SHANK1 Deletions in Males with Autism Spectrum Disorder
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Study of Chemotherapy Resistance in Acute Myeloid Leukemia
A Study of Chemotherapy Resistance in Acute Myeloid Leukemia A DISSERTATION SUBMITTED TO THE FACULTY OF THE GRADUATE SCHOOL OF THE UNIVERSITY OF MINNESOTA BY Susan Kay Rathe IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY David A. Largaespada October, 2013 © Susan Kay Rathe 2013 Acknowledgements Funding: The leukemia research was funded by The Leukemia & Lymphoma Society (grant 7019-04). The MMuFLR workflow was funded by The Children's Cancer Research Fund. University of Minnesota Resources: The BioMedical Genomics Center provided services for gene expression microarray, RNA sequencing, oligo preparation, and Sanger sequencing. The Minnesota Supercomputing Institute maintains the Galaxy Software, as well as provides data management services and training. The Masonic Cancer Center Bioinformatics Core provided guidance in the analysis of the RNA-seq data. The FDA Drug Screen was performed at the Institute for Therapeutics Discovery and Development. Colleagues: First and foremost, I would like to recognize Dr. David Largaespada for his mentorship and the opportunity to continue research he started while a postdoc in Dr. Neal Copeland’s lab. I would also like to thank the many members of the Largaespada lab who provided direct or indirect assistance to my research. Dr. Bin Yin continued Dr. Largaespada’s work on the BXH-2 AML cell lines by making drug resistant derivatives and provided training in cell culture and drug assay techniques. Dr. Won-Il Kim developed the TNM mouse model and provided training in mouse related experimental techniques. Miechaleen Diers did all of the tail vein injections for the AML transplants and assisted with necropsies. -

Tepzz¥ 6Z54za T
(19) TZZ¥ ZZ_T (11) EP 3 260 540 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 27.12.2017 Bulletin 2017/52 C12N 15/113 (2010.01) A61K 9/127 (2006.01) A61K 31/713 (2006.01) C12Q 1/68 (2006.01) (21) Application number: 17000579.7 (22) Date of filing: 12.11.2011 (84) Designated Contracting States: • Sarma, Kavitha AL AT BE BG CH CY CZ DE DK EE ES FI FR GB Philadelphia, PA 19146 (US) GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO • Borowsky, Mark PL PT RO RS SE SI SK SM TR Needham, MA 02494 (US) • Ohsumi, Toshiro Kendrick (30) Priority: 12.11.2010 US 412862 P Cambridge, MA 02141 (US) 20.12.2010 US 201061425174 P 28.07.2011 US 201161512754 P (74) Representative: Clegg, Richard Ian et al Mewburn Ellis LLP (62) Document number(s) of the earlier application(s) in City Tower accordance with Art. 76 EPC: 40 Basinghall Street 11840099.3 / 2 638 163 London EC2V 5DE (GB) (71) Applicant: The General Hospital Corporation Remarks: Boston, MA 02114 (US) •Thecomplete document including Reference Tables and the Sequence Listing can be downloaded from (72) Inventors: the EPO website • Lee, Jeannie T •This application was filed on 05-04-2017 as a Boston, MA 02114 (US) divisional application to the application mentioned • Zhao, Jing under INID code 62. San Diego, CA 92122 (US) •Claims filed after the date of receipt of the divisional application (Rule 68(4) EPC). (54) POLYCOMB-ASSOCIATED NON-CODING RNAS (57) This invention relates to long non-coding RNAs (IncRNAs), libraries of those ncRNAs that bind chromatin modifiers, such as Polycomb Repressive Complex 2, inhibitory nucleic acids and methods and compositions for targeting IncRNAs. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Human Lectins, Their Carbohydrate Affinities and Where to Find Them
