The Mismatch Repair and Meiotic Recombination Endonuclease Mlh1-Mlh3 Is Activated by Polymer Formation and Can Cleave DNA Substrates in Trans
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

An RNA Holliday Junction? Structural and Dynamic Considerations of the Bacteriophage T4 Gene 60 Interruption
J. Mol. Biol. (1990) 213, 199-201 COMMUNICATIONS An RNA Holliday Junction? Structural and Dynamic Considerations of the Bacteriophage T4 Gene 60 Interruption Donald H. Burke-Agiiero and John E. Hearst Department of Chemistry University of California, Laboratory of Chemical Biodynamics Lawrence Berkeley Laboratory, Berkeley, CA 94720, U.S.A. (Received 20 June 1988; accepted 20 November 1989) A novel interrupted gene motif has been reported in which a 50-nucleotide insertion into bacteriophage T4 gene 60 appears to be present in the message at the time of translation, yet it is not translated. We present here a dynamic model for how translation may be occurring in the neighborhood of the interruption. The model involves formation of an RNA structure with similarities to a Holliday junction, followed by migration of the branch point in a strand exchange between message and interruption. The advantage of this model over previous ones is that at no time is a new tRNA required to pair with a discontinuous template. Huang et al. (1988) have reported a novel gene tion; the order of events is not important to this interruption motif in gene 60 of bacteriophage T4, model. More extensive branch migration is hindered in which a 50-nucleotide intervening segment is by the lack of sequence homology between the neither removed nor translated. The sequence of the message and the dumbbell outside these five bases. interruption suggests that it can be folded to bring When tRNA G~y moves into the P site, the template the translated codons on either side of the inter- 3' to Gly46 will now be in position to receive the ruption into close proximity (Fig. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Holliday Junction Resolvases
Downloaded from http://cshperspectives.cshlp.org/ on September 23, 2021 - Published by Cold Spring Harbor Laboratory Press Holliday Junction Resolvases Haley D.M. Wyatt and Stephen C. West London Research Institute, Cancer Research UK, Clare Hall Laboratories, South Mimms, Herts EN6 3LD, United Kingdom Correspondence: [email protected] Four-way DNA intermediates, called Holliday junctions (HJs), can form during meiotic and mitotic recombination, and their removal is crucial for chromosome segregation. A group of ubiquitous and highly specialized structure-selective endonucleases catalyze the cleavage of HJs into two disconnected DNA duplexes in a reaction called HJ resolution. These enzymes, called HJ resolvases, have been identified in bacteria and their bacteriophages, archaea, and eukaryotes. In this review, we discuss fundamental aspects of the HJ structure and their interaction with junction-resolving enzymes. This is followed by a brief discussion of the eubacterial RuvABC enzymes, which provide the paradigm for HJ resolvases in other organisms. Finally, we review the biochemical and structural properties of some well-char- acterized resolvases from archaea, bacteriophage, and eukaryotes. omologous recombination (HR) is an es- homologous strand as a template for DNA syn- Hsential process that promotes genetic di- thesis. Recombination then proceeds in one of versity during meiosis (see Lam and Keeney several different ways, some of which involve 2014; Zickler and Kleckner 2014). However, in second-end capture, such that the other resect- somatic cells, HR plays a key role in conserv- ed 30 end anneals to the displaced strand of the ing genetic information by facilitating DNA re- D-loop (Szostak et al. -

