Xq28 Duplications
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
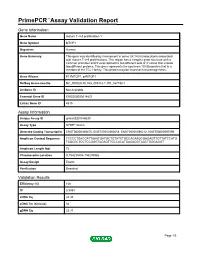
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name mature T-cell proliferation 1 Gene Symbol MTCP1 Organism Human Gene Summary This gene was identified by involvement in some t(X;14) translocations associated with mature T-cell proliferations. This region has a complex gene structure with a common promoter and 5' exon spliced to two different sets of 3' exons that encode two different proteins. This gene represents the upstream 13 kDa protein that is a member of the TCL1 family. This protein may be involved in leukemogenesis. Gene Aliases P13MTCP1, p8MTCP1 RefSeq Accession No. NC_000023.10, NG_005114.1, NT_167198.1 UniGene ID Not Available Ensembl Gene ID ENSG00000214827 Entrez Gene ID 4515 Assay Information Unique Assay ID qHsaCED0048630 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000369476, ENST00000362018, ENST00000596212, ENST00000597096 Amplicon Context Sequence TCCCCTGACCATTAAATGATGCTGTATCTGCCACAAGCGAGAGTTGTTATCCATG TAGCGCTCCTCCGGGTAGAGTTGCCACATGAGAGGTAGCTGGGAGGT Amplicon Length (bp) 72 Chromosome Location X:154293804-154293986 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 106 R2 0.9994 cDNA Cq 24.34 cDNA Tm (Celsius) 82 gDNA Cq 25.37 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report MTCP1, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard -

Structural Studies of the Complex Between Akt-In and the Akt2-PH Domain Suggest That the Peptide Acts As an Allosteric Inhibitor of the Akt Kinase
The Open Spectroscopy Journal, 2009, 3, 65-76 65 Open Access Structural Studies of the Complex Between Akt-in and the Akt2-PH Domain Suggest that the Peptide Acts as an Allosteric Inhibitor of the Akt Kinase Virginie Ropars1,2,3, Philippe Barthe1,2,3, Chi-Shien Wang4, Wenlung Chen4, Der-Lii M. Tzou4,5, Anne Descours6, Loïc Martin6, Masayuki Noguchi7, Daniel Auguin1,2,3,,¶ and Christian Roumestand*,1,2,3 1CNRS UMR 5048, Centre de Biochimie Structurale, Montpellier, France 2INSERM U554, Montpellier, France 3Université Montpellier I et II, Montpellier, France 4Department of Applied Chemistry, National Chiayi University, Chiayi 60004, Taiwan, ROC 5Institute of Chemistry, Academia Sinica, Nankang, Taipei 11529, Taiwan, ROC 6CEA, iBiTecs, Service d’Ingénierie Moléculaire des Protéines, 91191 Gif sur Yvette, France 7Division of Cancer Biology, Institute for Genetic Medicine, Hokkaido University, N15 W7, Kita-ku, Sapporo 060-0815, Japan Abstract: Serine/threonine kinase Akt plays a central role in the regulation of cell survival and proliferation. Hence, the search for Akt specific inhibitors constitutes an attractive strategy for anticancer therapy. We have previously demonstrated that the proto-oncogene TCL1 coactivates Akt upon binding to its Plekstrin Homology Domain, and we proposed a model for the structure of the complex TCL1:Akt2-PHD. This model led to the rational design of Akt-in, a peptide inhibitor spanning the A ß-strand of human TCL1 that binds Akt2 PH domain and inhibits the kinase activation. In the present report, we used NMR spectroscopy to determine the 3D structure of the peptide free in solution and bound to Akt2-PHD. -

