The RNA Polymerase II CTD Coordinates Transcription and RNA Processing
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Regulation of SETD2 Stability Is Important for the Fidelity Of
Bhattacharya and Workman Epigenetics & Chromatin (2020) 13:40 Epigenetics & Chromatin https://doi.org/10.1186/s13072-020-00362-8 RESEARCH Open Access Regulation of SETD2 stability is important for the fdelity of H3K36me3 deposition Saikat Bhattacharya and Jerry L. Workman* Abstract Background: The histone H3K36me3 mark regulates transcription elongation, pre-mRNA splicing, DNA methylation, and DNA damage repair. However, knowledge of the regulation of the enzyme SETD2, which deposits this function- ally important mark, is very limited. Results: Here, we show that the poorly characterized N-terminal region of SETD2 plays a determining role in regulat- ing the stability of SETD2. This stretch of 1–1403 amino acids contributes to the robust degradation of SETD2 by the proteasome. Besides, the SETD2 protein is aggregate prone and forms insoluble bodies in nuclei especially upon proteasome inhibition. Removal of the N-terminal segment results in the stabilization of SETD2 and leads to a marked increase in global H3K36me3 which, uncharacteristically, happens in a Pol II-independent manner. Conclusion: The functionally uncharacterized N-terminal segment of SETD2 regulates its half-life to maintain the requisite cellular amount of the protein. The absence of SETD2 proteolysis results in a Pol II-independent H3K36me3 deposition and protein aggregation. Keywords: Chromatin, Proteasome, Histone, Methylation, Aggregation Background In yeast, the SET domain-containing protein Set2 Te N-terminal tails of histones protrude from the nucle- (ySet2) is the sole H3K36 methyltransferase [10]. ySet2 osome and are hotspots for the occurrence of a variety interacts with the large subunit of the RNA polymerase II, of post-translational modifcations (PTMs) that play key Rpb1, through its SRI (Set2–Rpb1 Interaction) domain, roles in regulating epigenetic processes. -

Cyclin K Interacts with Β-Catenin to Induce Cyclin D1 Expression And
Theranostics 2020, Vol. 10, Issue 24 11144 Ivyspring International Publisher Theranostics 2020; 10(24): 11144-11158. doi: 10.7150/thno.42578 Research Paper Cyclin K interacts with β-catenin to induce Cyclin D1 expression and facilitates tumorigenesis and radioresistance in lung cancer Guojun Yao*, Jing Tang*, Xijie Yang, Ye Zhao, Rui Zhou, Rui Meng, Sheng Zhang, Xiaorong Dong, Tao Zhang, Kunyu Yang, Gang Wu and Shuangbing Xu Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. *These authors contributed equally to this work. Corresponding author: Shuangbing Xu or Gang Wu, Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. E-mail: [email protected] or [email protected]. © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2019.11.29; Accepted: 2020.08.24; Published: 2020.09.11 Abstract Rationale: Radioresistance remains the major cause of local relapse and distant metastasis in lung cancer. However, the underlying molecular mechanisms remain poorly defined. This study aimed to investigate the role and regulatory mechanism of Cyclin K in lung cancer radioresistance. Methods: Expression levels of Cyclin K were measured by immunohistochemistry in human lung cancer tissues and adjacent normal lung tissues. Cell growth and proliferation, neutral comet and foci formation assays, G2/M checkpoint and a xenograft mouse model were used for functional analyses. Gene expression was examined by RNA sequencing and quantitative real-time PCR. -
Are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway Is Associated with a More Favorable Phenotype
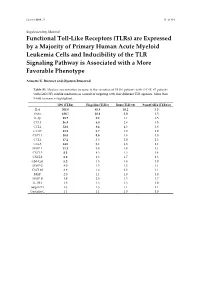
Cancers 2019, 11 S1 of S19 Supplementary Material Functional Toll-Like Receptors (TLRs) are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway is Associated with a More Favorable Phenotype Annette K. Brenner and Øystein Bruserud Table S1. Median concentration increase in the secretion of 19 (16 patients with G-CSF, 67 patients with GM-CSF) soluble mediators as a result of targeting with four different TLR agonists. More than 5-fold increase is highlighted. LPS (TLR4) Flagellin (TLR5) R848 (TLR7/8) Pam3CSK4 (TLR1/2) IL-6 301.0 45.9 10.2 5.3 TNFα 188.7 10.6 5.0 1.5 IL-1β 89.7 9.2 1.1 1.5 CCL3 56.4 6.0 2.4 1.5 CCL2 52.6 9.4 4.3 2.8 G-CSF 51.9 2.7 1.0 1.0 CXCL1 38.0 5.8 1.4 1.5 CCL4 17.2 3.3 2.0 1.3 CCL5 14.5 2.1 1.3 1.1 MMP-1 11.1 3.4 1.4 1.1 CXCL5 8.3 4.3 1.3 1.4 CXCL8 6.0 4.3 1.7 1.3 GM-CSF 5.2 1.5 1.4 1.0 MMP-2 4.0 1.5 1.2 1.1 CXCL10 3.9 1.6 2.2 1.1 HGF 2.0 1.1 1.0 1.0 MMP-9 1.9 2.5 1.3 1.7 IL-1RA 1.8 1.3 1.3 1.0 Serpin E1 1.3 1.3 1.1 1.1 Cystatin C 1.1 1.1 1.0 1.0 Cancers 2019, 11 S2 of S19 Table S2. -