A Mutation in the Putative MLH3 Endonuclease Domain Confers a Defect in Both Mismatch Repair and Meiosis in Saccharomyces Cerevisiae
Copyright Ó 2008 by the Genetics Society of America DOI: 10.1534/genetics.108.086645 A Mutation in the Putative MLH3 Endonuclease Domain Confers a Defect in Both Mismatch Repair and Meiosis in Saccharomyces cerevisiae K. T. Nishant, Aaron J. Plys and Eric Alani1 Department of Molecular Biology and Genetics, Cornell University, Ithaca, New York 14853-2703 Manuscript received January 2, 2008 Accepted for publication March 20, 2008 ABSTRACT Interference-dependent crossing over in yeast and mammalian meioses involves the mismatch repair protein homologs MSH4-MSH5 and MLH1-MLH3. The MLH3 protein contains a highly conserved metal- binding motif DQHA(X)2E(X)4E that is found in a subset of MLH proteins predicted to have endonuclease activities (Kadyrov et al. 2006). Mutations within this motif in human PMS2 and Saccharomyces cerevisiae PMS1 disrupted the endonuclease and mismatch repair activities of MLH1-PMS2 and MLH1-PMS1, re- spectively (Kadyrov et al. 2006, 2007; Erdeniz et al. 2007). As a first step in determining whether such an activity is required during meiosis, we made mutations in the MLH3 putative endonuclease domain motif (-D523N, -E529K) and found that single and double mutations conferred mlh3-null-like defects with respect to meiotic spore viability and crossing over. Yeast two-hybrid and chromatography analyses showed that the interaction between MLH1 and mlh3-D523N was maintained, suggesting that the mlh3-D523N mutation did not disrupt the stability of MLH3. The mlh3-D523N mutant also displayed a mutator phenotype in vegetative growth that was similar to mlh3D. Overexpression of this allele conferred a dominant-negative phenotype with respect to mismatch repair. -

Insights Into Holliday Junction Resolution and Chromosome Segregation in Yeast
September 17, 2012 SCIENCE SPOTLIGHT Insights into Holliday Junction Resolution and Chromosome Segregation in Yeast September 17, 2012 ME Arnegard Homologous DNA recombination is the highly conserved process by which similar or identical nucleotide sequences are exchanged between two DNA molecules. In all eukaryotes – from yeast to humans – this kind of genetic recombination serves two critical functions during meiosis, the class of cell division that leads to the formation of gametes such as sperm and eggs in the case of humans, or spores in yeast: Homologous recombination promotes the genetic diversity of gametes by allowing crossing-over between homologous chromosomes, and it is also critical to accurate chromosome segregation during meiosis. Anomalies in recombination often result in the complete loss of chromosomes during meiosis, or in chromosomal rearrangements such as deletions or translocations, which in turn can lead to birth defects or cancer. All current models for crossing-over involve two key intermediate structures: DNA double strand breaks (DSBs) and Holliday junctions (HJs). DSBs are the initiating events in crossing-over that are caused by Rec12 in fission yeast, whereas HJs are the crossed-strand structures by which DNA molecules from two homologous chromosomes become intertwined (see figure). At the Fred Hutchinson Cancer Research Center, the laboratory of Dr. Gerald Smith (Basic Sciences Division) has made many advances toward understanding a complex pathway composed of dozens of proteins that promote homologous recombination in fission yeast (Schizosaccharomyces pombe) by moving or pairing chromosomes, or forming or repairing DSBs. In a previous study, for example, Cromie et al. (2006) provided strong evidence that HJs in fission yeast are 'resolved' (i.e., precisely cut and reconnected into two independent helices) solely by the endonucleolytic activity of the partner proteins Mus81 and Eme1. -