biomolecules Review Human Lectins, Their Carbohydrate Affinities and Where to Review HumanFind Them Lectins, Their Carbohydrate Affinities and Where to FindCláudia ThemD. Raposo 1,*, André B. Canelas 2 and M. Teresa Barros 1 1, 2 1 Cláudia D. Raposo * , Andr1 é LAQVB. Canelas‐Requimte,and Department M. Teresa of Chemistry, Barros NOVA School of Science and Technology, Universidade NOVA de Lisboa, 2829‐516 Caparica, Portugal; [email protected] 12 GlanbiaLAQV-Requimte,‐AgriChemWhey, Department Lisheen of Chemistry, Mine, Killoran, NOVA Moyne, School E41 of ScienceR622 Co. and Tipperary, Technology, Ireland; canelas‐ [email protected] NOVA de Lisboa, 2829-516 Caparica, Portugal; [email protected] 2* Correspondence:Glanbia-AgriChemWhey, [email protected]; Lisheen Mine, Tel.: Killoran, +351‐212948550 Moyne, E41 R622 Tipperary, Ireland; [email protected] * Correspondence: [email protected]; Tel.: +351-212948550 Abstract: Lectins are a class of proteins responsible for several biological roles such as cell‐cell in‐ Abstract:teractions,Lectins signaling are pathways, a class of and proteins several responsible innate immune for several responses biological against roles pathogens. such as Since cell-cell lec‐ interactions,tins are able signalingto bind to pathways, carbohydrates, and several they can innate be a immuneviable target responses for targeted against drug pathogens. delivery Since sys‐ lectinstems. In are fact, able several to bind lectins to carbohydrates, were approved they by canFood be and a viable Drug targetAdministration for targeted for drugthat purpose. delivery systems.Information In fact, about several specific lectins carbohydrate were approved recognition by Food by andlectin Drug receptors Administration was gathered for that herein, purpose. plus Informationthe specific organs about specific where those carbohydrate lectins can recognition be found by within lectin the receptors human was body. -

Advances in Prognostic Methylation Biomarkers for Prostate Cancer
cancers Review Advances in Prognostic Methylation Biomarkers for Prostate Cancer 1 1,2 1,2, 1,2, , Dilys Lam , Susan Clark , Clare Stirzaker y and Ruth Pidsley * y 1 Epigenetics Research Laboratory, Genomics and Epigenetics Division, Garvan Institute of Medical Research, Sydney, New South Wales 2010, Australia; [email protected] (D.L.); [email protected] (S.C.); [email protected] (C.S.) 2 St. Vincent’s Clinical School, University of New South Wales, Sydney, New South Wales 2010, Australia * Correspondence: [email protected]; Tel.: +61-2-92958315 These authors have contributed equally. y Received: 22 September 2020; Accepted: 13 October 2020; Published: 15 October 2020 Simple Summary: Prostate cancer is a major cause of cancer-related death in men worldwide. There is an urgent clinical need for improved prognostic biomarkers to better predict the likely outcome and course of the disease and thus inform the clinical management of these patients. Currently, clinically recognised prognostic markers lack sensitivity and specificity in distinguishing aggressive from indolent disease, particularly in patients with localised, intermediate grade prostate cancer. Thus, there is major interest in identifying new molecular biomarkers to complement existing standard clinicopathological markers. DNA methylation is a frequent alteration in the cancer genome and offers potential as a reliable and robust biomarker. In this review, we provide a comprehensive overview of the current state of DNA methylation biomarker studies in prostate cancer prognosis. We highlight advances in this field that have enabled the discovery of novel prognostic genes and discuss the potential of methylation biomarkers for noninvasive liquid-biopsy testing. -
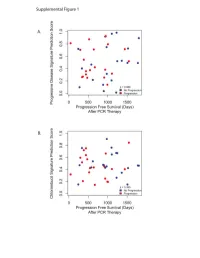
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

Identification of Novel Genes in Human Airway Epithelial Cells Associated with Chronic Obstructive Pulmonary Disease (COPD) Usin
