Car-Modified T Cells Capable of Distinguishing Normal Cells from Malignant Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

ENSG Gene Encodes Effector TCR Pathway Costimulation Inhibitory/Exhaustion Synapse/Adhesion Chemokines/Receptors
ENSG Gene Encodes Effector TCR pathway Costimulation Inhibitory/exhaustion Synapse/adhesion Chemokines/receptors ENSG00000111537 IFNG IFNg x ENSG00000109471 IL2 IL-2 x ENSG00000232810 TNF TNFa x ENSG00000271503 CCL5 CCL5 x x ENSG00000139187 KLRG1 Klrg1 x ENSG00000117560 FASLG Fas ligand x ENSG00000121858 TNFSF10 TRAIL x ENSG00000134545 KLRC1 Klrc1 / NKG2A x ENSG00000213809 KLRK1 Klrk1 / NKG2D x ENSG00000188389 PDCD1 PD-1 x x ENSG00000117281 CD160 CD160 x x ENSG00000134460 IL2RA IL-2 receptor x subunit alpha ENSG00000110324 IL10RA IL-10 receptor x subunit alpha ENSG00000115604 IL18R1 IL-18 receptor 1 x ENSG00000115607 IL18RAP IL-18 receptor x accessory protein ENSG00000081985 IL12RB2 IL-12 receptor x beta 2 ENSG00000186810 CXCR3 CXCR3 x x ENSG00000005844 ITGAL CD11a x ENSG00000160255 ITGB2 CD18; Integrin x x beta-2 ENSG00000156886 ITGAD CD11d x ENSG00000140678 ITGAX; CD11c x x Integrin alpha-X ENSG00000115232 ITGA4 CD49d; Integrin x x alpha-4 ENSG00000169896 ITGAM CD11b; Integrin x x alpha-M ENSG00000138378 STAT4 Stat4 x ENSG00000115415 STAT1 Stat1 x ENSG00000170581 STAT2 Stat2 x ENSG00000126561 STAT5a Stat5a x ENSG00000162434 JAK1 Jak1 x ENSG00000100453 GZMB Granzyme B x ENSG00000145649 GZMA Granzyme A x ENSG00000180644 PRF1 Perforin 1 x ENSG00000115523 GNLY Granulysin x ENSG00000100450 GZMH Granzyme H x ENSG00000113088 GZMK Granzyme K x ENSG00000057657 PRDM1 Blimp-1 x ENSG00000073861 TBX21 T-bet x ENSG00000115738 ID2 ID2 x ENSG00000176083 ZNF683 Hobit x ENSG00000137265 IRF4 Interferon x regulatory factor 4 ENSG00000140968 IRF8 Interferon -

The TNF and TNF Receptor Review Superfamilies: Integrating Mammalian Biology
Cell, Vol. 104, 487±501, February 23, 2001, Copyright 2001 by Cell Press The TNF and TNF Receptor Review Superfamilies: Integrating Mammalian Biology Richard M. Locksley,*²³k Nigel Killeen,²k The receptors and ligands in this superfamily have and Michael J. Lenardo§k unique structural attributes that couple them directly to *Department of Medicine signaling pathways for cell proliferation, survival, and ² Department of Microbiology and Immunology differentiation. Thus, they have assumed prominent ³ Howard Hughes Medical Institute roles in the generation of tissues and transient microen- University of California, San Francisco vironments. Most TNF/TNFR SFPs are expressed in the San Francisco, California 94143 immune system, where their rapid and potent signaling § Laboratory of Immunology capabilities are crucial in coordinating the proliferation National Institute of Allergy and Infectious Diseases and protective functions of pathogen-reactive cells. National Institutes of Health Here, we review the organization of the TNF/TNFR SF Bethesda, Maryland 20892 and how these proteins have been adapted for pro- cesses as seemingly disparate as host defense and or- ganogenesis. In interpreting this large and highly active Introduction area of research, we have focused on common themes that unite the actions of these genes in different tissues. Three decades ago, lymphotoxin (LT) and tumor necro- We also discuss the evolutionary success of this super- sis factor (TNF) were identified as products of lympho- familyÐsuccess that we infer from its expansion across cytes and macrophages that caused the lysis of certain the mammalian genome and from its many indispens- types of cells, especially tumor cells (Granger et al., able roles in mammalian biology. -