E2013080118.Full.Pdf
Replication-independent instability of Friedreich’s ataxia GAA repeats during chronological aging Alexander J. Neila,1, Julia A. Hiseya,1, Ishtiaque Quasema, Ryan J. McGintya, Marcin Hitczenkob, Alexandra N. Khristicha, and Sergei M. Mirkina,2 aDepartment of Biology, Tufts University, Medford, MA 02155; and bFederal Reserve Bank of Atlanta, Atlanta, GA 30309 Edited by Sue Jinks-Robertson, Duke University School of Medicine, Durham, NC, and approved December 28, 2020 (received for review June 25, 2020) Nearly 50 hereditary diseases result from the inheritance of abnor- For example, in the case of Huntington’s disease (HD), an in-frame mally long repetitive DNA microsatellites. While it was originally (CAG)n repeat causes the buildup of disruptive polyglutamine believed that the size of inherited repeats is the key factor in protein aggregates in the brain with age (7, 8). In FRDA, where the disease development, it has become clear that somatic instability of causative repetitive element is noncoding, it has been hypothesized these repeats throughout an individual’s lifetime strongly contrib- that frataxin deficiency drives disease progression in a similar utes to disease onset and progression. Importantly, somatic insta- fashion via accumulation of toxic iron due to the dysregulation of bility is commonly observed in terminally differentiated, postmitotic iron homeostasis (9). An emerging alternative to the toxicity hy- cells, such as neurons. To unravel the mechanisms of repeat insta- pothesis is that the primary driver of symptom onset and progres- bility in nondividing cells, we created an experimental system to sion could be age-related, somatic expansions of the inherited ’ analyze the mutability of Friedreich s ataxia (GAA)n repeats during disease-size microsatellite. -
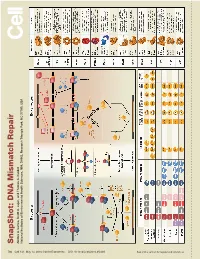
S Na P S H O T: D N a Mism a Tc H R E P a Ir
SnapShot: Repair DNA Mismatch Scott A. Lujan, and Thomas Kunkel A. Larrea, Andres Park, NC 27709, USA Triangle Health Sciences, NIH, DHHS, Research National Institutes of Environmental 730 Cell 141, May 14, 2010 ©2010 Elsevier Inc. DOI 10.1016/j.cell.2010.05.002 See online version for legend and references. SnapShot: DNA Mismatch Repair Andres A. Larrea, Scott A. Lujan, and Thomas A. Kunkel National Institutes of Environmental Health Sciences, NIH, DHHS, Research Triangle Park, NC 27709, USA Mismatch Repair in Bacteria and Eukaryotes Mismatch repair in the bacterium Escherichia coli is initiated when a homodimer of MutS binds as an asymmetric clamp to DNA containing a variety of base-base and insertion-deletion mismatches. The MutL homodimer then couples MutS recognition to the signal that distinguishes between the template and nascent DNA strands. In E. coli, the lack of adenine methylation, catalyzed by the DNA adenine methyltransferase (Dam) in newly synthesized GATC sequences, allows E. coli MutH to cleave the nascent strand. The resulting nick is used for mismatch removal involving the UvrD helicase, single-strand DNA-binding protein (SSB), and excision by single-stranded DNA exonucleases from either direction, depending upon the polarity of the nick relative to the mismatch. DNA polymerase III correctly resynthesizes DNA and ligase completes repair. In bacteria lacking Dam/MutH, as in eukaryotes, the signal for strand discrimination is uncertain but may be the DNA ends associated with replication forks. In these bacteria, MutL harbors a nick-dependent endonuclease that creates a nick that can be used for mismatch excision. Eukaryotic mismatch repair is similar, although it involves several dif- ferent MutS and MutL homologs: MutSα (MSH2/MSH6) recognizes single base-base mismatches and 1–2 base insertion/deletions; MutSβ (MSH2/MSH3) recognizes insertion/ deletion mismatches containing two or more extra bases. -

Coding and Noncoding Variants in HFM1, MLH3, MSH4, MSH5, RNF212, and RNF212B Affect Recombination Rate in Cattle
Downloaded from genome.cshlp.org on September 29, 2021 - Published by Cold Spring Harbor Laboratory Press Research Coding and noncoding variants in HFM1, MLH3, MSH4, MSH5, RNF212, and RNF212B affect recombination rate in cattle Naveen Kumar Kadri,1 Chad Harland,1,2 Pierre Faux,1 Nadine Cambisano,1,3 Latifa Karim,1,3 Wouter Coppieters,1,3 Sébastien Fritz,4,5 Erik Mullaart,6 Denis Baurain,7 Didier Boichard,5 Richard Spelman,2 Carole Charlier,1 Michel Georges,1 and Tom Druet1 1Unit of Animal Genomics, GIGA-R & Faculty of Veterinary Medicine, University of Liège (B34), 4000 Liège, Belgium; 2Livestock Improvement Corporation, Newstead, 3240 Hamilton, New Zealand; 3Genomics Platform, GIGA, University of Liège (B34), 4000 Liège, Belgium; 4Allice, 75012 Paris, France; 5GABI, INRA, AgroParisTech, Université Paris-Saclay, 78350 Jouy-en-Josas, France; 6CRV BV, 6800 AL Arnhem, the Netherlands; 7InBioS-Eukaryotic Phylogenomics, Department of Life Sciences and PhytoSYSTEMS, University of Liège (B22), 4000 Liège, Belgium We herein study genetic recombination in three cattle populations from France, New Zealand, and the Netherlands. We identify 2,395,177 crossover (CO) events in 94,516 male gametes, and 579,996 CO events in 25,332 female gametes. The average number of COs was found to be larger in males (23.3) than in females (21.4). The heritability of global recombi- nation rate (GRR) was estimated at 0.13 in males and 0.08 in females, with a genetic correlation of 0.66 indicating that shared variants are influencing GRR in both sexes. A genome-wide association study identified seven quantitative trait loci (QTL) for GRR. -

