The Oldest Platypus and Its Bearing on Divergence Timing of the Platypus and Echidna Clades
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Evolution of the Patellar Sesamoid Bone in Mammals
A peer-reviewed version of this preprint was published in PeerJ on 21 March 2017. View the peer-reviewed version (peerj.com/articles/3103), which is the preferred citable publication unless you specifically need to cite this preprint. Samuels ME, Regnault S, Hutchinson JR. 2017. Evolution of the patellar sesamoid bone in mammals. PeerJ 5:e3103 https://doi.org/10.7717/peerj.3103 Evolution of the patellar sesamoid bone in mammals Mark E Samuels 1, 2 , Sophie Regnault 3 , John R Hutchinson Corresp. 3 1 Department of Medicine, University of Montreal, Montreal, Quebec, Canada 2 Centre de Recherche du CHU Ste-Justine, Montreal, Quebec, Canada 3 Structure & Motion Laboratory, Department of Comparative Biomedical Sciences, The Royal Veterinary College, Hatfield, Hertfordshire, United Kingdom Corresponding Author: John R Hutchinson Email address: [email protected] The patella is a sesamoid bone located in the major extensor tendon of the knee joint, in the hindlimb of many tetrapods. Although numerous aspects of knee morphology are ancient and conserved among most tetrapods, the evolutionary occurrence of an ossified patella is highly variable. Among extant (crown clade) groups it is found in most birds, most lizards, the monotreme mammals and almost all placental mammals, but it is absent in most marsupial mammals as well as many reptiles. Here we integrate data from the literature and first-hand studies of fossil and recent skeletal remains to reconstruct the evolution of the mammalian patella. We infer that bony patellae most likely evolved between four to six times in crown group Mammalia: in monotremes, in the extinct multituberculates, in one or more stem-mammal genera outside of therian or eutherian mammals, and up to three times in therian mammals. -

Miocene Mammal Reveals a Mesozoic Ghost Lineage on Insular New Zealand, Southwest Pacific
Miocene mammal reveals a Mesozoic ghost lineage on insular New Zealand, southwest Pacific Trevor H. Worthy*†, Alan J. D. Tennyson‡, Michael Archer§, Anne M. Musser¶, Suzanne J. Hand§, Craig Jonesʈ, Barry J. Douglas**, James A. McNamara††, and Robin M. D. Beck§ *School of Earth and Environmental Sciences, Darling Building DP 418, Adelaide University, North Terrace, Adelaide 5005, South Australia, Australia; ‡Museum of New Zealand Te Papa Tongarewa, P.O. Box 467, Wellington 6015, New Zealand; §School of Biological, Earth and Environmental Sciences, University of New South Wales, New South Wales 2052, Australia; ¶Australian Museum, 6-8 College Street, Sydney, New South Wales 2010, Australia; ʈInstitute of Geological and Nuclear Sciences, P.O. Box 30368, Lower Hutt 5040, New Zealand; **Douglas Geological Consultants, 14 Jubilee Street, Dunedin 9011, New Zealand; and ††South Australian Museum, Adelaide, South Australia 5000, Australia Edited by James P. Kennett, University of California, Santa Barbara, CA, and approved October 11, 2006 (sent for review July 8, 2006) New Zealand (NZ) has long been upheld as the archetypical Ma) dinosaur material (13) and isolated moa bones from marine example of a land where the biota evolved without nonvolant sediments up to 2.5 Ma (1, 14), the terrestrial record older than terrestrial mammals. Their absence before human arrival is mys- 1 Ma is extremely limited. Until now, there has been no direct terious, because NZ was still attached to East Antarctica in the Early evidence for the pre-Pleistocene presence in NZ of any of its Cretaceous when a variety of terrestrial mammals occupied the endemic vertebrate lineages, particularly any group of terrestrial adjacent Australian portion of Gondwana. -

Versión Disponible En PDF
