Gene Specificity of Drosophila Bhlh/PAS Proteins
Downloaded from genesdev.cshlp.org on October 3, 2021 - Published by Cold Spring Harbor Laboratory Press The PAS domain confers target gene specificity of Drosophila bHLH/PAS proteins Elazar Zelzer, Pablo Wappner, and Ben-Zion Shilo1 Department of Molecular Genetics, Weizmann Institute of Science, Rehovot 76100, Israel Trachealess (Trh) and Single-minded (Sim) are highly similar Drosophila bHLH/PAS transcription factors. They activate nonoverlapping target genes and induce diverse cell fates. A single Drosophila gene encoding a bHLH/PAS protein homologous to the vertebrate ARNT protein was isolated and may serve as a partner for both Trh and Sim. We show that Trh and Sim complexes recognize similar DNA-binding sites in the embryo. To examine the basis for their distinct target gene specificity, the activity of Trh–Sim chimeric proteins was monitored in embryos. Replacement of the Trh PAS domain by the analogous region of Sim was sufficient to convert it into a functional Sim protein. The PAS domain thus mediates all the features conferring specificity and the distinct recognition of target genes. The normal expression pattern of additional proteins essential for the activity of the Trh or Sim complexes can be inferred from the induction pattern of target genes and binding-site reporters, triggered by ubiquitous expression of Trh or Sim. We postulate that the capacity of bHLH/PAS heterodimers to associate, through the PAS domain, with additional distinct proteins that bind target-gene DNA, is essential to confer specificity. [Key Words: Gene expression; bHLH/PAS; Trachealess; Single minded; HIF1a; ARNT; trachea; midline] Received February 20, 1997; revised version accepted July 1, 1997. -

Anti-Inflammatory Role of Curcumin in LPS Treated A549 Cells at Global Proteome Level and on Mycobacterial Infection
Anti-inflammatory Role of Curcumin in LPS Treated A549 cells at Global Proteome level and on Mycobacterial infection. Suchita Singh1,+, Rakesh Arya2,3,+, Rhishikesh R Bargaje1, Mrinal Kumar Das2,4, Subia Akram2, Hossain Md. Faruquee2,5, Rajendra Kumar Behera3, Ranjan Kumar Nanda2,*, Anurag Agrawal1 1Center of Excellence for Translational Research in Asthma and Lung Disease, CSIR- Institute of Genomics and Integrative Biology, New Delhi, 110025, India. 2Translational Health Group, International Centre for Genetic Engineering and Biotechnology, New Delhi, 110067, India. 3School of Life Sciences, Sambalpur University, Jyoti Vihar, Sambalpur, Orissa, 768019, India. 4Department of Respiratory Sciences, #211, Maurice Shock Building, University of Leicester, LE1 9HN 5Department of Biotechnology and Genetic Engineering, Islamic University, Kushtia- 7003, Bangladesh. +Contributed equally for this work. S-1 70 G1 S 60 G2/M 50 40 30 % of cells 20 10 0 CURI LPSI LPSCUR Figure S1: Effect of curcumin and/or LPS treatment on A549 cell viability A549 cells were treated with curcumin (10 µM) and/or LPS or 1 µg/ml for the indicated times and after fixation were stained with propidium iodide and Annexin V-FITC. The DNA contents were determined by flow cytometry to calculate percentage of cells present in each phase of the cell cycle (G1, S and G2/M) using Flowing analysis software. S-2 Figure S2: Total proteins identified in all the three experiments and their distribution betwee curcumin and/or LPS treated conditions. The proteins showing differential expressions (log2 fold change≥2) in these experiments were presented in the venn diagram and certain number of proteins are common in all three experiments. -