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

Tools for Cell Therapy and Immunoregulation
RnDSy-lu-2945 Tools for Cell Therapy and Immunoregulation Target Cell TIM-4 SLAM/CD150 BTNL8 PD-L2/B7-DC B7-H1/PD-L1 (Human) Unknown PD-1 B7-1/CD80 TIM-1 SLAM/CD150 Receptor TIM Family SLAM Family Butyrophilins B7/CD28 Families T Cell Multiple Co-Signaling Molecules Co-stimulatory Co-inhibitory Ig Superfamily Regulate T Cell Activation Target Cell T Cell Target Cell T Cell B7-1/CD80 B7-H1/PD-L1 T cell activation requires two signals: 1) recognition of the antigenic peptide/ B7-1/CD80 B7-2/CD86 CTLA-4 major histocompatibility complex (MHC) by the T cell receptor (TCR) and 2) CD28 antigen-independent co-stimulation induced by interactions between B7-2/CD86 B7-H1/PD-L1 B7-1/CD80 co-signaling molecules expressed on target cells, such as antigen-presenting PD-L2/B7-DC PD-1 ICOS cells (APCs), and their T cell-expressed receptors. Engagement of the TCR in B7-H2/ICOS L 2Ig B7-H3 (Mouse) the absence of this second co-stimulatory signal typically results in T cell B7-H1/PD-L1 B7/CD28 Families 4Ig B7-H3 (Human) anergy or apoptosis. In addition, T cell activation can be negatively regulated Unknown Receptors by co-inhibitory molecules present on APCs. Therefore, integration of the 2Ig B7-H3 Unknown B7-H4 (Mouse) Receptors signals transduced by co-stimulatory and co-inhibitory molecules following TCR B7-H5 4Ig B7-H3 engagement directs the outcome and magnitude of a T cell response Unknown Ligand (Human) B7-H5 including the enhancement or suppression of T cell proliferation, B7-H7 Unknown Receptor differentiation, and/or cytokine secretion. -

Prediction of Distant Metastasis by Using Reverse Transcriptase Polymerase Chain Reaction for Epithelial and Variant CD44 Mrna I
ORIGINAL ARTICLE Prediction of Distant Metastasis by Using Reverse Transcriptase–Polymerase Chain Reaction for Epithelial and Variant CD44 mRNA in the Peripheral Blood of Patients With Colorectal Cancer Shozo Yokoyama, MD; Hiroki Yamaue, MD Background: Reverse transcriptase–polymerase chain Main Outcome Measure: The clinical significance of reaction (RT-PCR) has been used to identify small num- RT-PCR for epithelial and variant CD44 mRNA in pe- bers of tumor cells. Molecular detection is thought to pro- ripheral blood. vide useful information for the clinical management of postoperative adjuvant therapy regimens. Results: During 3 years of follow-up, 2 patients whose peripheral blood had carcinoembryonic antigen and Objective: To use RT-PCR to identify messenger RNA CD44 variant mRNA also had distant metastases (lung (mRNA) coding for carcinoembryonic antigen, epithelial or spleen). Expression of epithelial and variant CD44 and variant CD44, and matrix metalloproteinase 7 in the mRNA in peripheral blood was more highly correlated portal venous and peripheral blood of patients with colo- with the clinical cancer stage than with expression rectal carcinoma to predict live or distant metastasis. of carcinoembryonic antigen and matrix metallopro- teinase 7. Design: Prospective consecutive series. Conclusions: Molecular detection of epithelial and Setting: University hospital. variant CD44 mRNA in the peripheral blood may help determine distant metastases in patients with colorectal Patients and Methods: Portal venous and peripheral carcinoma. Molecular detection in the peripheral blood blood samples were obtained from 22 patients with colo- at surgical treatment suggests that systemic hemato- rectal cancer during surgical manipulation. Using comple- genic tumor cell dissemination is an early event of dis- mentary DNA primers specific for carcinoembryonic tant metastasis. -
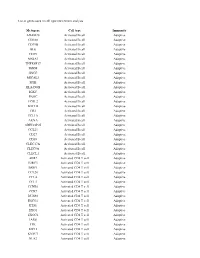
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