www.nature.com/scientificreports OPEN Identifcation of Novel Genes in Human Airway Epithelial Cells associated with Chronic Obstructive Received: 6 July 2018 Accepted: 7 October 2018 Pulmonary Disease (COPD) using Published: xx xx xxxx Machine-Based Learning Algorithms Shayan Mostafaei1, Anoshirvan Kazemnejad1, Sadegh Azimzadeh Jamalkandi2, Soroush Amirhashchi 3, Seamas C. Donnelly4,5, Michelle E. Armstrong4 & Mohammad Doroudian4 The aim of this project was to identify candidate novel therapeutic targets to facilitate the treatment of COPD using machine-based learning (ML) algorithms and penalized regression models. In this study, 59 healthy smokers, 53 healthy non-smokers and 21 COPD smokers (9 GOLD stage I and 12 GOLD stage II) were included (n = 133). 20,097 probes were generated from a small airway epithelium (SAE) microarray dataset obtained from these subjects previously. Subsequently, the association between gene expression levels and smoking and COPD, respectively, was assessed using: AdaBoost Classifcation Trees, Decision Tree, Gradient Boosting Machines, Naive Bayes, Neural Network, Random Forest, Support Vector Machine and adaptive LASSO, Elastic-Net, and Ridge logistic regression analyses. Using this methodology, we identifed 44 candidate genes, 27 of these genes had been previously been reported as important factors in the pathogenesis of COPD or regulation of lung function. Here, we also identifed 17 genes, which have not been previously identifed to be associated with the pathogenesis of COPD or the regulation of lung function. The most signifcantly regulated of these genes included: PRKAR2B, GAD1, LINC00930 and SLITRK6. These novel genes may provide the basis for the future development of novel therapeutics in COPD and its associated morbidities. -

Abstracts from the 51St European Society of Human Genetics Conference: Electronic Posters
European Journal of Human Genetics (2019) 27:870–1041 https://doi.org/10.1038/s41431-019-0408-3 MEETING ABSTRACTS Abstracts from the 51st European Society of Human Genetics Conference: Electronic Posters © European Society of Human Genetics 2019 June 16–19, 2018, Fiera Milano Congressi, Milan Italy Sponsorship: Publication of this supplement was sponsored by the European Society of Human Genetics. All content was reviewed and approved by the ESHG Scientific Programme Committee, which held full responsibility for the abstract selections. Disclosure Information: In order to help readers form their own judgments of potential bias in published abstracts, authors are asked to declare any competing financial interests. Contributions of up to EUR 10 000.- (Ten thousand Euros, or equivalent value in kind) per year per company are considered "Modest". Contributions above EUR 10 000.- per year are considered "Significant". 1234567890();,: 1234567890();,: E-P01 Reproductive Genetics/Prenatal Genetics then compared this data to de novo cases where research based PO studies were completed (N=57) in NY. E-P01.01 Results: MFSIQ (66.4) for familial deletions was Parent of origin in familial 22q11.2 deletions impacts full statistically lower (p = .01) than for de novo deletions scale intelligence quotient scores (N=399, MFSIQ=76.2). MFSIQ for children with mater- nally inherited deletions (63.7) was statistically lower D. E. McGinn1,2, M. Unolt3,4, T. B. Crowley1, B. S. Emanuel1,5, (p = .03) than for paternally inherited deletions (72.0). As E. H. Zackai1,5, E. Moss1, B. Morrow6, B. Nowakowska7,J. compared with the NY cohort where the MFSIQ for Vermeesch8, A. -

Detailed Characterization of Human Induced Pluripotent Stem Cells Manufactured for Therapeutic Applications