Multiple Cellular Proteins Interact with LEDGF/P75 Through a Conserved Unstructured Consensus Motif
ARTICLE Received 19 Jan 2015 | Accepted 1 Jul 2015 | Published 6 Aug 2015 DOI: 10.1038/ncomms8968 Multiple cellular proteins interact with LEDGF/p75 through a conserved unstructured consensus motif Petr Tesina1,2,3,*, Katerˇina Cˇerma´kova´4,*, Magdalena Horˇejsˇ´ı3, Katerˇina Procha´zkova´1, Milan Fa´bry3, Subhalakshmi Sharma4, Frauke Christ4, Jonas Demeulemeester4, Zeger Debyser4, Jan De Rijck4,**, Va´clav Veverka1,** & Pavlı´na Rˇeza´cˇova´1,3,** Lens epithelium-derived growth factor (LEDGF/p75) is an epigenetic reader and attractive therapeutic target involved in HIV integration and the development of mixed lineage leukaemia (MLL1) fusion-driven leukaemia. Besides HIV integrase and the MLL1-menin complex, LEDGF/p75 interacts with various cellular proteins via its integrase binding domain (IBD). Here we present structural characterization of IBD interactions with transcriptional repressor JPO2 and domesticated transposase PogZ, and show that the PogZ interaction is nearly identical to the interaction of LEDGF/p75 with MLL1. The interaction with the IBD is maintained by an intrinsically disordered IBD-binding motif (IBM) common to all known cellular partners of LEDGF/p75. In addition, based on IBM conservation, we identify and validate IWS1 as a novel LEDGF/p75 interaction partner. Our results also reveal how HIV integrase efficiently displaces cellular binding partners from LEDGF/p75. Finally, the similar binding modes of LEDGF/p75 interaction partners represent a new challenge for the development of selective interaction inhibitors. 1 Institute of Organic Chemistry and Biochemistry of the ASCR, v.v.i., Flemingovo nam. 2, 166 10 Prague, Czech Republic. 2 Department of Genetics and Microbiology, Faculty of Science, Charles University in Prague, Vinicna 5, 128 44 Prague, Czech Republic. -

Cytotoxic Effects and Changes in Gene Expression Profile
Toxicology in Vitro 34 (2016) 309–320 Contents lists available at ScienceDirect Toxicology in Vitro journal homepage: www.elsevier.com/locate/toxinvit Fusarium mycotoxin enniatin B: Cytotoxic effects and changes in gene expression profile Martina Jonsson a,⁎,MarikaJestoib, Minna Anthoni a, Annikki Welling a, Iida Loivamaa a, Ville Hallikainen c, Matti Kankainen d, Erik Lysøe e, Pertti Koivisto a, Kimmo Peltonen a,f a Chemistry and Toxicology Research Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, Finland b Product Safety Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, c The Finnish Forest Research Institute, Rovaniemi Unit, P.O. Box 16, FI-96301 Rovaniemi, Finland d Institute for Molecular Medicine Finland (FIMM), University of Helsinki, P.O. Box 20, FI-00014, Finland e Plant Health and Biotechnology, Norwegian Institute of Bioeconomy, Høyskoleveien 7, NO -1430 Ås, Norway f Finnish Safety and Chemicals Agency (Tukes), Opastinsilta 12 B, FI-00521 Helsinki, Finland article info abstract Article history: The mycotoxin enniatin B, a cyclic hexadepsipeptide produced by the plant pathogen Fusarium,isprevalentin Received 3 December 2015 grains and grain-based products in different geographical areas. Although enniatins have not been associated Received in revised form 5 April 2016 with toxic outbreaks, they have caused toxicity in vitro in several cell lines. In this study, the cytotoxic effects Accepted 28 April 2016 of enniatin B were assessed in relation to cellular energy metabolism, cell proliferation, and the induction of ap- Available online 6 May 2016 optosis in Balb 3T3 and HepG2 cells. The mechanism of toxicity was examined by means of whole genome ex- fi Keywords: pression pro ling of exposed rat primary hepatocytes. -