Propranolol-Mediated Attenuation of MMP-9 Excretion in Infants with Hemangiomas
Supplementary Online Content Thaivalappil S, Bauman N, Saieg A, Movius E, Brown KJ, Preciado D. Propranolol-mediated attenuation of MMP-9 excretion in infants with hemangiomas. JAMA Otolaryngol Head Neck Surg. doi:10.1001/jamaoto.2013.4773 eTable. List of All of the Proteins Identified by Proteomics This supplementary material has been provided by the authors to give readers additional information about their work. © 2013 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 10/01/2021 eTable. List of All of the Proteins Identified by Proteomics Protein Name Prop 12 mo/4 Pred 12 mo/4 Δ Prop to Pred mo mo Myeloperoxidase OS=Homo sapiens GN=MPO 26.00 143.00 ‐117.00 Lactotransferrin OS=Homo sapiens GN=LTF 114.00 205.50 ‐91.50 Matrix metalloproteinase‐9 OS=Homo sapiens GN=MMP9 5.00 36.00 ‐31.00 Neutrophil elastase OS=Homo sapiens GN=ELANE 24.00 48.00 ‐24.00 Bleomycin hydrolase OS=Homo sapiens GN=BLMH 3.00 25.00 ‐22.00 CAP7_HUMAN Azurocidin OS=Homo sapiens GN=AZU1 PE=1 SV=3 4.00 26.00 ‐22.00 S10A8_HUMAN Protein S100‐A8 OS=Homo sapiens GN=S100A8 PE=1 14.67 30.50 ‐15.83 SV=1 IL1F9_HUMAN Interleukin‐1 family member 9 OS=Homo sapiens 1.00 15.00 ‐14.00 GN=IL1F9 PE=1 SV=1 MUC5B_HUMAN Mucin‐5B OS=Homo sapiens GN=MUC5B PE=1 SV=3 2.00 14.00 ‐12.00 MUC4_HUMAN Mucin‐4 OS=Homo sapiens GN=MUC4 PE=1 SV=3 1.00 12.00 ‐11.00 HRG_HUMAN Histidine‐rich glycoprotein OS=Homo sapiens GN=HRG 1.00 12.00 ‐11.00 PE=1 SV=1 TKT_HUMAN Transketolase OS=Homo sapiens GN=TKT PE=1 SV=3 17.00 28.00 ‐11.00 CATG_HUMAN Cathepsin G OS=Homo -

Pan-KIR2DL NK-Receptor Antibodies and Their Use in Diagnostik and Therapy
(19) TZZ _ T (11) EP 2 287 195 A2 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 23.02.2011 Bulletin 2011/08 C07K 16/28 (2006.01) G01N 33/52 (2006.01) (21) Application number: 10178924.6 (22) Date of filing: 01.07.2005 (84) Designated Contracting States: • Romagne, François AT BE BG CH CY CZ DE DK EE ES FI FR GB GR 13600 La Ciotat (FR) HU IE IS IT LI LT LU LV MC NL PL PT RO SE SI • Wagtmann, Peter, Andreas, Nicolai, Reumert SK TR 2960 Rungsted Kyst (DK) • Svendsen, Ivan (30) Priority: 06.01.2005 DK 200500025 2765 Smorum (DK) 01.07.2004 PCT/DK2004/000470 • Zahn, Stefan 01.07.2004 PCT/IB2004/002464 2750 Ballerup (DK) • Svensson, Anders (62) Document number(s) of the earlier application(s) in 21746 Malmö (DK) accordance with Art. 76 EPC: • Thorolfsson, Matthias 05758642.2 / 1 791 868 2920 Charlottenlund (DK) • Berg Padkaer, Soren (71) Applicants: 3500 Vaerlose (DK) • Novo Nordisk A/S • Kjaergaard, Kristian 2880 Bagsvaerd (DK) 2750 Ballerup (DK) • Innate Pharma • Spee, Pieter 13009 Marseille (FR) 3450 Allerod (DK) • Universita di Genova • Wilken, Michael 16132 Genova (IT) 3390 Hundested (DK) (72) Inventors: (74) Representative: Gallois, Valérie et al • Moretta, Alessandro Cabinet BECKER & ASSOCIES 16133 Genova (IT) 25, rue Louis Le Grand • Della Chiesa, Mariella 75002 Paris (FR) 16132 Genova (IT) • Andre, Pascale Remarks: 13006 Marseille (FR) This application was filed on 23-09-2010 as a • Gauthier, Laurent divisional application to the application mentioned 13008 Marseille (FR) under INID code 62. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

