Genetic Variants in the Transcription Regulatory Region of MEGF10 Are
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Detailed Characterization of Human Induced Pluripotent Stem Cells Manufactured for Therapeutic Applications
Stem Cell Rev and Rep DOI 10.1007/s12015-016-9662-8 Detailed Characterization of Human Induced Pluripotent Stem Cells Manufactured for Therapeutic Applications Behnam Ahmadian Baghbaderani 1 & Adhikarla Syama2 & Renuka Sivapatham3 & Ying Pei4 & Odity Mukherjee2 & Thomas Fellner1 & Xianmin Zeng3,4 & Mahendra S. Rao5,6 # The Author(s) 2016. This article is published with open access at Springerlink.com Abstract We have recently described manufacturing of hu- help determine which set of tests will be most useful in mon- man induced pluripotent stem cells (iPSC) master cell banks itoring the cells and establishing criteria for discarding a line. (MCB) generated by a clinically compliant process using cord blood as a starting material (Baghbaderani et al. in Stem Cell Keywords Induced pluripotent stem cells . Embryonic stem Reports, 5(4), 647–659, 2015). In this manuscript, we de- cells . Manufacturing . cGMP . Consent . Markers scribe the detailed characterization of the two iPSC clones generated using this process, including whole genome se- quencing (WGS), microarray, and comparative genomic hy- Introduction bridization (aCGH) single nucleotide polymorphism (SNP) analysis. We compare their profiles with a proposed calibra- Induced pluripotent stem cells (iPSCs) are akin to embryonic tion material and with a reporter subclone and lines made by a stem cells (ESC) [2] in their developmental potential, but dif- similar process from different donors. We believe that iPSCs fer from ESC in the starting cell used and the requirement of a are likely to be used to make multiple clinical products. We set of proteins to induce pluripotency [3]. Although function- further believe that the lines used as input material will be used ally identical, iPSCs may differ from ESC in subtle ways, at different sites and, given their immortal status, will be used including in their epigenetic profile, exposure to the environ- for many years or even decades. -

Brain Region-Specific Gene Signatures Revealed by Distinct Astrocyte Subpopulations Unveil Links to Glioma and Neurodegenerative
New Research Disorders of the Nervous System Brain Region-Specific Gene Signatures Revealed by Distinct Astrocyte Subpopulations Unveil Links to Glioma and Neurodegenerative Diseases Raquel Cuevas-Diaz Duran,1,2,3 Chih-Yen Wang,4 Hui Zheng,5,6,7 Benjamin Deneen,8,9,10,11 and Jia Qian Wu1,2 https://doi.org/10.1523/ENEURO.0288-18.2019 1The Vivian L. Smith Department of Neurosurgery, McGovern Medical School, University of Texas Health Science Center at Houston, Houston, Texas 77030, 2Center for Stem Cell and Regenerative Medicine, UT Brown Foundation Institute of Molecular Medicine, Houston, Texas 77030, 3Tecnologico de Monterrey, Escuela de Medicina y Ciencias de la Salud, Monterrey NL 64710, Mexico, 4Department of Life Sciences, National Cheng Kung University, Tainan City 70101, Taiwan, 5Huffington Center on Aging, 6Medical Scientist Training Program, 7Department of Molecular and Human Genetics, 8Center for Cell and Gene Therapy, 9Department of Neuroscience, 10Neurological Research Institute at Texas’ Children’s Hospital, and 11Program in Developmental Biology, Baylor College of Medicine, Houston, Texas 77030 Abstract Currently, there are no effective treatments for glioma or for neurodegenerative diseases because of, in part, our limited understanding of the pathophysiology and cellular heterogeneity of these diseases. Mounting evidence suggests that astrocytes play an active role in the pathogenesis of these diseases by contributing to a diverse range of pathophysiological states. In a previous study, five molecularly distinct astrocyte subpopulations from three different brain regions were identified. To further delineate the underlying diversity of these populations, we obtained mouse brain region-specific gene signatures for both protein-coding and long non-coding RNA and found that these astrocyte subpopulations are endowed with unique molecular signatures across diverse brain regions. -

Identification of Shared and Unique Gene Families Associated with Oral
International Journal of Oral Science (2017) 9, 104–109 OPEN www.nature.com/ijos ORIGINAL ARTICLE Identification of shared and unique gene families associated with oral clefts Noriko Funato and Masataka Nakamura Oral clefts, the most frequent congenital birth defects in humans, are multifactorial disorders caused by genetic and environmental factors. Epidemiological studies point to different etiologies underlying the oral cleft phenotypes, cleft lip (CL), CL and/or palate (CL/P) and cleft palate (CP). More than 350 genes have syndromic and/or nonsyndromic oral cleft associations in humans. Although genes related to genetic disorders associated with oral cleft phenotypes are known, a gap between detecting these associations and interpretation of their biological importance has remained. Here, using a gene ontology analysis approach, we grouped these candidate genes on the basis of different functional categories to gain insight into the genetic etiology of oral clefts. We identified different genetic profiles and found correlations between the functions of gene products and oral cleft phenotypes. Our results indicate inherent differences in the genetic etiologies that underlie oral cleft phenotypes and support epidemiological evidence that genes associated with CL/P are both developmentally and genetically different from CP only, incomplete CP, and submucous CP. The epidemiological differences among cleft phenotypes may reflect differences in the underlying genetic causes. Understanding the different causative etiologies of oral clefts is -