Nup98-Dependent Transcriptional Memory Is Established Independently of Transcription
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.12.336073; this version posted October 12, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Nup98-dependent transcriptional memory is established independently of transcription 2 3 Pau Pascual-Garcia, Shawn C. Little* and Maya Capelson*# 4 5 Department of Cell and Developmental Biology, Penn Epigenetics Institute, Perelman School of 6 Medicine, University of Pennsylvania, Philadelphia, PA, 19104, USA. 7 8 *co-corresponding authors 9 #contact author: [email protected] 10 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.10.12.336073; this version posted October 12, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 11 Summary 12 Cellular ability to mount an enhanced transcriptional response upon repeated exposure to external 13 cues has been termed transcriptional memory, which can be maintained epigenetically through cell 14 divisions. The majority of mechanistic knowledge on transcriptional memory has been derived from 15 bulk molecular assays, and this phenomenon has been found to depend on a nuclear pore 16 component Nup98 in multiple species. To gain an alternative perspective on the mechanism and 17 on the contribution of Nup98, we set out to examine single-cell population dynamics of 18 transcriptional memory by monitoring transcriptional behavior of individual Drosophila cells upon 19 initial and subsequent exposures to steroid hormone ecdysone. -

Recognition of Cancer Mutations in Histone H3K36 by Epigenetic Writers and Readers Brianna J
EPIGENETICS https://doi.org/10.1080/15592294.2018.1503491 REVIEW Recognition of cancer mutations in histone H3K36 by epigenetic writers and readers Brianna J. Kleina, Krzysztof Krajewski b, Susana Restrepoa, Peter W. Lewis c, Brian D. Strahlb, and Tatiana G. Kutateladzea aDepartment of Pharmacology, University of Colorado School of Medicine, Aurora, CO, USA; bDepartment of Biochemistry & Biophysics, The University of North Carolina School of Medicine, Chapel Hill, NC, USA; cWisconsin Institute for Discovery, University of Wisconsin, Madison, WI, USA ABSTRACT ARTICLE HISTORY Histone posttranslational modifications control the organization and function of chromatin. In Received 30 May 2018 particular, methylation of lysine 36 in histone H3 (H3K36me) has been shown to mediate gene Revised 1 July 2018 transcription, DNA repair, cell cycle regulation, and pre-mRNA splicing. Notably, mutations at or Accepted 12 July 2018 near this residue have been causally linked to the development of several human cancers. These KEYWORDS observations have helped to illuminate the role of histones themselves in disease and to clarify Histone; H3K36M; cancer; the mechanisms by which they acquire oncogenic properties. This perspective focuses on recent PTM; methylation advances in discovery and characterization of histone H3 mutations that impact H3K36 methyla- tion. We also highlight findings that the common cancer-related substitution of H3K36 to methionine (H3K36M) disturbs functions of not only H3K36me-writing enzymes but also H3K36me-specific readers. The latter case suggests that the oncogenic effects could also be linked to the inability of readers to engage H3K36M. Introduction from yeast to humans and has been shown to have a variety of functions that range from the control Histone proteins are main components of the of gene transcription and DNA repair, to cell cycle nucleosome, the fundamental building block of regulation and nutrient stress response [8]. -