CD44 Predicts Early Recurrence in Pancreatic Cancer Patients Undergoing Radical Surgery
in vivo 32 : 1533-1540 (2018) doi:10.21873/invivo.11411 CD44 Predicts Early Recurrence in Pancreatic Cancer Patients Undergoing Radical Surgery CHIH-PO HSU 1* , LI-YU LEE 2* , JUN-TE HSU 1, YU-PAO HSU 1, YU-TUNG WU 1, SHANG-YU WANG 1, CHUN-NAN YEH 1, TSE-CHING CHEN 2 and TSANN-LONG HWANG 1 1Department of General Surgery, Chang Gung Memorial Hospital at Linkou, Chang Gung University College of Medicine, Taoyuan, Taiwan, R.O.C.; 2Department of Pathology, Chang Gung Memorial Hospital at Linkou, Chang Gung University College of Medicine, Taoyuan, Taiwan, R.O.C. Abstract. Background/Aim: Pancreatic ductal adeno- predicted ER. Conclusion: High CA19-9 levels, CD44 H- carcinoma (PDAC) is one of the most aggressive types of scores and poor differentiation are independent predictors digestive cancer. Recurrence within one year after surgery is for ER in PDAC patients undergoing radical resection. inevitable in most PDAC patients. Recently, cluster of Therefore, the determination of CD44 expression might help differentiation 44 (CD44) has been shown to be associated in identifying patients at a high risk of ER for more with tumor initiation, metastasis and prognosis. This study aggressive treatment after radical surgery. aimed to explore the correlation of CD44 expression with clinicopathological factors and the role of CD44 in Pancreatic ductal adenocarcinoma (PDAC) is the fourth most predicting early recurrence (ER) in PDAC patients after common cause of cancer-related death worldwide, with an radical surgery. Materials and Methods: PDAC patients who 8% 5-year survival rate for all stages of disease (1). underwent radical resection between January 1999 and Although various treatment modalities are available, only March 2015 were enrolled in this study. -

Human TNFRSF18 ELISA Kit (ARG81453)
Product datasheet [email protected] ARG81453 Package: 96 wells Human TNFRSF18 ELISA Kit Store at: 4°C Component Cat. No. Component Name Package Temp ARG81453-001 Antibody-coated 8 X 12 strips 4°C. Unused strips microplate should be sealed tightly in the air-tight pouch. ARG81453-002 Standard 2 X 10 ng/vial 4°C ARG81453-003 Standard/Sample 30 ml (Ready to use) 4°C diluent ARG81453-004 Antibody conjugate 1 vial (100 µl) 4°C concentrate (100X) ARG81453-005 Antibody diluent 12 ml (Ready to use) 4°C buffer ARG81453-006 HRP-Streptavidin 1 vial (100 µl) 4°C concentrate (100X) ARG81453-007 HRP-Streptavidin 12 ml (Ready to use) 4°C diluent buffer ARG81453-008 25X Wash buffer 20 ml 4°C ARG81453-009 TMB substrate 10 ml (Ready to use) 4°C (Protect from light) ARG81453-010 STOP solution 10 ml (Ready to use) 4°C ARG81453-011 Plate sealer 4 strips Room temperature Summary Product Description ARG81453 Human TNFRSF18 ELISA Kit is an Enzyme Immunoassay kit for the quantification of Human TNFRSF18 in serum, plasma (heparin, EDTA) and cell culture supernatants. Tested Reactivity Hu Tested Application ELISA Specificity There is no detectable cross-reactivity with other relevant proteins. Target Name TNFRSF18 Conjugation HRP Conjugation Note Substrate: TMB and read at 450 nm. Sensitivity 31.25 pg/ml Sample Type Serum, plasma (heparin, EDTA) and cell culture supernatants. Standard Range 62.5 - 4000 pg/ml Sample Volume 100 µl www.arigobio.com 1/3 Precision Intra-Assay CV: 6.3% Inter-Assay CV: 7.0% Alternate Names Tumor necrosis factor receptor superfamily member 18; AITR; CD357; CD antigen CD357; Activation- inducible TNFR family receptor; GITR-D; GITR; Glucocorticoid-induced TNFR-related protein Application Instructions Assay Time ~ 5 hours Properties Form 96 well Storage instruction Store the kit at 2-8°C. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like,