nicolás r chimento, Federico L agnolin y Fernando e novas Museo Argentino de Ciencias Naturales Bernardino Rivadavia Necrolestes un mamífero patagónico que sobrevivió a la extinción de los dinosaurios Un descubrimiento patagónico desembocadura del río Santa Cruz, descubrieron esque- letos fósiles prácticamente completos del Necrolestes. El es- En 1891, Florentino Ameghino (1854-1911) dio a tudio de esos esqueletos llevó a pensar que se trataba de conocer unos restos fósiles encontrados por su hermano un mamífero muy arcaico en la historia de la evolución, Carlos (1865-1936) en las barrancas de Monte Obser- más que un ancestro de los topos, como había supuesto vación, en la provincia de Santa Cruz, en yacimientos Ameghino. Por determinados rasgos se pensó que podía de unos 17 millones de años de antigüedad. Determinó haber sido un marsupial, es decir, un pariente lejano de las que pertenecían a un pequeño –escasos 10cm de largo, comadrejas, los canguros y los coalas actuales. del hocico a la cola– y desconocido mamífero extin- Ciertos investigadores aceptaron esta última hipótesis guido. Estudió los diminutos huesos y consideró que de parentesco, pero otros se mostraron escépticos acerca el animal habría sido un pariente lejano de los topos de ella y se inclinaron por considerar inciertas las rela- africanos vivientes. Le dio el nombre científicoN ecrolestes ciones genealógicas del diminuto mamífero. Así, su po- patagonensis, es decir, ladrón de tumbas de la Patagonia, en sición en el árbol evolutivo de los mamíferos fue objeto alusión a sus hábitos excavadores. El hallazgo, publica- de debate durante gran parte del siglo XX. Para algunos, do por Ameghino en el número de la Revista Argentina de era pariente lejano de las mulitas; otros seguían pensan- Historia Natural citado entre las lecturas sugeridas, atrajo do que podía estar relacionado con los topos, y para un la atención del ámbito científico, ya que hasta ese mo- tercer grupo, sus vínculos eran con los marsupiales aus- mento en Sudamérica no se habían encontrado restos tralianos. -

Using Dental Enamel Wrinkling to Define Sauropod Tooth Morphotypes from the Cañadón Asfalto Formation, Patagonia, Argentina
RESEARCH ARTICLE Using Dental Enamel Wrinkling to Define Sauropod Tooth Morphotypes from the Cañadón Asfalto Formation, Patagonia, Argentina Femke M. Holwerda1,2,3*, Diego Pol4,5, Oliver W. M. Rauhut1,3 1 Staatliche Naturwissenschaftliche Sammlungen Bayerns (SNSB), Bayerische Staatssammlung für Paläontologie und Geologie, München, Germany, 2 GeoBioTec, Departamento de Ciências da Terra, Faculdade de Ciências e Tecnologia (FCT), Universidade Nova de Lisboa, Caparica, Portugal, 3 Department of Earth and Environmental Sciences and GeoBioCenter, Ludwig Maximilians Universität, München, Germany, 4 Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET), Buenos Aires, Argentina, 5 Museo Paleontológico Egidio Feruglio, Trelew, Argentina * [email protected] Abstract OPEN ACCESS The early Middle Jurassic is regarded as the period when sauropods diversified and be- Citation: Holwerda FM, Pol D, Rauhut OWM (2015) came major components of the terrestrial ecosystems. Not many sites yield sauropod mate- Using Dental Enamel Wrinkling to Define Sauropod Tooth Morphotypes from the Cañadón Asfalto rial of this time; however, both cranial and postcranial material of eusauropods have been Formation, Patagonia, Argentina. PLoS ONE 10(2): found in the Cañadón Asfalto Formation (latest Early Jurassic–early Middle Jurassic) in e0118100. doi:10.1371/journal.pone.0118100 Central Patagonia (Argentina), which may help to shed light on the early evolution of Academic Editor: Peter Dodson, University of eusauropods. These eusauropod remains include teeth associated with cranial and man- Pennsylvania, UNITED STATES dibular material as well as isolated teeth found at different localities. In this study, an assem- Received: September 1, 2014 blage of sauropod teeth from the Cañadón Asfalto Formation found in four different Accepted: January 7, 2015 localities in the area of Cerro Condor (Chubut, Argentina) is used as a mean of assessing sauropod species diversity at these sites. -