Working with Molecular Genetics Chapter 8. Recombination of DNA CHAPTER 8 RECOMBINATION of DNA
Working with Molecular Genetics Chapter 8. Recombination of DNA CHAPTER 8 RECOMBINATION OF DNA The previous chapter on mutation and repair of DNA dealt mainly with small changes in DNA sequence, usually single base pairs, resulting from errors in replication or damage to DNA. The DNA sequence of a chromosome can change in large segments as well, by the processes of recombination and transposition. Recombination is the production of new DNA molecule(s) from two parental DNA molecules or different segments of the same DNA molecule; this will be the topic of this chapter. Transposition is a highly specialized form of recombination in which a segment of DNA moves from one location to another, either on the same chromosome or a different chromosome; this will be discussed in the next chapter. Types and examples of recombination At least four types of naturally occurring recombination have been identified in living organisms (Fig. 8.1). General or homologous recombination occurs between DNA molecules of very similar sequence, such as homologous chromosomes in diploid organisms. General recombination can occur throughout the genome of diploid organisms, using one or a small number of common enzymatic pathways. This chapter will be concerned almost entirely with general recombination. Illegitimate or nonhomologous recombination occurs in regions where no large- scale sequence similarity is apparent, e.g. translocations between different chromosomes or deletions that remove several genes along a chromosome. However, when the DNA sequence at the breakpoints for these events is analyzed, short regions of sequence similarity are found in some cases. For instance, recombination between two similar genes that are several million bp apart can lead to deletion of the intervening genes in somatic cells. -

Protein-Assisted Room-Temperature Assembly of Rigid, Immobile Holliday Junctions and Hierarchical DNA Nanostructures
molecules Article Protein-Assisted Room-Temperature Assembly of Rigid, Immobile Holliday Junctions and Hierarchical DNA Nanostructures 1,2, 3,4, 1, Saminathan Ramakrishnan y, Sivaraman Subramaniam y, Charlotte Kielar z, Guido Grundmeier 1 , A. Francis Stewart 3,4 and Adrian Keller 1,* 1 Technical and Macromolecular Chemistry, Paderborn University, Warburger Str. 100, 33098 Paderborn, Germany; [email protected] (S.R.); [email protected] (C.K.); [email protected] (G.G.) 2 Structural Biophysics Laboratory, Center for Cancer Research, National Cancer Institute, Frederick, MD 21702, USA 3 Biotechnology Center, Department of Genomics, Technische Universität Dresden, Tatzberg 47-51, 01307 Dresden, Germany; [email protected] (S.S.); [email protected] (A.F.S.) 4 Cluster of Excellence Physics of Life, Technische Universität Dresden, 01062 Dresden, Germany * Correspondence: [email protected] These authors contributed equally to this work. y Present address: Institute of Resource Ecology, Helmholtz-Zentrum Dresden-Rossendorf, z Bautzner Landstraße 400, 01328 Dresden, Germany. Academic Editor: Ramon Eritja Received: 29 September 2020; Accepted: 30 October 2020; Published: 3 November 2020 Abstract: Immobile Holliday junctions represent not only the most fundamental building block of structural DNA nanotechnology but are also of tremendous importance for the in vitro investigation of genetic recombination and epigenetics. Here, we present a detailed study on the room-temperature assembly of immobile Holliday junctions with the help of the single-strand annealing protein Redβ. Individual DNA single strands are initially coated with protein monomers and subsequently hybridized to form a rigid blunt-ended four-arm junction. We investigate the efficiency of this approach for different DNA/protein ratios, as well as for different DNA sequence lengths. -