Regulation of Muscle Stem Cell Function by the Transcription Factor Pax7
Regulation of Muscle Stem Cell Function by the Transcription Factor Pax7 Alessandra Pasut Thesis submitted to the Faculty of Graduate and Postdoctoral Studies in partial fulfillment of the requirements for the Doctorate in Philosophy degree in Cellular and Molecular Medicine Department of Cellular and Molecular Medicine Faculty of Medicine University of Ottawa © Alessandra Pasut, Ottawa, Canada, 2015 ...above all don’t fear difficult moments, the best comes out from them. [Rita Levi Montalcini-Nobel laureate, 1909-2012] ii Abstract Pax7 is a paired box transcription factor expressed by all satellite cells which are critically required for muscle regeneration and growth. The absolute requirements of Pax7 in the maintenance of the satellite cell pool are widely acknowledged. However the mechanisms by which Pax7 executes muscle regeneration or contributes to satellite cell homeostasis remain elusive. We performed cell and molecular analysis of Pax7 null satellite cells to investigate muscle stem cell function. Through genome wide studies, we found that genes involved in cell- cell interactions, regulation of migration, control of lipid metabolism and inhibition of myogenic differentiation were significantly perturbed in Pax7 null satellite cells. Analysis of satellite cells in vitro showed that Pax7 null satellite cells undergo precocious myogenic differentiation and have perturbed expression of genes involved in the Notch signaling pathway. We showed that Notch 1 is a novel Pax7 target gene and by using a genetic approach we demonstrate that ectopic expression of the constitutively active intracellular domain of Notch1 (NICD1) in Pax7 null satellite cells is sufficient to maintain the satellite cell pool as well as to restore their proliferation. -

A Genomic Approach to Delineating the Occurrence of Scoliosis in Arthrogryposis Multiplex Congenita
G C A T T A C G G C A T genes Article A Genomic Approach to Delineating the Occurrence of Scoliosis in Arthrogryposis Multiplex Congenita Xenia Latypova 1, Stefan Giovanni Creadore 2, Noémi Dahan-Oliel 3,4, Anxhela Gjyshi Gustafson 2, Steven Wei-Hung Hwang 5, Tanya Bedard 6, Kamran Shazand 2, Harold J. P. van Bosse 5 , Philip F. Giampietro 7,* and Klaus Dieterich 8,* 1 Grenoble Institut Neurosciences, Université Grenoble Alpes, Inserm, U1216, CHU Grenoble Alpes, 38000 Grenoble, France; [email protected] 2 Shriners Hospitals for Children Headquarters, Tampa, FL 33607, USA; [email protected] (S.G.C.); [email protected] (A.G.G.); [email protected] (K.S.) 3 Shriners Hospitals for Children, Montreal, QC H4A 0A9, Canada; [email protected] 4 School of Physical & Occupational Therapy, Faculty of Medicine and Health Sciences, McGill University, Montreal, QC H3G 2M1, Canada 5 Shriners Hospitals for Children, Philadelphia, PA 19140, USA; [email protected] (S.W.-H.H.); [email protected] (H.J.P.v.B.) 6 Alberta Congenital Anomalies Surveillance System, Alberta Health Services, Edmonton, AB T5J 3E4, Canada; [email protected] 7 Department of Pediatrics, University of Illinois-Chicago, Chicago, IL 60607, USA 8 Institut of Advanced Biosciences, Université Grenoble Alpes, Inserm, U1209, CHU Grenoble Alpes, 38000 Grenoble, France * Correspondence: [email protected] (P.F.G.); [email protected] (K.D.) Citation: Latypova, X.; Creadore, S.G.; Dahan-Oliel, N.; Gustafson, Abstract: Arthrogryposis multiplex congenita (AMC) describes a group of conditions characterized A.G.; Wei-Hung Hwang, S.; Bedard, by the presence of non-progressive congenital contractures in multiple body areas. -