Constraints on the Timescale of Animal Evolutionary History
Palaeontologia Electronica palaeo-electronica.org Constraints on the timescale of animal evolutionary history Michael J. Benton, Philip C.J. Donoghue, Robert J. Asher, Matt Friedman, Thomas J. Near, and Jakob Vinther ABSTRACT Dating the tree of life is a core endeavor in evolutionary biology. Rates of evolution are fundamental to nearly every evolutionary model and process. Rates need dates. There is much debate on the most appropriate and reasonable ways in which to date the tree of life, and recent work has highlighted some confusions and complexities that can be avoided. Whether phylogenetic trees are dated after they have been estab- lished, or as part of the process of tree finding, practitioners need to know which cali- brations to use. We emphasize the importance of identifying crown (not stem) fossils, levels of confidence in their attribution to the crown, current chronostratigraphic preci- sion, the primacy of the host geological formation and asymmetric confidence intervals. Here we present calibrations for 88 key nodes across the phylogeny of animals, rang- ing from the root of Metazoa to the last common ancestor of Homo sapiens. Close attention to detail is constantly required: for example, the classic bird-mammal date (base of crown Amniota) has often been given as 310-315 Ma; the 2014 international time scale indicates a minimum age of 318 Ma. Michael J. Benton. School of Earth Sciences, University of Bristol, Bristol, BS8 1RJ, U.K. [email protected] Philip C.J. Donoghue. School of Earth Sciences, University of Bristol, Bristol, BS8 1RJ, U.K. [email protected] Robert J. -

Like Globin Genes in Monotremes and Therian Mammals
Genomic evidence for independent origins of -like globin genes in monotremes and therian mammals Juan C. Opazo*, Federico G. Hoffmann, and Jay F. Storz† School of Biological Sciences, University of Nebraska, Lincoln, NE 68588 Edited by Morris Goodman, Wayne State University School of Medicine, Detroit, MI, and approved December 13, 2007 (received for review November 5, 2007) Phylogenetic reconstructions of the -globin gene family in ver- embryonic -globin gene is exclusively expressed in primitive tebrates have revealed that developmentally regulated systems of erythroid cells derived from the yolk sac. However, the ‘‘- hemoglobin synthesis have been reinvented multiple times in globin’’ gene in birds is not orthologous to the -globin gene in independent lineages. For example, the functional differentiation mammals (2, 12), because they are independently derived from of embryonic and adult -like globin genes occurred independently lineage-specific duplications of a proto -globin gene. in birds and mammals. In both taxa, the embryonic -globin gene In placental mammals (subclass Eutheria), the -globin gene is exclusively expressed in primitive erythroid cells derived from cluster includes a linked set of three early expressed (prenatal) the yolk sac. However, the ‘‘-globin’’ gene in birds is not ortholo- genes, -␥-,atthe5Ј end of the cluster, and a pair of late gous to the -globin gene in mammals, because they are indepen- expressed (adult) genes, ␦ and ,atthe3Ј end. There is extensive dently derived from lineage-specific duplications of a proto variation in the copy number of these different paralogs among -globin gene. Here, we report evidence that the early and late species, and in a number of placental mammal lineages, the - expressed -like globin genes in monotremes and therian mam- and ␦-globin genes have been inactivated or deleted (13–15). -