A Common X-Linked Inborn Error of Carnitine Biosynthesis May Be a Risk Factor for Nondysmorphic Autism
A common X-linked inborn error of carnitine biosynthesis may be a risk factor for nondysmorphic autism Patrícia B. S. Celestino-Sopera,1, Sara Violanteb,c,1, Emily L. Crawfordd, Rui Luoe, Anath C. Lionelf, Elsa Delabyg, Guiqing Caih, Bekim Sadikovica, Kwanghyuk Leea, Charlene Loa, Kun Gaoe, Richard E. Persona, Timothy J. Mossa, Jennifer R. Germana, Ni Huangi, Marwan Shinawia,j,2, Diane Treadwell-Deeringj,k, Peter Szatmaril, Wendy Robertsm, Bridget Fernandezn, Richard J. Schroero, Roger E. Stevensono, Joseph D. Buxbaumh, Catalina Betancurg, Stephen W. Schererf,m, Stephan J. Sandersp, Daniel H. Geschwinde, James S. Sutcliffed, Matthew E. Hurlesi, Ronald J. A. Wandersb, Chad A. Shawa, Suzanne M. Leala, Edwin H. Cook, Jr.q, Robin P. Goin-Kochela,j,r, Frédéric M. Vazb,1, and Arthur L. Beaudeta,j,r,1,3 Departments of aMolecular and Human Genetics, kPsychiatry, and rPediatrics, Baylor College of Medicine, Houston, TX 77030; jTexas Children’s Hospital, Houston, TX 77030; bLaboratory Genetic Metabolic Disease, Departments of Clinical Chemistry and Pediatrics, Academic Medical Center, University of Amsterdam, 1105 AZ, Amsterdam, The Netherlands; cMetabolism and Genetics Group, Research Institute for Medicines and Pharmaceutical Sciences (iMed.UL), Faculdade de Farmácia, Universidade de Lisboa, 1649-003 Lisbon, Portugal; dDepartment of Molecular Physiology and Biophysics, Center for Molecular Neuroscience, Vanderbilt University, Nashville, TN 37232; eDepartment of Human Genetics, David Geffen School of Medicine, University of California, Los Angeles, -

The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid
The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid Leukemia Cells Supported Only by Exogenous Cytokines Is Associated with a Patient Subset with Adverse Outcome Annette K. Brenner, Elise Aasebø, Maria Hernandez-Valladares, Frode Selheim, Frode Berven, Ida-Sofie Grønningsæter, Sushma Bartaula-Brevik and Øystein Bruserud Supplementary Material S2 of S31 Table S1. Detailed information about the 68 AML patients included in the study. # of blasts Viability Proliferation Cytokine Viable cells Change in ID Gender Age Etiology FAB Cytogenetics Mutations CD34 Colonies (109/L) (%) 48 h (cpm) secretion (106) 5 weeks phenotype 1 M 42 de novo 241 M2 normal Flt3 pos 31.0 3848 low 0.24 7 yes 2 M 82 MF 12.4 M2 t(9;22) wt pos 81.6 74,686 low 1.43 969 yes 3 F 49 CML/relapse 149 M2 complex n.d. pos 26.2 3472 low 0.08 n.d. no 4 M 33 de novo 62.0 M2 normal wt pos 67.5 6206 low 0.08 6.5 no 5 M 71 relapse 91.0 M4 normal NPM1 pos 63.5 21,331 low 0.17 n.d. yes 6 M 83 de novo 109 M1 n.d. wt pos 19.1 8764 low 1.65 693 no 7 F 77 MDS 26.4 M1 normal wt pos 89.4 53,799 high 3.43 2746 no 8 M 46 de novo 26.9 M1 normal NPM1 n.d. n.d. 3472 low 1.56 n.d. no 9 M 68 MF 50.8 M4 normal D835 pos 69.4 1640 low 0.08 n.d. -

Snps) Distant from Xenobiotic Response Elements Can Modulate Aryl Hydrocarbon Receptor Function: SNP-Dependent CYP1A1 Induction S
Supplemental material to this article can be found at: http://dmd.aspetjournals.org/content/suppl/2018/07/06/dmd.118.082164.DC1 1521-009X/46/9/1372–1381$35.00 https://doi.org/10.1124/dmd.118.082164 DRUG METABOLISM AND DISPOSITION Drug Metab Dispos 46:1372–1381, September 2018 Copyright ª 2018 by The American Society for Pharmacology and Experimental Therapeutics Single Nucleotide Polymorphisms (SNPs) Distant from Xenobiotic Response Elements Can Modulate Aryl Hydrocarbon Receptor Function: SNP-Dependent CYP1A1 Induction s Duan Liu, Sisi Qin, Balmiki Ray,1 Krishna R. Kalari, Liewei Wang, and Richard M. Weinshilboum Division of Clinical Pharmacology, Department of Molecular Pharmacology and Experimental Therapeutics (D.L., S.Q., B.R., L.W., R.M.W.) and Division of Biomedical Statistics and Informatics, Department of Health Sciences Research (K.R.K.), Mayo Clinic, Rochester, Minnesota Received April 22, 2018; accepted June 28, 2018 ABSTRACT Downloaded from CYP1A1 expression can be upregulated by the ligand-activated aryl fashion. LCLs with the AA genotype displayed significantly higher hydrocarbon receptor (AHR). Based on prior observations with AHR-XRE binding and CYP1A1 mRNA expression after 3MC estrogen receptors and estrogen response elements, we tested treatment than did those with the GG genotype. Electrophoretic the hypothesis that single-nucleotide polymorphisms (SNPs) map- mobility shift assay (EMSA) showed that oligonucleotides with the ping hundreds of base pairs (bp) from xenobiotic response elements AA genotype displayed higher LCL nuclear extract binding after (XREs) might influence AHR binding and subsequent gene expres- 3MC treatment than did those with the GG genotype, and mass dmd.aspetjournals.org sion. -

