2159-8290.CD-19-0680.Full-Text.Pdf
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

ENSG Gene Encodes Effector TCR Pathway Costimulation Inhibitory/Exhaustion Synapse/Adhesion Chemokines/Receptors
ENSG Gene Encodes Effector TCR pathway Costimulation Inhibitory/exhaustion Synapse/adhesion Chemokines/receptors ENSG00000111537 IFNG IFNg x ENSG00000109471 IL2 IL-2 x ENSG00000232810 TNF TNFa x ENSG00000271503 CCL5 CCL5 x x ENSG00000139187 KLRG1 Klrg1 x ENSG00000117560 FASLG Fas ligand x ENSG00000121858 TNFSF10 TRAIL x ENSG00000134545 KLRC1 Klrc1 / NKG2A x ENSG00000213809 KLRK1 Klrk1 / NKG2D x ENSG00000188389 PDCD1 PD-1 x x ENSG00000117281 CD160 CD160 x x ENSG00000134460 IL2RA IL-2 receptor x subunit alpha ENSG00000110324 IL10RA IL-10 receptor x subunit alpha ENSG00000115604 IL18R1 IL-18 receptor 1 x ENSG00000115607 IL18RAP IL-18 receptor x accessory protein ENSG00000081985 IL12RB2 IL-12 receptor x beta 2 ENSG00000186810 CXCR3 CXCR3 x x ENSG00000005844 ITGAL CD11a x ENSG00000160255 ITGB2 CD18; Integrin x x beta-2 ENSG00000156886 ITGAD CD11d x ENSG00000140678 ITGAX; CD11c x x Integrin alpha-X ENSG00000115232 ITGA4 CD49d; Integrin x x alpha-4 ENSG00000169896 ITGAM CD11b; Integrin x x alpha-M ENSG00000138378 STAT4 Stat4 x ENSG00000115415 STAT1 Stat1 x ENSG00000170581 STAT2 Stat2 x ENSG00000126561 STAT5a Stat5a x ENSG00000162434 JAK1 Jak1 x ENSG00000100453 GZMB Granzyme B x ENSG00000145649 GZMA Granzyme A x ENSG00000180644 PRF1 Perforin 1 x ENSG00000115523 GNLY Granulysin x ENSG00000100450 GZMH Granzyme H x ENSG00000113088 GZMK Granzyme K x ENSG00000057657 PRDM1 Blimp-1 x ENSG00000073861 TBX21 T-bet x ENSG00000115738 ID2 ID2 x ENSG00000176083 ZNF683 Hobit x ENSG00000137265 IRF4 Interferon x regulatory factor 4 ENSG00000140968 IRF8 Interferon -

The TNF and TNF Receptor Review Superfamilies: Integrating Mammalian Biology
Cell, Vol. 104, 487±501, February 23, 2001, Copyright 2001 by Cell Press The TNF and TNF Receptor Review Superfamilies: Integrating Mammalian Biology Richard M. Locksley,*²³k Nigel Killeen,²k The receptors and ligands in this superfamily have and Michael J. Lenardo§k unique structural attributes that couple them directly to *Department of Medicine signaling pathways for cell proliferation, survival, and ² Department of Microbiology and Immunology differentiation. Thus, they have assumed prominent ³ Howard Hughes Medical Institute roles in the generation of tissues and transient microen- University of California, San Francisco vironments. Most TNF/TNFR SFPs are expressed in the San Francisco, California 94143 immune system, where their rapid and potent signaling § Laboratory of Immunology capabilities are crucial in coordinating the proliferation National Institute of Allergy and Infectious Diseases and protective functions of pathogen-reactive cells. National Institutes of Health Here, we review the organization of the TNF/TNFR SF Bethesda, Maryland 20892 and how these proteins have been adapted for pro- cesses as seemingly disparate as host defense and or- ganogenesis. In interpreting this large and highly active Introduction area of research, we have focused on common themes that unite the actions of these genes in different tissues. Three decades ago, lymphotoxin (LT) and tumor necro- We also discuss the evolutionary success of this super- sis factor (TNF) were identified as products of lympho- familyÐsuccess that we infer from its expansion across cytes and macrophages that caused the lysis of certain the mammalian genome and from its many indispens- types of cells, especially tumor cells (Granger et al., able roles in mammalian biology. -