Fossil Focus: Dinosaurs Down Under Author(S): Stephen F
www.palaeontologyonline.com |Page 1 Title: Fossil Focus: Dinosaurs Down Under Author(s): Stephen F. Propat *1 Volume: 5 Article: 1 Page(s): 1-11 Published Date: 01/01/2015 PermaLink: http://www.palaeontologyonline.com/articles/2015/fossil-focus-dinosaurs/ IMPORTANT Your use of the Palaeontology [online] archive indicates your acceptance of Palaeontology [online]'s Terms and Conditions of Use, available at http://www.palaeontologyonline.com/site-information/terms-and-conditions/. COPYRIGHT Palaeontology [online] (www.palaeontologyonline.com) publishes all work, unless otherwise stated, under the Creative Commons Attribution 3.0 Unported (CC BY 3.0) license. This license lets others distribute, remix, tweak, and build upon the published work, even commercially, as long as they credit Palaeontology[online] for the original creation. This is the most accommodating of licenses offered by Creative Commons and is recommended for maximum dissemination of published material. Further details are available at http://www.palaeontologyonline.com/site-information/copyright/. CITATION OF ARTICLE Please cite the following published work as: Propat, Stephen F.. 2015. Fossil Focus: Dinosaurs Down Under, Palaeontology Online, Volume 5, Article 1, 1- 11. Published by: Palaeontology [online] www.palaeontologyonline.com |Page 2 Fossil Focus: Dinosaurs Down Under by Stephen F. Poropat*1,2 Introduction: Ask the average person in the street to name an Australian dinosaur, and you will be lucky if you get a correct answer. If they say crocodile, they are in the right postcode but have the wrong address. If they say emu, then they are correct, strictly speaking, but they are either lucky or being smart. If they say kangaroo, back away slowly and avoid eye contact. -

Lower Triassic Postcanine Teeth with Allotherian-Like Crowns
Research Letters South African Journal of Science 103, May/June 2007 245 Lower Triassic postcanine teeth with allotherian-like crowns F. Abdala*‡, H. Mocke*§ and P.J. Hancox* The Allotheria are fossil mammals with upper and lower post- canines usually showing two longitudinal rows of cusps separated by a central valley. The group comprises the poorly known haramiyids, mostly represented by isolated teeth, and the notably diverse and long-lived multituberculates; its monophyly is uncer- tain. The oldest records of this particular group are the Late Triassic (Norian–Rhaetian) haramiyids. We present here postcanines with haramiyid-like crowns that were recovered from the Lower Triassic of South Africa. A distinguishing feature of the new teeth is that they are single-rooted. This is the oldest record of mammal-like teeth with crowns having parallel rows of cusps, representing a temporal extension of some 43 million years from similar crown patterns of haramiyids and tritylodontids. This finding reinforces evidence of the remarkable faunal turnover of therapsids in the Early/Middle Triassic, at which time an explosive origin followed by a rapid early diversification of herbivorous/omnivorous forms with occluding expanded postcanines took place. Introduction The Beaufort Group of the South African Karoo shows an abundance and diversity of non-mammalian synapsids, which have allowed for biostratigraphic subdivisions ranging from Middle Permian to Middle Triassic.1 The youngest of these Fig. 1.Allotherian-like teeth.A, Occlusal and lateral views of BP/1/6515 (Pattern 1); B, occlusal and lateral views of BP/1/6516 (Pattern 2). biozones, the Cynognathus Assemblage Zone (AZ), comprises the full extent of the Burgersdorp Formation of the Tarkastad Sub- found to be most parsimonious from an unconstrained search, group (J. -

Early Cretaceous Amphilestid ('Triconodont') Mammals from Mongolia