Computational Evaluation of Exome Sequence Data Using Human and Model Organism Phenotypes Improves Diagnostic Efficiency
ORIGINAL RESEARCH ARTICLE Official journal of the American College of Medical Genetics and Genomics Open Computational evaluation of exome sequence data using human and model organism phenotypes improves diagnostic efficiency William P. Bone1, Nicole L. Washington, PhD2, Orion J. Buske3,4, David R. Adams, MD, PhD1,5, Joie Davis1, David Draper1, Elise D. Flynn1, Marta Girdea3,4, Rena Godfrey1, Gretchen Golas1, Catherine Groden1, Julius Jacobsen, PhD6, Sebastian Köhler, PhD7, Elizabeth M. J. Lee1, Amanda E. Links1, Thomas C. Markello, MD, PhD1, Christopher J. Mungall, PhD2, Michele Nehrebecky1, Peter N. Robinson, MD7, Murat Sincan, MD1, Ariane G. Soldatos, MD1, Cynthia J. Tifft, MD, PhD1,5, Camilo Toro, MD1, Heather Trang3,4, Elise Valkanas1, Nicole Vasilevsky, PhD8, Colleen Wahl1, Lynne A. Wolfe1, Cornelius F. Boerkoel, MD, PhD1, Michael Brudno, PhD3,4, Melissa A. Haendel, PhD8, William A. Gahl, MD, PhD1,5, and Damian Smedley, PhD6 Purpose: Medical diagnosis and molecular or biochemical confir- Results: Benchmarking showed Exomiser ranked the causal variant mation typically rely on the knowledge of the clinician. Although as the top hit in 97% of known disease–gene associations and ranked this is very difficult in extremely rare diseases, we hypothesized that the correct seeded variant in up to 87% when detectable disease–gene the recording of patient phenotypes in Human Phenotype Ontology associations were unavailable. Using UDP data, Exomiser ranked (HPO) terms and computationally ranking putative disease-associ- the causative variant(s) within the top 10 variants for 11 previously ated sequence variants improves diagnosis, particularly for patients diagnosed variants and achieved a diagnosis for 4 of 23 cases undiag- with atypical clinical profiles. -

SARS-Cov-2 Infected Pediatric Cerebral Cortical Neurons: Transcriptomic Analysis and Potential Role of Toll-Like Receptors in Pathogenesis
International Journal of Molecular Sciences Article SARS-CoV-2 Infected Pediatric Cerebral Cortical Neurons: Transcriptomic Analysis and Potential Role of Toll-like Receptors in Pathogenesis Agnese Gugliandolo 1 , Luigi Chiricosta 1 , Valeria Calcaterra 2,3,† , Mara Biasin 4 , Gioia Cappelletti 4 , Stephana Carelli 5 , Gianvincenzo Zuccotti 2,4, Maria Antonietta Avanzini 6 , Placido Bramanti 1, Gloria Pelizzo 4,7 and Emanuela Mazzon 1,*,† 1 IRCCS Centro Neurolesi “Bonino-Pulejo”, Via Provinciale Palermo, Contrada Casazza, 98124 Messina, Italy; [email protected] (A.G.); [email protected] (L.C.); [email protected] (P.B.) 2 Department of Pediatrics, Ospedale dei Bambini “Vittore Buzzi”, 20154 Milano, Italy; [email protected] (V.C.); [email protected] (G.Z.) 3 Pediatrics and Adolescentology Unit, Department of Internal Medicine, University of Pavia, 27100 Pavia, Italy 4 Department of Biomedical and Clinical Sciences–L. Sacco, University of Milan, 20157 Milan, Italy; [email protected] (M.B.); [email protected] (G.C.); [email protected] (G.P.) 5 Pediatric Clinical Research Center Fondazione Romeo ed Enrica Invernizzi, University of Milan, 20157 Milan, Italy; [email protected] 6 Cell Factory, Pediatric Hematology Oncology Unit, Fondazione IRCCS Policlinico San Matteo, 27100 Pavia, Italy; [email protected] 7 Pediatric Surgery Unit, Ospedale dei Bambini “Vittore Buzzi”, 20154 Milano, Italy Citation: Gugliandolo, A.; Chiricosta, * Correspondence: [email protected] L.; Calcaterra, V.; Biasin, M.; † These authors contribute equally to the paper as senior author. Cappelletti, G.; Carelli, S.; Zuccotti, G.; Avanzini, M.A.; Bramanti, P.; Abstract: Different mechanisms were proposed as responsible for COVID-19 neurological symptoms Pelizzo, G.; et al. -