Stem Cell Rev and Rep DOI 10.1007/s12015-016-9662-8 Detailed Characterization of Human Induced Pluripotent Stem Cells Manufactured for Therapeutic Applications Behnam Ahmadian Baghbaderani 1 & Adhikarla Syama2 & Renuka Sivapatham3 & Ying Pei4 & Odity Mukherjee2 & Thomas Fellner1 & Xianmin Zeng3,4 & Mahendra S. Rao5,6 # The Author(s) 2016. This article is published with open access at Springerlink.com Abstract We have recently described manufacturing of hu- help determine which set of tests will be most useful in mon- man induced pluripotent stem cells (iPSC) master cell banks itoring the cells and establishing criteria for discarding a line. (MCB) generated by a clinically compliant process using cord blood as a starting material (Baghbaderani et al. in Stem Cell Keywords Induced pluripotent stem cells . Embryonic stem Reports, 5(4), 647–659, 2015). In this manuscript, we de- cells . Manufacturing . cGMP . Consent . Markers scribe the detailed characterization of the two iPSC clones generated using this process, including whole genome se- quencing (WGS), microarray, and comparative genomic hy- Introduction bridization (aCGH) single nucleotide polymorphism (SNP) analysis. We compare their profiles with a proposed calibra- Induced pluripotent stem cells (iPSCs) are akin to embryonic tion material and with a reporter subclone and lines made by a stem cells (ESC) [2] in their developmental potential, but dif- similar process from different donors. We believe that iPSCs fer from ESC in the starting cell used and the requirement of a are likely to be used to make multiple clinical products. We set of proteins to induce pluripotency [3]. Although function- further believe that the lines used as input material will be used ally identical, iPSCs may differ from ESC in subtle ways, at different sites and, given their immortal status, will be used including in their epigenetic profile, exposure to the environ- for many years or even decades. -

Association Mapping Based on a Common-Garden Migration Experiment Reveals
G3: Genes|Genomes|Genetics Early Online, published on July 9, 2019 as doi:10.1534/g3.119.400369 Association mapping based on a common-garden migration experiment reveals candidate genes for migration tendency in brown trout Alexandre Lemopoulos1,2,*, Silva Uusi-Heikkilä2,3, Pekka Hyvärinen4, Nico Alioravainen1, Jenni M. Prokkola1,5, Chris K. Elvidge1, Anti Vasemägi2,6,7,‡, Anssi Vainikka1, ‡ 1. Department of Environmental and Biological Sciences, University of Eastern Finland, P.O. Box 111, FI-80101 Joensuu, Finland. 2. Department of Biology, University of Turku, FI- 20014, Turku, Finland. 3. Department of Biological and Environmental Science, University of Jyväskylä, P.O. Box 35, FI-40014, Jyväskylä, Finland. 4. Natural Resources Institute Finland, Manamansalontie 90, FI-88300, Paltamo, Finland. 5. Institute of Integrative Biology, University of Liverpool, Bioscience building, Crown street, L69 7BZ Liverpool, UK 6. Department of Aquaculture, Institute of Veterinary Medicine and Animal Sciences, Estonian University of Life Sciences, 51014 Tartu, Estonia. 7. Swedish University of Agricultural Sciences, Department of Aquatic Resources, Institute of Freshwater Research, 17893 Drottningholm, Sweden ‡ Shared senior authorship *Corresponding author: [email protected] Key words: Life-history strategies, RADseq, GWAS, salmonids © The Author(s) 2013. Published by the Genetics Society of America. Abstract A better understanding of the environmental and genetic contribution to migratory behavior and the evolution of traits linked to migration is crucial for fish conservation and fisheries management. Up to date, a few genes with unequivocal influence on the adoption of alternative migration strategies have been identified in salmonids. Here, we used a common garden set-up to measure individual migration distances of generally highly polymorphic brown trout Salmo trutta from two populations. -

Identification of DNA Methylation Associated Gene Signatures in Endometrial Cancer Via Integrated Analysis of DNA Methylation and Gene Expression Systematically