Association of Gene Ontology Categories with Decay Rate for Hepg2 Experiments These Tables Show Details for All Gene Ontology Categories
Supplementary Table 1: Association of Gene Ontology Categories with Decay Rate for HepG2 Experiments These tables show details for all Gene Ontology categories. Inferences for manual classification scheme shown at the bottom. Those categories used in Figure 1A are highlighted in bold. Standard Deviations are shown in parentheses. P-values less than 1E-20 are indicated with a "0". Rate r (hour^-1) Half-life < 2hr. Decay % GO Number Category Name Probe Sets Group Non-Group Distribution p-value In-Group Non-Group Representation p-value GO:0006350 transcription 1523 0.221 (0.009) 0.127 (0.002) FASTER 0 13.1 (0.4) 4.5 (0.1) OVER 0 GO:0006351 transcription, DNA-dependent 1498 0.220 (0.009) 0.127 (0.002) FASTER 0 13.0 (0.4) 4.5 (0.1) OVER 0 GO:0006355 regulation of transcription, DNA-dependent 1163 0.230 (0.011) 0.128 (0.002) FASTER 5.00E-21 14.2 (0.5) 4.6 (0.1) OVER 0 GO:0006366 transcription from Pol II promoter 845 0.225 (0.012) 0.130 (0.002) FASTER 1.88E-14 13.0 (0.5) 4.8 (0.1) OVER 0 GO:0006139 nucleobase, nucleoside, nucleotide and nucleic acid metabolism3004 0.173 (0.006) 0.127 (0.002) FASTER 1.28E-12 8.4 (0.2) 4.5 (0.1) OVER 0 GO:0006357 regulation of transcription from Pol II promoter 487 0.231 (0.016) 0.132 (0.002) FASTER 6.05E-10 13.5 (0.6) 4.9 (0.1) OVER 0 GO:0008283 cell proliferation 625 0.189 (0.014) 0.132 (0.002) FASTER 1.95E-05 10.1 (0.6) 5.0 (0.1) OVER 1.50E-20 GO:0006513 monoubiquitination 36 0.305 (0.049) 0.134 (0.002) FASTER 2.69E-04 25.4 (4.4) 5.1 (0.1) OVER 2.04E-06 GO:0007050 cell cycle arrest 57 0.311 (0.054) 0.133 (0.002) -

Regulation of the Akt Kinase by Interacting Proteins
Oncogene (2005) 24, 7401–7409 & 2005 Nature Publishing Group All rights reserved 0950-9232/05 $30.00 www.nature.com/onc Regulation of the Akt kinase by interacting proteins Keyong Du1 and Philip N Tsichlis*,1 1Molecular Oncology Research Institute, Tufts-New England Medical Center, Boston, MA 02111, USA Ten years ago, it was observed that the Akt kinase revolutionized our understanding of cell function at is activated by phosphorylation via a phosphoinositide the molecular level.These technologies also allowed 3-kinase (PI-3K)-dependent process. This discovery gene- the identification of a large array of protein–protein rated enormous interest because it provided a link between interaction motifs that facilitate our understanding of PI-3K, an enzyme known to play a critical role in cellular signal transduction by providing clues on the potential physiology, and its downstream targets. Subsequently, it function of novel signaling proteins (Barnouin, 2004; was shown that the activity of the core components of the Miller and Stagljar, 2004). ‘PI-3K/Akt pathway’ is modulated by a complex network Akt1, also known as protein kinase Ba (PKBa) of regulatory proteins and pathways. Some of the Akt- (Bellacosa et al., 1991; Coffer and Woodgett, 1991; binding partners modulate its activation by external Jones et al., 1991), is the founding member of a protein signals by interacting with different domains of the Akt kinase family composed of three members, Akt1, Akt2, protein. This review focuses on the Akt interacting and Akt3.Akt family members regulate a diverse proteins and the mechanisms by which they regulate Akt array of cellular functions, including apoptosis, cellular activation. -