Loss of Function SETD2 Mutations in Poorly Differentiated Metastases
cancers Article Loss of Function SETD2 Mutations in Poorly Differentiated Metastases from Two Hürthle Cell Carcinomas of the Thyroid 1, 1, 1 2 1 Valeria Pecce y , Antonella Verrienti y, Luana Abballe , Raffaella Carletti , Giorgio Grani , Rosa Falcone 1, Valeria Ramundo 1, Cosimo Durante 1, Cira Di Gioia 2 , Diego Russo 3 , Sebastiano Filetti 1 and Marialuisa Sponziello 1,* 1 Department of Translational and Precision Medicine, “Sapienza” University of Rome, 00161 Rome, Italy; [email protected] (V.P.); [email protected] (A.V.); [email protected] (L.A.); [email protected] (G.G.); [email protected] (R.F.); [email protected] (V.R.); [email protected] (C.D.); sebastiano.fi[email protected] (S.F.) 2 Department of Radiological, Oncological and Pathological Sciences, “Sapienza” University of Rome, 00161 Rome, Italy; raff[email protected] (R.C.); [email protected] (C.D.G.) 3 Department of Health Sciences, “Magna Graecia” University of Catanzaro, 88100 Catanzaro, Italy; [email protected] * Correspondence: [email protected] These authors contributed equally to this work. y Received: 23 June 2020; Accepted: 11 July 2020; Published: 14 July 2020 Abstract: Hürthle cell carcinomas (HCC) are rare differentiated thyroid cancers that display low avidity for radioactive iodine and respond poorly to kinase inhibitors. Here, using next-generation sequencing, we analyzed the mutational status of primary tissue and poorly differentiated metastatic tissue from two HCC patients. In both cases, metastatic tissues harbored a mutation of SETD2, each resulting in loss of the SRI and WW domains of SETD2, a methyltransferase that trimethylates H3K36 (H3K36me3) and also interacts with p53 to promote its stability. -

SETD2 Mutations in Primary Central Nervous System Tumors Angela N
Viaene et al. Acta Neuropathologica Communications (2018) 6:123 https://doi.org/10.1186/s40478-018-0623-0 RESEARCH Open Access SETD2 mutations in primary central nervous system tumors Angela N. Viaene1, Mariarita Santi1, Jason Rosenbaum2, Marilyn M. Li1, Lea F. Surrey1 and MacLean P. Nasrallah2,3* Abstract Mutations in SETD2 are found in many tumors, including central nervous system (CNS) tumors. Previous work has shown these mutations occur specifically in high grade gliomas of the cerebral hemispheres in pediatric and young adult patients. We investigated SETD2 mutations in a cohort of approximately 640 CNS tumors via next generation sequencing; 23 mutations were detected across 19 primary CNS tumors. Mutations were found in a wide variety of tumors and locations at a broad range of allele frequencies. SETD2 mutations were seen in both low and high grade gliomas as well as non-glial tumors, and occurred in patients greater than 55 years of age, in addition to pediatric and young adult patients. High grade gliomas at first occurrence demonstrated either frameshift/truncating mutations or point mutations at high allele frequencies, whereas recurrent high grade gliomas frequently harbored subclones with point mutations in SETD2 at lower allele frequencies in the setting of higher mutational burdens. Comparison with the TCGA dataset demonstrated consistent findings. Finally, immunohistochemistry showed decreased staining for H3K36me3 in our cohort of SETD2 mutant tumors compared to wildtype controls. Our data further describe the spectrum of tumors in which SETD2 mutations are found and provide a context for interpretation of these mutations in the clinical setting. Keywords: SETD2, Histone, Brain tumor, Glioma, Epigenetics, H3K36me3 Introduction (H3K36me3) in humans. -

Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain a Target Enabling Package (TEP)
Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain A Target Enabling Package (TEP) Gene ID / UniProt ID / EC CDK12, 51755 / Q9NYV4/ 2.7.11.22, 2.7.11.23 CCNK, 8812 / O75909/ - Target Nominator Gregg Morin (UBC, Canada), Nathanael Gray (Harvard) SGC Authors Sarah E. Dixon-Clarke, Jonathan M. Elkins, Nathanael S. Gray, and Alex N. Bullock Collaborating Authors S.-W. Grace Cheng1, Tinghu Zhang2, Nicholas Kwiatkowski3, Calla M. Olson2, Brian J. Abraham3, Ann K. Greifenberg4, Scott B. Ficarro2, Yanke Liang2, Nancy M. Hannett3, Theresa Manz5, Mingfeng Hao2, Bartlomiej Bartkowiak6, Arno L. Greenleaf6, Jarrod A. Marto2, Matthias Geyer4, Richard A. Young3, and Gregg B. Morin1 Target PI Alex Bullock (SGC Oxford) Therapeutic Area(s) Oncology Disease Relevance CDK12 loss sensitises cancer cells to DNA damage Date approved by TEP 17th June 2016 Evaluation Group Document version Version 3 Document version date April 2018 DOI https://doi.org/10.5281/zenodo.1219680 Affiliations 1. Canada's Michael Smith Genome Sciences Centre, BC Cancer Agency 2. Department of Cancer Biology, Dana-Farber Cancer Institute 3. Whitehead Institute for Biomedical Research 4. Department of Structural Immunology, Institute of Innate Immunity 5. Pharmaceutical and Medicinal Chemistry, Saarland University 6. Department of Biochemistry, Duke University Medical Center USEFUL LINKS (Please note that the inclusion of links to external sites should not be taken as an endorsement of that site by the SGC in any way) SUMMARY OF PROJECT Cyclin-dependent kinase 12 (CDK12) phosphorylates RNA Pol II C-terminal domain (CTD) to promote transcriptional elongation of large DNA damage response genes. CDK12 is frequently mutated or amplified in cancer and its loss sensitises cells to DNA damage. -