(12) Patent Application Publication (10) Pub. No.: US 2008/0261923 A1 Etzkorn Et Al
US 20080261923A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2008/0261923 A1 Etzkorn et al. (43) Pub. Date: Oct. 23, 2008 54) ALKENE MIMICS Related U.S. Applicationpp Data (76) Inventors: Felicia A. Etzkorn, Blacksburg, VA (60) Provisional application No. 60/598.421, filed on Aug. (US); Xiaodong X. Wang, 4, 2004. Maricopa, AZ (US); Bulling Xu, Publication Classification Blacksburg, VA (US) (51) Int. Cl. Correspondence Address: A63/675 (2006.01) WHITHAM, CURTIS & CHRISTOFFERSON & C07F 9/06 (2006.01) COOK, PC A6IP35/00 (2006.01) 9 Lew e 11491 SUNSET HILLS ROAD, SUITE 340 A6II 3/662 (2006.01) RESTON, VA 20190 (US) (52) U.S. Cl. ............... 514/80; 546/22: 548/414: 546/23; 548/112:558/166; 514/89: 514/114 (22) PCT Filed: Jul. 29, 2005 Ac-Phe-Tyr-phosphoSer-CH=C-Pro-Arg-NHAND Fmoc-bis(pivaloylmethoxy)phosphoSer-CH=C-Pro-2- (86). PCT No.: PCT/USOS/26821 aminoethyl-(3-indole); and their Phospho-(D)-serine stereoi Somers are novel compounds. I refers to a pseudo amide. S371 (c)(1), Such novel compounds advantageously may be used as alk (2), (4) Date: Sep. 26, 2007 ene mimics. US 2008/0261923 A1 Oct. 23, 2008 ALKENE MIMICS 0005. The possibility of Pin1 activity led to interest and work on certain alkene mimics. (Wang, Supra); Wang, X. J., FIELD OF THE INVENTION Xu, B., Mullins, A. B., Neiler, F.K., and Etzkorn, F.A. (2004), Conformationally Locked Isostere of PhosphoSer-cis-Pro 0001. This invention relates to the design and synthesis of Inhibits Pin1 23-Fold Better than PhosphoSer-trans-Pro Isos compounds that are alkene mimics. -

Role of Protein Repair Enzymes in Oxidative Stress Survival And
Shome et al. Annals of Microbiology (2020) 70:55 Annals of Microbiology https://doi.org/10.1186/s13213-020-01597-2 REVIEW ARTICLE Open Access Role of protein repair enzymes in oxidative stress survival and virulence of Salmonella Arijit Shome1* , Ratanti Sarkhel1, Shekhar Apoorva1, Sonu Sukumaran Nair2, Tapan Kumar Singh Chauhan1, Sanjeev Kumar Bhure1 and Manish Mahawar1 Abstract Purpose: Proteins are the principal biomolecules in bacteria that are affected by the oxidants produced by the phagocytic cells. Most of the protein damage is irreparable though few unfolded proteins and covalently modified amino acids can be repaired by chaperones and repair enzymes respectively. This study reviews the three protein repair enzymes, protein L-isoaspartyl O-methyl transferase (PIMT), peptidyl proline cis-trans isomerase (PPIase), and methionine sulfoxide reductase (MSR). Methods: Published articles regarding protein repair enzymes were collected from Google Scholar and PubMed. The information obtained from the research articles was analyzed and categorized into general information about the enzyme, mechanism of action, and role played by the enzymes in bacteria. Special emphasis was given to the importance of these enzymes in Salmonella Typhimurium. Results: Protein repair is the direct and energetically preferred way of replenishing the cellular protein pool without translational synthesis. Under the oxidative stress mounted by the host during the infection, protein repair becomes very crucial for the survival of the bacterial pathogens. Only a few covalent modifications of amino acids are reversible by the protein repair enzymes, and they are highly specific in activity. Deletion mutants of these enzymes in different bacteria revealed their importance in the virulence and oxidative stress survival. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Of Antigen Receptor-Driven T Cells Hypoxia-Inducible Factor Regulates Survival
Hypoxia-Inducible Factor Regulates Survival of Antigen Receptor-Driven T Cells Yuichi Makino, Hiroshi Nakamura, Eiji Ikeda, Kei Ohnuma, Kenji Yamauchi, Yutaka Yabe, Lorenz Poellinger, Yasunori This information is current as Okada, Chikao Morimoto and Hirotoshi Tanaka of September 28, 2021. J Immunol 2003; 171:6534-6540; ; doi: 10.4049/jimmunol.171.12.6534 http://www.jimmunol.org/content/171/12/6534 Downloaded from References This article cites 46 articles, 17 of which you can access for free at: http://www.jimmunol.org/content/171/12/6534.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 28, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2003 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Hypoxia-Inducible Factor Regulates Survival of Antigen Receptor-Driven T Cells1 ‡,Kenji Yamauchi ء,Eiji Ikeda,† Kei Ohnuma ء,Hiroshi Nakamura ء,Yuichi Makino and ء,Yutaka Yabe,‡ Lorenz Poellinger,§ Yasunori Okada,† Chikao Morimoto ءHirotoshi Tanaka2 Peripheral T lymphocytes undergo activation by antigenic stimulation and function in hypoxic areas of inflammation. -