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

A1068-CD86 Polyclonal Antibody
BioVision 05/16 For research use only CD86 Polyclonal Antibody CATALOG NO: A1068-100 ALTERNATIVE NAMES: T-lymphocyte activation antigen CD86, Activation B7-2 antigen, B70, BU63, CTLA-4 counter-receptor B72, FUN-1, CD86, CD86, CD28LG2 Western blot analysis of CD86 in NCI-H292 cell line lysate AMOUNT: 100 µl IMMUNOGEN: KLH conjugated synthetic peptide between 269-298 amino acids from the C-terminal region of human CD86. MOLECULAR WEIGHT: 37 kDa HOST/ISOTYPE: Rabbit IgG SPECIES REACTIVITY: Human PURIFICATION: This antibody is purified through a protein A column, followed by peptide affinity purification. FORM: Liquid FORMULATION: Supplied in PBS with 0.09% (W/V) sodium azide. STORAGE CONDITIONS: Maintain refrigerated at 2-8°C for up to 6 months. For long term RELATED PRODUCTS storage store at -20°C in small aliquots to prevent freeze-thaw cycles. DESCRIPTION: This gene encodes a type I membrane protein that is a member of the immunoglobulin superfamily. This protein is expressed by Human CellExp™ B7-2 /CD86, human recombinant (Cat. No. 7496-10, -50) antigen-presenting cells, and it is the ligand for two proteins at the CD86 (Human) ELISA Kit (Cat. No. K4175-100) cell surface of T cells, CD28 antigen and cytotoxic T-lymphocyte- Human CellExp™LAG3 /CD223, human recombinant (Cat. No. 7278-10, -50) associated protein 4. Binding of this protein with CD28 antigen is a costimulatory signal for activation of the T-cell. Binding of this CD223 (LAG3) Polyclonal Antibody (Cat. No. A1067-100) protein with cytotoxic T-lymphocyte-associated protein 4 negatively regulates T-cell activation and diminishes the immune response. -

Molecular and Clinical Characterization of LAG3 in Breast Cancer Through 2994 Samples
Molecular and Clinical Characterization of LAG3 in Breast Cancer Through 2994 Samples Qiang Liu Chinese Academy of Medical Sciences & Peking Union Medical College Yihang Qi ( [email protected] ) Chinese Academy of Medical Sciences and Peking Union Medical College https://orcid.org/0000-0001- 7589-0333 Jie Zhai Chinese Academy of Medical Sciences & Peking Union Medical College Xiangyi Kong Chinese Academy of Medical Sciences & Peking Union Medical College Xiangyu Wang Chinese Academy of Medical Sciences & Peking Union Medical College Yi Fang Chinese Academy of Medical Sciences & Peking Union Medical College Jing Wang Chinese Academy of Medical Sciences & Peking Union Medical College Research Keywords: Cancer immunotherapy, CD223, LAG3, Immune response, Inammatory activity Posted Date: June 19th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-36422/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/33 Abstract Background Despite the promising impact of cancer immunotherapy targeting CTLA4 and PD1/PDL1, a large number of cancer patients fail to respond. LAG3 (Lymphocyte Activating 3), also named CD233, is a protein Coding gene served as alternative inhibitory receptors to be targeted in the clinic. The impact of LAG3 on immune cell populations and co-regulation of immune response in breast cancer remained largely unknown. Methods To characterize the role of LAG3 in breast cancer, we investigated transcriptome data and associated clinical information derived from a total of 2994 breast cancer patients. Results We observed that LAG3 was closely correlated with major molecular and clinical characteristics, and was more likely to be enriched in higher malignant subtype, suggesting LAG3 was a potential biomarker of triple-negative breast cancer. -
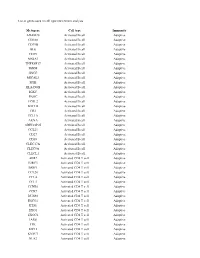
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

LAG-3-Expressing Tumor-Infiltrating T Cells Are Associated with Reduced
cancers Article LAG-3-Expressing Tumor-Infiltrating T Cells Are Associated with Reduced Disease-Free Survival in Pancreatic Cancer Lena Seifert 1,2,3,†, Ioana Plesca 4,†, Luise Müller 4, Ulrich Sommer 5, Max Heiduk 1,2, Janusz von Renesse 1, David Digomann 1, Jessica Glück 1, Anna Klimova 6,7, Jürgen Weitz 1,2,3, Marc Schmitz 2,3,4 and Adrian M. Seifert 1,2,3,* 1 Department of Visceral, Thoracic and Vascular Surgery, University Hospital Carl Gustav Carus, TU Dresden, 01307 Dresden, Germany; [email protected] (L.S.); [email protected] (M.H.); [email protected] (J.v.R.); [email protected] (D.D.); [email protected] (J.G.); [email protected] (J.W.) 2 National Center for Tumor Diseases (NCT), Partner Site Dresden, 69120 Heidelberg, Germany; [email protected] 3 German Cancer Consortium (DKTK), Partner Site Dresden, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany 4 Faculty of Medicine Carl Gustav Carus, Institute of Immunology, TU Dresden, 01307 Dresden, Germany; [email protected] (I.P.); [email protected] (L.M.) 5 Faculty of Medicine Carl Gustav Carus, Institute of Pathology, TU Dresden, 01307 Dresden, Germany; [email protected] 6 Faculty of Medicine Carl Gustav Carus, Institute for Medical Informatics and Biometry, TU Dresden, 01307 Dresden, Germany; [email protected] 7 National Center for Tumor Diseases (NCT), Core Unit for Data Management and Analytics (CDMA), Citation: Seifert, L.; Plesca, I.; Müller, 01307 Dresden, Germany L.; Sommer, U.; Heiduk, M.; von * Correspondence: [email protected] Renesse, J.; Digomann, D.; Glück, J.; † These authors have contributed equally to this work. -