The Crystal Structures of DNA Holliday Junctions P Shing Ho* and Brandt F Eichman†
302 The crystal structures of DNA Holliday junctions P Shing Ho* and Brandt F Eichman† Nearly 40 years ago, Holliday proposed a four-stranded Molecular models of the four-way junction complex or junction as the central intermediate in the general It is now generally accepted that there are two structural mechanism of genetic recombination. During the past two forms of the Holliday junction (reviewed in [9•]). Low salt years, six single-crystal structures of such DNA junctions have conditions favor an open-X form, in which the four arms been determined by three different research groups. These are extended in a square planar geometry (Figure 1b), structures all essentially adopt the antiparallel stacked-X thereby minimizing the repulsion between the negatively conformation, but can be classified into three distinct charged phosphates concentrated at the junction. This is categories: RNA–DNA junctions; ACC trinucleotide junctions; also the form observed in complexes with the enzymes and drug-induced junctions. Together, these structures provide RuvA [10,11,12•] and Cre [13]. insight into how local and distant interactions help to define the detailed and general physical features of Holliday junctions In contrast, the presence of multivalent cations effective- at the atomic level. ly shields these negative charges, allowing the arms to coaxially pair and stack into the more compact stacked-X Addresses junction (Figure 1c). Before the recent crystal studies, *Department of Biochemistry and Biophysics, ALS 2011, Oregon the most detailed models of four-way junctions were State University, Corvallis, Oregon 97331, USA; derived from studies using gel electrophoresis, fluores- e-mail: [email protected] cence resonance energy transfer, NMR spectroscopy and †Department of Biological Chemistry and Molecular Pharmacology, • Harvard Medical School, Boston, Massachusetts 02115, USA atomic force microscopy [9 ]. -

Mammalian BLM Helicase Is Critical for Integrating Multiple Pathways of Meiotic Recombination
JCB: Article Mammalian BLM helicase is critical for integrating multiple pathways of meiotic recombination J. Kim Holloway, Meisha A. Morelli, Peter L. Borst, and Paula E. Cohen Department of Biomedical Sciences, College of Veterinary Medicine, Cornell University, Ithaca, NY 14853 loom’s syndrome (BS) is an autosomal recessive prompting us to explore the meiotic phenotype of mice disorder characterized by growth retardation, bearing a conditional mutant allele of Blm. In this study, B cancer predisposition, and sterility. BS mutated we show that BLM deficiency does not affect entry into (Blm), the gene mutated in BS patients, is one of five mam- prophase I but causes severe defects in meiotic pro- malian RecQ helicases. Although BLM has been shown gression. This is exemplified by improper pairing and to promote genome stability by assisting in the repair of synapsis of homologous chromosomes and altered pro- DNA structures that arise during homologous recombina- cessing of recombination intermediates, resulting in tion in somatic cells, less is known about its role in meiotic increased chiasmata. Our data provide the first analy- recombination primarily because of the embryonic lethal- sis of BLM function in mammalian meiosis and strongly ity associated with Blm deletion. However, the localiza- argue that BLM is involved in proper pairing, synapsis, tion of BLM protein on meiotic chromosomes together with and segregation of homologous chromosomes; how- evidence from yeast and other organisms implicates a ever, it is dispensable for the accumulation of recombi- role for BLM helicase in meiotic recombination events, nation intermediates. Introduction In humans, mutation of the RecQ helicase Bloom’s syndrome indicating that this protein may play an important role in mei (BS) mutated (BLM) results in the carrier suffering from the otic progression.