Early Cretaceous amphilestid ('triconodont') mammals from Mongolia ZOFIAKIELAN-JAWOROWSKA and DEMBERLYIN DASHZEVEG Kielan-Jaworowską Z. &Daslueveg, D. 1998. Early Cretaceous amphilestid (.tricono- dont') mammals from Mongotia. - Acta Pal.aeontol.ogicaPolonica,43,3, 413438. Asmall collection of ?Aptianor ?Albian amphilestid('triconodont') mammals consisting of incomplete dentaries and maxillae with teeth, from the Khoboor localiĘ Guchin Us counĘ in Mongolia, is described. Grchinodon Troftmov' 1978 is regarded a junior subjective synonym of GobiconodonTroftmov, 1978. Heavier wear of the molariforms M3 andM4than of themore anteriorone-M2 in Gobiconodonborissiaki gives indirect evidence formolariformreplacement in this taxon. The interlocking mechanismbetween lower molariforms n Gobiconodon is of the pattern seen in Kuchneotherium and Ttnodon. The ińterlocking mechanism and the type of occlusion ally Amphilestidae with Kuehneotheriidae, from which they differ in having lower molariforms with main cusps aligned and the dentary-squamosal jaw joint (double jaw joint in Kuehneotheńdae). The main cusps in upper molariforms M3-M5 of Gobiconodon, however, show incipient tńangular arrangement. The paper gives some support to Mills' idea on the therian affinities of the Amphilestidae, although it cannot be excluded that the characters that unite the two groups developed in parallel. Because of scanty material and arnbiguĘ we assign the Amphilestidae to order incertae sedis. Key words : Mammali4 .triconodonts', Amphilestidae, Kuehneotheriidae, Early Cretaceous, Mongolia. Zofia Kiel,an-Jaworowska [zkielnn@twarda,pan.pl], InsĘtut Paleobiologii PAN, ul. Twarda 5 I /5 5, PL-00-8 I 8 Warszawa, Poland. DemberĘin Dash7eveg, Geological Institute, Mongolian Academy of Sciences, Ulan Bator, Mongolia. Introduction Beliajeva et al. (1974) reportedthe discovery of Early Cretaceous mammals at the Khoboor locality (referred to also sometimes as Khovboor), in the Guchin Us Soinon (County), Gobi Desert, Mongolia. -
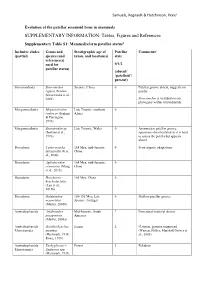
SUPPLEMENTARY INFORMATION: Tables, Figures and References
Samuels, Regnault & Hutchinson, PeerJ Evolution of the patellar sesamoid bone in mammals SUPPLEMENTARY INFORMATION: Tables, Figures and References Supplementary Table S1: Mammaliaform patellar status$ Inclusive clades Genus and Stratigraphic age of Patellar Comments# (partial) species (and taxon, and location(s) state reference(s) used for 0/1/2 patellar status) (absent/ ‘patelloid’/ present) Sinoconodonta Sinoconodon Jurassic, China 0 Patellar groove absent, suggests no rigneyi (Kielan- patella Jaworowska et al., 2004) Sinoconodon is included on our phylogeny within tritylodontids. Morganucodonta Megazostrodon Late Triassic, southern 0 rudnerae (Jenkins Africa & Parrington, 1976) Morganucodonta Eozostrodon sp. Late Triassic, Wales 0 Asymmetric patellar groove, (Jenkins et al., specimens disarticulated so it is hard 1976) to assess the patella but appears absent Docodonta Castorocauda 164 Mya, mid-Jurassic, 0 Semi-aquatic adaptations lutrasimilis (Ji et China al., 2006) Docodonta Agilodocodon 164 Mya, mid-Jurassic, 0 scansorius (Meng China et al., 2015) Docodonta Docofossor 160 Mya, China 0 brachydactylus (Luo et al., 2015b) Docodonta Haldanodon 150-155 Mya, Late 0 Shallow patellar groove exspectatus Jurassic, Portugal (Martin, 2005b) Australosphenida Asfaltomylos Mid-Jurassic, South ? Postcranial material absent patagonicus America (Martin, 2005a) Australosphenida Ornithorhynchus Extant 2 Platypus, genome sequenced Monotremata anatinus (Warren, Hillier, Marshall Graves et (Herzmark, 1938; al., 2008) Rowe, 1988) Australosphenida Tachyglossus -
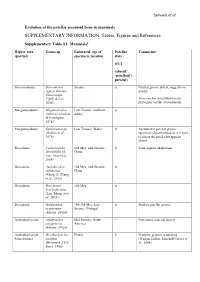
SUPPLEMENTARY INFORMATION: Tables, Figures and References