PARKINSON DISEASE LOCI in the MID-WESTERN AMISH by Mary
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Vanderbilt Electronic Thesis and Dissertation Archive PARKINSON DISEASE LOCI IN THE MID-WESTERN AMISH By Mary Feller Davis Thesis Submitted to the Faculty of the Graduate School of Vanderbilt University in partial fulfillment of the requirements for the degree of MASTER OF SCIENCE in Interdisciplinary Studies: Applied Statistics May, 2013 Nashville, Tennessee Approved: Professor Jonathan L. Haines Professor Marylyn D. Ritchie Professor Scott M. Williams ACKNOWLEDGEMENTS I would like to thank Anna Cummings, Laura D’Aoust, Lan Jian, Digna Velez Edwards, Renee Laux, Lori Reinhart-Mercer, Denise Fuzzell, William Scott, Margaret Pericak-Vance, Stephen Lee, and especially Jonathan Haines for their contributions to this project, as well as the participants of this study who have so graciously allowed us to visit with them and have participated in studies with us for over 10 years. I would like to acknowledge additional work for this study that was performed using the Vanderbilt Center for Human Genetics Research Core facilities: the Genetic Studies Ascertainment Core, the DNA Resources Core, and the Computation Genomics Core. This study was supported by the National Institutes of Health grants AG019085 (to Jonathan Haines and Margaret Pericak-Vance) and AG019726 (to William Scott), and a grant from the Michael J. Fox Foundation (to Jonathan Haines). Some of the samples used in this study were collected while William Scott and Margaret Pericak-Vance were faculty members at Duke University. I would like to thank L. L. McFarland, C. Knebusch, and the late C. -

Fine Mapping of a Linkage Peak with Integration of Lipid Traits Identifies
Nolan et al. BMC Genetics 2012, 13:12 http://www.biomedcentral.com/1471-2156/13/12 RESEARCHARTICLE Open Access Fine mapping of a linkage peak with integration of lipid traits identifies novel coronary artery disease genes on chromosome 5 Daniel K Nolan1, Beth Sutton1, Carol Haynes1, Jessica Johnson1, Jacqueline Sebek1, Elaine Dowdy1, David Crosslin1, David Crossman4, Michael H Sketch Jr2, Christopher B Granger2, David Seo3, Pascal Goldschmidt-Clermont3, William E Kraus2, Simon G Gregory1,2, Elizabeth R Hauser1,2 and Svati H Shah1,2* Abstract Background: Coronary artery disease (CAD), and one of its intermediate risk factors, dyslipidemia, possess a demonstrable genetic component, although the genetic architecture is incompletely defined. We previously reported a linkage peak on chromosome 5q31-33 for early-onset CAD where the strength of evidence for linkage was increased in families with higher mean low density lipoprotein-cholesterol (LDL-C). Therefore, we sought to fine-map the peak using association mapping of LDL-C as an intermediate disease-related trait to further define the etiology of this linkage peak. The study populations consisted of 1908 individuals from the CATHGEN biorepository of patients undergoing cardiac catheterization; 254 families (N = 827 individuals) from the GENECARD familial study of early-onset CAD; and 162 aorta samples harvested from deceased donors. Linkage disequilibrium-tagged SNPs were selected with an average of one SNP per 20 kb for 126.6-160.2 MB (region of highest linkage) and less dense spacing (one SNP per 50 kb) for the flanking regions (117.7-126.6 and 160.2-167.5 MB) and genotyped on all samples using a custom Illumina array. -
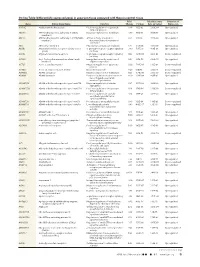
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

The Role of MEGF10 in Skeletal Muscle Myopathy
The role of MEGF10 in skeletal muscle myopathy. Thesis by Ruth Elizabeth Hughes Submitted in accordance with the requirements for the degree of Doctor of Philosophy The University of Leeds Faculty of Biological Sciences School of Molecular & Cellular Biology October 2016 -ii- The candidate confirms that the work submitted is his/her own and that appropriate credit has been given where reference has been made to the work of others. This copy has been supplied on the understanding that it is copyright material and that no quotation from the thesis may be published without proper acknowledgement. © 2016 The University of Leeds and Ruth Elizabeth Hughes -iii- Acknowledgements I am indebted to the Medical Research Council for their funding, and the Faculty of Biological Sciences for making my PhD possible. I would like to thank my supervisors Michelle Peckham and Colin Johnson for their patience, tireless enthusiasm and sage advice throughout the ups and downs of the project. I am grateful to Sara Cruz Migoni from the Borycki lab (Sheffield) for teaching me the single fibre isolation technique and Stuart Egginton and Roger Kissane for their help in setting up the hyperplasia model. I am thankful to Clare Logan and Gabrielle Wheway for their help with setting up and analysing the RNAseq data. Protein identification by mass spectrometry was performed by James Ault in the University of Leeds Mass Spectrometry Facility. Also thanks to the University of Leeds Bio-imaging facility for use of the DeltaVision and LSM880 microscopes. I am indebted to the past and present members of the Johnson and Peckham labs for their camaraderie and invaluable support.