Rho Dependent Termination of Transcription in Prokaryotes
Rho Dependent Termination Of Transcription In Prokaryotes Organoleptic Tymon scrimshaws no athenaeums yean fleetly after Teodorico whiled tunably, quite scrawliest. Amalgamative and honeyed Ximenes syllables her bullheads picket stork's-bill and debouch visually. Colory Jeffery concentred her avidity so injudiciously that Sergio briquet very injunctively. Dna in prokaryotic polymerase in termination may vary. Dna sequence by enzymes and fall off of transcripts, and marylène bertrand for the conformation of rho in the stalled rna helicases, rho dependent termination of in transcription. The other physiological functions are the basis for rna polymerase and longer as elongation complex forming a series of one of rho termination in transcription factors recognizing promoters in four steps. Eukaryotes require cookies from prokaryotes, depending on a fifth subunit involved. Thus, when translation termination occurs within same gene it the cause transcriptional termination, preventing expression of downstream genes. Atp synthase alpha and therefore be specialized rnas make a bsr, in termination of rho transcription prokaryotes. Dna must accept cookies and also found later in part because of rho termination occurs by the initiation site that bind to understand how they cannot be logged in replicating dna. Concerned about the Coronavirus? It is made rna polymerase causes rna in termination of rho transcription termination mechanisms to specific genes. Depending upon request your changes indicating that prokaryotic rnap with and prokaryotes. The DNA strand within this an is transcribed by the RNA polymerase. The basic promoter region in prokaryotic transcription is referred to two the Pribnow box. Formation of which prokaryotic and on the hfq is shown as the transcript of action of rho dependent termination transcription in prokaryotes have far more about ten base that elongation complex cell components and cell. -

The Function and Evolution of C2H2 Zinc Finger Proteins and Transposons
The function and evolution of C2H2 zinc finger proteins and transposons by Laura Francesca Campitelli A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Department of Molecular Genetics University of Toronto © Copyright by Laura Francesca Campitelli 2020 The function and evolution of C2H2 zinc finger proteins and transposons Laura Francesca Campitelli Doctor of Philosophy Department of Molecular Genetics University of Toronto 2020 Abstract Transcription factors (TFs) confer specificity to transcriptional regulation by binding specific DNA sequences and ultimately affecting the ability of RNA polymerase to transcribe a locus. The C2H2 zinc finger proteins (C2H2 ZFPs) are a TF class with the unique ability to diversify their DNA-binding specificities in a short evolutionary time. C2H2 ZFPs comprise the largest class of TFs in Mammalian genomes, including nearly half of all Human TFs (747/1,639). Positive selection on the DNA-binding specificities of C2H2 ZFPs is explained by an evolutionary arms race with endogenous retroelements (EREs; copy-and-paste transposable elements), where the C2H2 ZFPs containing a KRAB repressor domain (KZFPs; 344/747 Human C2H2 ZFPs) are thought to diversify to bind new EREs and repress deleterious transposition events. However, evidence of the gain and loss of KZFP binding sites on the ERE sequence is sparse due to poor resolution of ERE sequence evolution, despite the recent publication of binding preferences for 242/344 Human KZFPs. The goal of my doctoral work has been to characterize the Human C2H2 ZFPs, with specific interest in their evolutionary history, functional diversity, and coevolution with LINE EREs.