Samuels et al. Evolution of the patellar sesamoid bone in mammals SUPPLEMENTARY INFORMATION: Tables, Figures and References Supplementary Table S1: Mammals$ Higher taxa Genus sp. Estimated. age of Patellar Comments# (partial) specimen, location state 0/1/2 (absent/ ‘patelloid’/ present) Sinoconodonta Sinoconodon Jurassic 0 Patellar groove absent, suggests no rigneyi (Kielan- patella Jaworowska, Cifelli & Luo, Sinoconodon is included on our 2004) phylogeny within tritylodontids. Morganucodonta Megazostrodon Late Triassic, southern 0 rudnerae (Jenkins Africa & Parrington, 1976) Morganucodonta Eozostrodon sp. Late Triassic, Wales 0 Asymmetric patellar groove, (Jenkins et al., specimens disarticulated so it is hard 1976) to assess the patella but appears absent Docodonta Castorocauda 164 Mya, mid-Jurassic, 0 Semi-aquatic adaptations lutrasimilis (Ji, China Luo, Yuan et al., 2006) Docodonta Agilodocodon 164 Mya, mid-Jurassic, 0 scansorius China (Meng, Ji, Zhang et al., 2015) Docodonta Docofossor 160 Mya 0 brachydactylus (Luo, Meng, Ji et al., 2015) Docodonta Haldanodon 150-155 Mya, Late 0 Shallow patellar groove exspectatus Jurassic, Portugal (Martin, 2005b) Australosphenida Asfaltomylos Mid-Jurassic, South ? Postcranial material absent patagonicus America (Martin, 2005a) Australosphenida Ornithorhynchus Extant 2 Platypus, genome sequenced Monotremata anatinus (Warren, Hillier, Marshall Graves et (Herzmark, 1938; al., 2008) Rowe, 1988) Samuels et al. Australosphenida Tachyglossus + Extant 2 Echidnas Monotremata Zaglossus spp. (Herzmark, 1938; Rowe, 1988) Mammaliaformes Fruitafossor 150 Mya, Late Jurassic, 0 Phylogenetic status uncertain indet. windscheffeli (Luo Colorado & Wible, 2005) Mammaliaformes Volaticotherium Late Jurassic/Early ? Hindlimb material incomplete indet. antiquus (Meng, Cretaceous Hu, Wang et al., 2006) Eutriconodonta Jeholodens 120-125 Mya, Early 0 Poorly developed patellar groove jenkinsi (Ji, Luo Cretaceous, China & Ji, 1999) Eutriconodonta Gobiconodon spp. -

Two New Species of Gobiconodon (Mammalia, Eutriconodonta, Gobiconodontidae) from the Lower Cretaceous Shahai and Fuxin Formations, Northeastern China
Historical Biology An International Journal of Paleobiology ISSN: 0891-2963 (Print) 1029-2381 (Online) Journal homepage: http://www.tandfonline.com/loi/ghbi20 Two new species of Gobiconodon (Mammalia, Eutriconodonta, Gobiconodontidae) from the Lower Cretaceous Shahai and Fuxin formations, northeastern China Nao Kusuhashi, Yuan-Qing Wang, Chuan-Kui Li & Xun Jin To cite this article: Nao Kusuhashi, Yuan-Qing Wang, Chuan-Kui Li & Xun Jin (2016) Two new species of Gobiconodon (Mammalia, Eutriconodonta, Gobiconodontidae) from the Lower Cretaceous Shahai and Fuxin formations, northeastern China, Historical Biology, 28:1-2, 14-26 To link to this article: http://dx.doi.org/10.1080/08912963.2014.977881 Published online: 01 Oct 2015. Submit your article to this journal View related articles View Crossmark data Full Terms & Conditions of access and use can be found at http://www.tandfonline.com/action/journalInformation?journalCode=ghbi20 Download by: [University of Sussex Library] Date: 01 October 2015, At: 18:24 Historical Biology, 2016 Vol. 28, Nos. 1–2, 14–26, http://dx.doi.org/10.1080/08912963.2014.977881 Two new species of Gobiconodon (Mammalia, Eutriconodonta, Gobiconodontidae) from the Lower Cretaceous Shahai and Fuxin formations, northeastern China Nao Kusuhashia*, Yuan-Qing Wangb, Chuan-Kui Lib and Xun Jinb aDepartment of Earth’s Evolution and Environment, Graduate School of Science and Engineering, Ehime University, Ehime 790-8577, Japan; bKey Laboratory of Vertebrate Evolution and Human Origins of Chinese Academy of Sciences, Institute of Vertebrate Paleontology and Paleoanthropology, Chinese Academy of Sciences, Beijing 100044, P.R. China (Received 29 July 2014; accepted 14 October 2014) Two new gobiconodontid mammals, Gobiconodon tomidai sp.