430.Full.Pdf
430 LETTER TO JMG J Med Genet: first published as 10.1136/jmg.39.6.430 on 1 June 2002. Downloaded from A novel 2 bp deletion in the TM4SF2 gene is associated with MRX58 F E Abidi, E Holinski-Feder, O Rittinger, F Kooy, H A Lubs, R E Stevenson, C E Schwartz ............................................................................................................................. J Med Genet 2002;39:430–433 linked mental retardation (XLMR) represents around 5% detection of any alteration at the splice junction. Since the of all MR, with a prevalence of 1 in 600 males.12Fifteen PCR products for exons 2 and 5 were greater than 310 bp, they to twenty percent of the total XLMR is the result of the were digested with HaeIII and PvuII respectively (table 1). X 3 fragile X syndrome. Non-fragile X mental retardation was Amplification was done in a PTC-200 thermocycler (MJ subdivided into syndromal and non-syndromal conditions by Research). Following PCR, 10 µl of the IPS loading dye (95% Neri et al4 in 1991. The syndromal XLMR entities (MRXS) are formamide, 10 mmol/l NaOH, 0.25% bromophenol blue, 0.25% those in which there is a specific pattern of physical, xylene cyanol) was added. The samples were denatured at neurological, or metabolic abnormalities associated with the 96°C for three minutes and loaded on a 0.5 × MDE gel (FMC) presence of mental retardation.5 Non-syndromic XLMR prepared in 0.6 × TBE. The gel was run at 8 W for 15-20 hours (MRX) are conditions in which a gene mutation causes men- at room temperature. -

Gene Ontology Functional Annotations and Pleiotropy
Network based analysis of genetic disease associations Sarah Gilman Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy under the Executive Committee of the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2014 © 2013 Sarah Gilman All Rights Reserved ABSTRACT Network based analysis of genetic disease associations Sarah Gilman Despite extensive efforts and many promising early findings, genome-wide association studies have explained only a small fraction of the genetic factors contributing to common human diseases. There are many theories about where this “missing heritability” might lie, but increasingly the prevailing view is that common variants, the target of GWAS, are not solely responsible for susceptibility to common diseases and a substantial portion of human disease risk will be found among rare variants. Relatively new, such variants have not been subject to purifying selection, and therefore may be particularly pertinent for neuropsychiatric disorders and other diseases with greatly reduced fecundity. Recently, several researchers have made great progress towards uncovering the genetics behind autism and schizophrenia. By sequencing families, they have found hundreds of de novo variants occurring only in affected individuals, both large structural copy number variants and single nucleotide variants. Despite studying large cohorts there has been little recurrence among the genes implicated suggesting that many hundreds of genes may underlie these complex phenotypes. The question -

Anti-GDI1 (Aa 399-447) Polyclonal Antibody (DPABH-15243) This Product Is for Research Use Only and Is Not Intended for Diagnostic Use
Anti-GDI1 (aa 399-447) polyclonal antibody (DPABH-15243) This product is for research use only and is not intended for diagnostic use. PRODUCT INFORMATION Antigen Description Regulates the GDP/GTP exchange reaction of most Rab proteins by inhibiting the dissociation of GDP from them, and the subsequent binding of GTP to them. Immunogen A region within a synthetic peptide: FCSCSYDATT HFETTCNDIK DIYKRMAGTA FDFENMKRKQ NDVFGEAEQ, corresponding to C terminal amino acids 399-447 of Human GDI1 Isotype IgG Source/Host Rabbit Species Reactivity Human Purification IgG fraction Conjugate Unconjugated Applications WB Format Liquid Size 100 μg Buffer Constituents: 2% Sucrose, PBS Preservative None Storage Shipped at 4°C. Upon delivery aliquot and store at -20°C or -80°C. Avoid repeated freeze / thaw cycles. GENE INFORMATION Gene Name GDI1 GDP dissociation inhibitor 1 [ Homo sapiens ] Official Symbol GDI1 Synonyms GDI1; GDP dissociation inhibitor 1; GDIL, MRX41, MRX48; rab GDP dissociation inhibitor alpha; FLJ41411; mental retardation; X linked 41; X linked 48; OPHN2; rab GDP dissociation inhibitor; 45-1 Ramsey Road, Shirley, NY 11967, USA Email: [email protected] Tel: 1-631-624-4882 Fax: 1-631-938-8221 1 © Creative Diagnostics All Rights Reserved alpha; RABGDIA; XAP 4; GDI-1; protein XAP-4; rab GDI alpha; oligophrenin-2; mental retardation, X-linked 48; rab GDP-dissociation inhibitor, alpha; guanosine diphosphate dissociation inhibitor 1; 1A; GDIL; MRX41; MRX48; XAP-4; RABGD1A; Entrez Gene ID 2664 Protein Refseq NP_001484 UniProt ID P31150 Chromosome Location Xq28 Pathway Rho GTPase cycle; Signal Transduction; Signaling by Rho GTPases; Signaling mediated by p38-alpha and p38-beta; Function GDP-dissociation inhibitor activity; GTPase activator activity; Rab GDP-dissociation inhibitor activity; protein binding; 45-1 Ramsey Road, Shirley, NY 11967, USA Email: [email protected] Tel: 1-631-624-4882 Fax: 1-631-938-8221 2 © Creative Diagnostics All Rights Reserved. -