Molecular Evolution of PAS Domain-Containing Proteins of Filamentous Cyanobacteria Through Domain Shuffling and Domain Duplication
DNA Research 11, 69–81 (2004) Molecular Evolution of PAS Domain-Containing Proteins of Filamentous Cyanobacteria Through Domain Shuffling and Domain Duplication Rei Narikawa, Shinobu Okamoto, Masahiko Ikeuchi,∗ and Masayuki Ohmori Department of Life Sciences (Biology), Graduate School of Arts and Sciences, University of Tokyo, Komaba, Meguro-ku, Tokyo 153-8902, Japan (Received 15 December 2003; revised 11 March 2004) Downloaded from https://academic.oup.com/dnaresearch/article/11/2/69/534432 by guest on 28 September 2021 Abstract When the entire genome of a filamentous heterocyst-forming N2-fixing cyanobacterium, Anabaena sp. PCC 7120 (Anabaena) was determined in 2001, a large number of PAS domains were detected in signal-transducing proteins. The draft genome sequence is also available for the cyanobacterium, Nostoc punctiforme strain ATCC 29133 (Nostoc), that is closely related to Anabaena. In this study, we extracted all PAS domains from the Nostoc genome sequence and analyzed them together with those of Anabaena. Clustering analysis of all the PAS domains gave many specific pairings, indicative of evolutionary conser- vations. Ortholog analysis of PAS-containing proteins showed composite multidomain architecture in some cases of conserved domains and domains of disagreement between the two species. Further inspection of the domains of disagreement allowed us to trace them back in evolution. Thus, multidomain proteins could have been generated by duplication or shuffling in these cyanobacteria. The conserved PAS domains in the orthologous proteins were analyzed by structural fitting to the known PAS domains. We detected several subclasses with unique sequence features, which will be the target of experimental analysis. Key words: cyanobacterium; PAS domain; domain shuffling; ortholog pair; molecular evolution 12,13 1. -
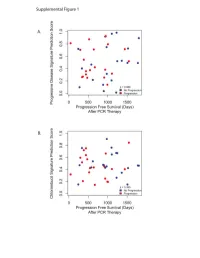
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

Cyclophilins and Other Foldases: Cell Signaling Catalysts and Drug Targets
International Symposium on Cyclophilins and other Foldases: Cell Signaling Catalysts and Drug Targets Halle (Saale), Germany, September 19-21, 2013 CONFERENCE MATERIAL Cyclophilins and other Foldases CONFERENCE VENUE The symposium will be held at the Martin-Luther-University Halle-Wittenberg Lecture hall XXII Universitätsplatz 1 06108 Halle (Saale) Germany Conference office telephone: 0157-30056381 2 Cyclophilins and other Foldases CONFERENCE VENUE LOCATION 3 Cyclophilins and other Foldases ORGANIZING COMMITTEE Jochen Balbach (MLU Halle-Wittenberg) Gunter Fischer (MPI BPC Göttingen, BO Halle) Franz X. Schmid (University of Bayreuth) The conference is supported by SCYNEXIS Inc. Selcia Ltd. AstraZeneca Takeda Pharma Debiopharm Fonds der Chemischen Industrie The City of Halle 4 Cyclophilins and other Foldases WELCOME TO THE CONFERENCE We take great pleasure in setting aside our regular lab work and welcoming you all to the International Conference on Cyclophilins and other Foldases - Cell Signaling Catalysts and Drug Targets, which will be held at the Martin-Luther-University Halle- Wittenberg, Germany from September 19th to 22th, 2013. In the last few years, the scientific and biotechnological communities have become increasingly interested in the challenges involved in conformational regulation of bioactivities by endogenous factors such as foldases. This is reflected in the number of over 600 papers published during 2012 about the enzymes addressed in this conference. We are just beginning to understand how catalyzed conformational interconversions of peptide bonds may affect protein functioning under physiological and pathophysiological conditions. This conference is thought to provide an unique forum for researchers to present and exchange new data and to call attention on all aspects of foldase enzymes and their inhibitors.