J Gynecol Oncol. 2017 Nov;28(6):e83 https://doi.org/10.3802/jgo.2017.28.e83 pISSN 2005-0380·eISSN 2005-0399 Original Article Identification of DNA methylation associated gene signatures in endometrial cancer via integrated analysis of DNA methylation and gene expression systematically Chuandi Men ,1,2 Hongjuan Chai ,1 Xumin Song ,1 Yue Li ,1 Huawen Du ,1 Qing Ren 1 1Department of Gynecology and Obstetrics, Shanghai Ninth People's Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China Received: May 11, 2017 2Graduate School, Bengbu Medical College, Bengbu, China Revised: Aug 2, 2017 Accepted: Aug 10, 2017 Correspondence to ABSTRACT Qing Ren Department of Gynecology and Obstetrics, Objective: Endometrial cancer (EC) is a common gynecologic cancer worldwide. However, Shanghai Ninth People's Hospital, Shanghai the pathogenesis of EC has not been epigenetically elucidated. Here, this study aims Jiao Tong University School of Medicine, No. to describe the DNA methylation profile and identify favorable gene signatures highly 280, Mohe Road, Baoshan District, Shanghai 201900, China. associated with aberrant DNA methylation changes in EC. E-mail: [email protected] Methods: The data regarding DNA methylation and gene expression were downloaded from The Cancer Genome Atlas (TCGA) database. Differentially methylated CpG sites (DMCs), Copyright © 2017. Asian Society of differentially methylated regions (DMRs), and differentially expressed genes (DEGs) were Gynecologic Oncology, Korean Society of Gynecologic Oncology identified, and the relationship between the 2 omics was further analyzed. In addition, This is an Open Access article distributed weighted CpG site co-methylation network (WCCN) was constructed followed by an under the terms of the Creative Commons integrated analysis of DNA methylation and gene expression data. -

Deciphering the Molecular Profile of Plaques, Memory Decline And
ORIGINAL RESEARCH ARTICLE published: 16 April 2014 AGING NEUROSCIENCE doi: 10.3389/fnagi.2014.00075 Deciphering the molecular profile of plaques, memory decline and neuron loss in two mouse models for Alzheimer’s disease by deep sequencing Yvonne Bouter 1†,Tim Kacprowski 2,3†, Robert Weissmann4, Katharina Dietrich1, Henning Borgers 1, Andreas Brauß1, Christian Sperling 4, Oliver Wirths 1, Mario Albrecht 2,5, Lars R. Jensen4, Andreas W. Kuss 4* andThomas A. Bayer 1* 1 Division of Molecular Psychiatry, Georg-August-University Goettingen, University Medicine Goettingen, Goettingen, Germany 2 Department of Bioinformatics, Institute of Biometrics and Medical Informatics, University Medicine Greifswald, Greifswald, Germany 3 Department of Functional Genomics, Interfaculty Institute for Genetics and Functional Genomics, University Medicine Greifswald, Greifswald, Germany 4 Human Molecular Genetics, Department for Human Genetics of the Institute for Genetics and Functional Genomics, Institute for Human Genetics, University Medicine Greifswald, Ernst-Moritz-Arndt University Greifswald, Greifswald, Germany 5 Institute for Knowledge Discovery, Graz University of Technology, Graz, Austria Edited by: One of the central research questions on the etiology of Alzheimer’s disease (AD) is the Isidro Ferrer, University of Barcelona, elucidation of the molecular signatures triggered by the amyloid cascade of pathological Spain events. Next-generation sequencing allows the identification of genes involved in disease Reviewed by: Isidro Ferrer, University of Barcelona, processes in an unbiased manner. We have combined this technique with the analysis of Spain two AD mouse models: (1) The 5XFAD model develops early plaque formation, intraneu- Dietmar R. Thal, University of Ulm, ronal Ab aggregation, neuron loss, and behavioral deficits. (2)TheTg4–42 model expresses Germany N-truncated Ab4–42 and develops neuron loss and behavioral deficits albeit without plaque *Correspondence: formation.