Diabetes in the Absence of LAG-3 Cutting Edge
Cutting Edge: Accelerated Autoimmune Diabetes in the Absence of LAG-3 Maria Bettini, Andrea L. Szymczak-Workman, Karen Forbes, Ashley H. Castellaw, Mark Selby, Xiaoyu Pan, This information is current as Charles G. Drake, Alan J. Korman and Dario A. A. Vignali of September 24, 2021. J Immunol 2011; 187:3493-3498; Prepublished online 26 August 2011; doi: 10.4049/jimmunol.1100714 http://www.jimmunol.org/content/187/7/3493 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2011/08/26/jimmunol.110071 Material 4.DC1 http://www.jimmunol.org/ References This article cites 25 articles, 11 of which you can access for free at: http://www.jimmunol.org/content/187/7/3493.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 24, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2011 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Cutting Edge: Accelerated Autoimmune Diabetes in the Absence of LAG-3 Maria Bettini,* Andrea L. -

Effector CD4 T-Cell Transition to Memory Requires Late Cognate Interactions That Induce Autocrine IL-2
ARTICLE Received 3 Jun 2014 | Accepted 24 Sep 2014 | Published 5 Nov 2014 DOI: 10.1038/ncomms6377 Effector CD4 T-cell transition to memory requires late cognate interactions that induce autocrine IL-2 K. Kai McKinstry1,*, Tara M. Strutt1,*, Bianca Bautista1, Wenliang Zhang1, Yi Kuang1, Andrea M. Cooper2 & Susan L. Swain1 It is unclear how CD4 T-cell memory formation is regulated following pathogen challenge, and when critical mechanisms act to determine effector T-cell fate. Here, we report that following influenza infection most effectors require signals from major histocompatibility complex class II molecules and CD70 during a late window well after initial priming to become memory. During this timeframe, effector cells must produce IL-2 or be exposed to high levels of paracrine or exogenously added IL-2 to survive an otherwise rapid default contraction phase. Late IL-2 promotes survival through acute downregulation of apoptotic pathways in effector T cells and by permanently upregulating their IL-7 receptor expression, enabling IL-7 to sustain them as memory T cells. This new paradigm defines a late checkpoint during the effector phase at which cognate interactions direct CD4 T-cell memory generation. 1 Department of Pathology, University of Massachusetts Medical School, 55 Lake Avenue North, Worcester, Massachusetts 01655, USA. 2 Trudeau Institute, 154 Algonquin Avenue, Saranac Lake, New York 12983, USA. * These authors contributed equally to this work. Correspondence and requests for materials should be addressed to K.K.M. (email: [email protected]). NATURE COMMUNICATIONS | 5:5377 | DOI: 10.1038/ncomms6377 | www.nature.com/naturecommunications 1 & 2014 Macmillan Publishers Limited. -

Single-Cell RNA Sequencing Demonstrates the Molecular and Cellular Reprogramming of Metastatic Lung Adenocarcinoma
ARTICLE https://doi.org/10.1038/s41467-020-16164-1 OPEN Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma Nayoung Kim 1,2,3,13, Hong Kwan Kim4,13, Kyungjong Lee 5,13, Yourae Hong 1,6, Jong Ho Cho4, Jung Won Choi7, Jung-Il Lee7, Yeon-Lim Suh8,BoMiKu9, Hye Hyeon Eum 1,2,3, Soyean Choi 1, Yoon-La Choi6,10,11, Je-Gun Joung1, Woong-Yang Park 1,2,6, Hyun Ae Jung12, Jong-Mu Sun12, Se-Hoon Lee12, ✉ ✉ Jin Seok Ahn12, Keunchil Park12, Myung-Ju Ahn 12 & Hae-Ock Lee 1,2,3,6 1234567890():,; Advanced metastatic cancer poses utmost clinical challenges and may present molecular and cellular features distinct from an early-stage cancer. Herein, we present single-cell tran- scriptome profiling of metastatic lung adenocarcinoma, the most prevalent histological lung cancer type diagnosed at stage IV in over 40% of all cases. From 208,506 cells populating the normal tissues or early to metastatic stage cancer in 44 patients, we identify a cancer cell subtype deviating from the normal differentiation trajectory and dominating the metastatic stage. In all stages, the stromal and immune cell dynamics reveal ontological and functional changes that create a pro-tumoral and immunosuppressive microenvironment. Normal resident myeloid cell populations are gradually replaced with monocyte-derived macrophages and dendritic cells, along with T-cell exhaustion. This extensive single-cell analysis enhances our understanding of molecular and cellular dynamics in metastatic lung cancer and reveals potential diagnostic and therapeutic targets in cancer-microenvironment interactions. 1 Samsung Genome Institute, Samsung Medical Center, Seoul 06351, Korea. -