Downloaded from the App Store and Nucleobase, Nucleotide and Nucleic Acid Metabolism 7 Google Play
Hoytema van Konijnenburg et al. Orphanet J Rare Dis (2021) 16:170 https://doi.org/10.1186/s13023-021-01727-2 REVIEW Open Access Treatable inherited metabolic disorders causing intellectual disability: 2021 review and digital app Eva M. M. Hoytema van Konijnenburg1†, Saskia B. Wortmann2,3,4†, Marina J. Koelewijn2, Laura A. Tseng1,4, Roderick Houben6, Sylvia Stöckler‑Ipsiroglu5, Carlos R. Ferreira7 and Clara D. M. van Karnebeek1,2,4,8* Abstract Background: The Treatable ID App was created in 2012 as digital tool to improve early recognition and intervention for treatable inherited metabolic disorders (IMDs) presenting with global developmental delay and intellectual disabil‑ ity (collectively ‘treatable IDs’). Our aim is to update the 2012 review on treatable IDs and App to capture the advances made in the identifcation of new IMDs along with increased pathophysiological insights catalyzing therapeutic development and implementation. Methods: Two independent reviewers queried PubMed, OMIM and Orphanet databases to reassess all previously included disorders and therapies and to identify all reports on Treatable IDs published between 2012 and 2021. These were included if listed in the International Classifcation of IMDs (ICIMD) and presenting with ID as a major feature, and if published evidence for a therapeutic intervention improving ID primary and/or secondary outcomes is avail‑ able. Data on clinical symptoms, diagnostic testing, treatment strategies, efects on outcomes, and evidence levels were extracted and evaluated by the reviewers and external experts. The generated knowledge was translated into a diagnostic algorithm and updated version of the App with novel features. Results: Our review identifed 116 treatable IDs (139 genes), of which 44 newly identifed, belonging to 17 ICIMD categories. -

Epigenetic Co-Activation of Genes MAGEA6 and CT-GABRA3 Defines Orientation of a Segmental Duplication on the Human X Chromosome
Cytogenetic and Genome Research (Karger) October 2019 (doi : 10.1159/000502933) Epigenetic co-activation of genes MAGEA6 and CT-GABRA3 defines orientation of a segmental duplication on the human X chromosome Jean S. Faina, Aurélie Van Tongelena, Axelle Loriota and Charles De Smeta,b a Group of Genetics and Epigenetics, de Duve Institute, Université catholique de Louvain (UCLouvain), Brussels, Belgium b Corresponding author: [email protected] Keywords: Cancer-germline genes, MAGEA3, MAGEA6, segmental duplication, bidirectional promoter, genome misassembly Abstract The human genome harbors many duplicated segments, which sometimes show very high sequence identity. This may complicate assignment during genome assembly. One such example is on Xq28, where the arrangement of two recently duplicated segments varies between genome assembly versions. The duplicated segments comprise highly similar genes, including MAGEA3 and MAGEA6, which display specific expression in testicular germline cells, and also become aberrantly activated in a variety of tumors. Recently, a new gene was identified, CT-GABRA3, the transcription of which initiates inside the segmental duplication but extends far outside. According to the latest genome annotation, CT-GABRA3 starts near MAGEA3, with which it shares a bidirectional promoter. In an earlier annotation however, the duplicated segment was positioned in the opposite orientation, and CT-GABRA3 was instead coupled with MAGEA6. To resolve this discrepancy, and based on the contention that genes connected by a bidirectional promoter are almost always co-expressed, we decided to compare the expression profiles of CT-GABRA3, MAGEA3, and MAGEA6. We found that in tumor tissues and cell lines of different origins, the expression of CT- GABRA3 was better correlated with that of MAGEA6.