Human TNFRSF18 ELISA Kit (ARG81453)
Product datasheet [email protected] ARG81453 Package: 96 wells Human TNFRSF18 ELISA Kit Store at: 4°C Component Cat. No. Component Name Package Temp ARG81453-001 Antibody-coated 8 X 12 strips 4°C. Unused strips microplate should be sealed tightly in the air-tight pouch. ARG81453-002 Standard 2 X 10 ng/vial 4°C ARG81453-003 Standard/Sample 30 ml (Ready to use) 4°C diluent ARG81453-004 Antibody conjugate 1 vial (100 µl) 4°C concentrate (100X) ARG81453-005 Antibody diluent 12 ml (Ready to use) 4°C buffer ARG81453-006 HRP-Streptavidin 1 vial (100 µl) 4°C concentrate (100X) ARG81453-007 HRP-Streptavidin 12 ml (Ready to use) 4°C diluent buffer ARG81453-008 25X Wash buffer 20 ml 4°C ARG81453-009 TMB substrate 10 ml (Ready to use) 4°C (Protect from light) ARG81453-010 STOP solution 10 ml (Ready to use) 4°C ARG81453-011 Plate sealer 4 strips Room temperature Summary Product Description ARG81453 Human TNFRSF18 ELISA Kit is an Enzyme Immunoassay kit for the quantification of Human TNFRSF18 in serum, plasma (heparin, EDTA) and cell culture supernatants. Tested Reactivity Hu Tested Application ELISA Specificity There is no detectable cross-reactivity with other relevant proteins. Target Name TNFRSF18 Conjugation HRP Conjugation Note Substrate: TMB and read at 450 nm. Sensitivity 31.25 pg/ml Sample Type Serum, plasma (heparin, EDTA) and cell culture supernatants. Standard Range 62.5 - 4000 pg/ml Sample Volume 100 µl www.arigobio.com 1/3 Precision Intra-Assay CV: 6.3% Inter-Assay CV: 7.0% Alternate Names Tumor necrosis factor receptor superfamily member 18; AITR; CD357; CD antigen CD357; Activation- inducible TNFR family receptor; GITR-D; GITR; Glucocorticoid-induced TNFR-related protein Application Instructions Assay Time ~ 5 hours Properties Form 96 well Storage instruction Store the kit at 2-8°C. -

An Ontogenetic Switch Drives the Positive and Negative Selection of B Cells
An ontogenetic switch drives the positive and negative selection of B cells Xijin Xua, Mukta Deobagkar-Lelea, Katherine R. Bulla, Tanya L. Crockforda, Adam J. Meadb, Adam P. Cribbsc, David Simsc, Consuelo Anzilottia, and Richard J. Cornalla,1 aMedical Research Council Human Immunology Unit, Weatherall Institute of Molecular Medicine, University of Oxford, OX3 9DS Oxford, United Kingdom; bMedical Research Council Molecular Haematology Unit, Weatherall Institute of Molecular Medicine, University of Oxford, OX3 9DS Oxford, United Kingdom; and cMedical Research Council, Weatherall Institute of Molecular Medicine, Centre for Computational Biology, Weatherall Institute of Molecular Medicine, University of Oxford, OX3 9DS Oxford, United Kingdom Edited by Michael Reth, University of Freiburg, Freiburg, Germany, and approved January 6, 2020 (received for review September 3, 2019) + Developing B cells can be positively or negatively selected by self- BM HSCs increased CD5 B-1a B cell development (15), while antigens, but the mechanisms that determine these outcomes are expression of let-7b in FL pro-B cells blocked the development of incompletely understood. Here, we show that a B cell intrinsic B-1 B cells (17). These findings support the notion of hard-wired switch between positive and negative selection during ontogeny differences during ontogeny, but possibly downstream of the HSC is determined by a change from Lin28b to let-7 gene expression. commitment stage. Ectopic expression of a Lin28b transgene in murine B cells restored Several lines of evidence also suggest that B-1 B cells can un- the positive selection of autoreactive B-1 B cells by self-antigen in dergo positive selection, which is linked to their B cell receptor adult bone marrow.