A Cellular Protein, Activating Transcription Factor, Activates
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplemental Materials ZNF281 Enhances Cardiac Reprogramming
Supplemental Materials ZNF281 enhances cardiac reprogramming by modulating cardiac and inflammatory gene expression Huanyu Zhou, Maria Gabriela Morales, Hisayuki Hashimoto, Matthew E. Dickson, Kunhua Song, Wenduo Ye, Min S. Kim, Hanspeter Niederstrasser, Zhaoning Wang, Beibei Chen, Bruce A. Posner, Rhonda Bassel-Duby and Eric N. Olson Supplemental Table 1; related to Figure 1. Supplemental Table 2; related to Figure 1. Supplemental Table 3; related to the “quantitative mRNA measurement” in Materials and Methods section. Supplemental Table 4; related to the “ChIP-seq, gene ontology and pathway analysis” and “RNA-seq” and gene ontology analysis” in Materials and Methods section. Supplemental Figure S1; related to Figure 1. Supplemental Figure S2; related to Figure 2. Supplemental Figure S3; related to Figure 3. Supplemental Figure S4; related to Figure 4. Supplemental Figure S5; related to Figure 6. Supplemental Table S1. Genes included in human retroviral ORF cDNA library. Gene Gene Gene Gene Gene Gene Gene Gene Symbol Symbol Symbol Symbol Symbol Symbol Symbol Symbol AATF BMP8A CEBPE CTNNB1 ESR2 GDF3 HOXA5 IL17D ADIPOQ BRPF1 CEBPG CUX1 ESRRA GDF6 HOXA6 IL17F ADNP BRPF3 CERS1 CX3CL1 ETS1 GIN1 HOXA7 IL18 AEBP1 BUD31 CERS2 CXCL10 ETS2 GLIS3 HOXB1 IL19 AFF4 C17ORF77 CERS4 CXCL11 ETV3 GMEB1 HOXB13 IL1A AHR C1QTNF4 CFL2 CXCL12 ETV7 GPBP1 HOXB5 IL1B AIMP1 C21ORF66 CHIA CXCL13 FAM3B GPER HOXB6 IL1F3 ALS2CR8 CBFA2T2 CIR1 CXCL14 FAM3D GPI HOXB7 IL1F5 ALX1 CBFA2T3 CITED1 CXCL16 FASLG GREM1 HOXB9 IL1F6 ARGFX CBFB CITED2 CXCL3 FBLN1 GREM2 HOXC4 IL1F7 -
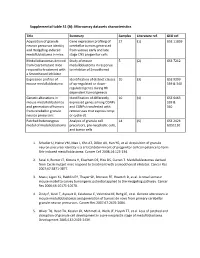
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Transcriptional and Post-Transcriptional Regulation of ATP-Binding Cassette Transporter Expression
Transcriptional and Post-transcriptional Regulation of ATP-binding Cassette Transporter Expression by Aparna Chhibber DISSERTATION Submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILOSOPHY in Pharmaceutical Sciences and Pbarmacogenomies in the Copyright 2014 by Aparna Chhibber ii Acknowledgements First and foremost, I would like to thank my advisor, Dr. Deanna Kroetz. More than just a research advisor, Deanna has clearly made it a priority to guide her students to become better scientists, and I am grateful for the countless hours she has spent editing papers, developing presentations, discussing research, and so much more. I would not have made it this far without her support and guidance. My thesis committee has provided valuable advice through the years. Dr. Nadav Ahituv in particular has been a source of support from my first year in the graduate program as my academic advisor, qualifying exam committee chair, and finally thesis committee member. Dr. Kathy Giacomini graciously stepped in as a member of my thesis committee in my 3rd year, and Dr. Steven Brenner provided valuable input as thesis committee member in my 2nd year. My labmates over the past five years have been incredible colleagues and friends. Dr. Svetlana Markova first welcomed me into the lab and taught me numerous laboratory techniques, and has always been willing to act as a sounding board. Michael Martin has been my partner-in-crime in the lab from the beginning, and has made my days in lab fly by. Dr. Yingmei Lui has made the lab run smoothly, and has always been willing to jump in to help me at a moment’s notice. -
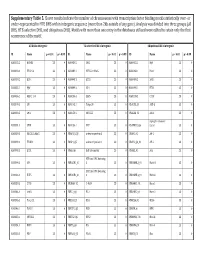
Supplementary Table 5. Clover Results Indicate the Number Of
Supplementary Table 5. Clover results indicate the number of chromosomes with transcription factor binding motifs statistically over‐ or under‐represented in HTE DHS within intergenic sequence (more than 2kb outside of any gene). Analysis was divided into three groups (all DHS, HTE‐selective DHS, and ubiquitous DHS). Motifs with more than one entry in the databases utilized were edited to retain only the first occurrence of the motif. All DHS x Intergenic TEselective DHS x Intergenic Ubiquitous DHS x Intergenic ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 MA0002.2 RUNX1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0003.1 TFAP2A 23 0 MA0089.1 NFE2L1::MafG 23 0 MA0068.1 Pax4 23 0 MA0039.2 Klf4 23 0 MA0098.1 ETS1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0099.2 AP1 23 0 MA0098.1 ETS1 23 0 MA0056.1 MZF1_1‐4 23 0 MA0136.1 ELF5 23 0 MA0139.1 CTCF 23 0 MA0079.2 SP1 23 0 MA0145.1 Tcfcp2l1 23 0 V$ALX3_01 ALX‐3 23 0 MA0080.2 SPI1 23 0 MA0150.1 NFE2L2 23 0 V$ALX4_02 Alx‐4 23 0 myocyte enhancer MA0081.1 SPIB 23 0 MA0156.1 FEV 23 0 V$AMEF2_Q6 factor 23 0 MA0089.1 NFE2L1::MafG 23 0 V$AP1FJ_Q2 activator protein 1 23 0 V$AP1_01 AP‐1 23 0 MA0090.1 TEAD1 23 0 V$AP4_Q5 activator protein 4 23 0 V$AP2_Q6_01 AP‐2 23 0 MA0098.1 ETS1 23 0 V$AR_Q6 half‐site matrix 23 0 V$ARX_01 Arx 23 0 BTB and CNC homolog MA0099.2 AP1 23 0 V$BACH1_01 1 23 0 V$BARHL1_01 Barhl‐1 23 0 BTB and CNC homolog MA0136.1 ELF5 23 0 V$BACH2_01 2 23 0 V$BARHL2_01 Barhl2 23 0 MA0139.1 CTCF 23 0 V$CMAF_02 C‐MAF 23 0 V$BARX1_01 Barx1 23 0 MA0144.1 Stat3 23 0 -

Cardiac SARS‐Cov‐2 Infection Is Associated with Distinct Tran‐ Scriptomic Changes Within the Heart
Cardiac SARS‐CoV‐2 infection is associated with distinct tran‐ scriptomic changes within the heart Diana Lindner, PhD*1,2, Hanna Bräuninger, MS*1,2, Bastian Stoffers, MS1,2, Antonia Fitzek, MD3, Kira Meißner3, Ganna Aleshcheva, PhD4, Michaela Schweizer, PhD5, Jessica Weimann, MS1, Björn Rotter, PhD9, Svenja Warnke, BSc1, Carolin Edler, MD3, Fabian Braun, MD8, Kevin Roedl, MD10, Katharina Scher‐ schel, PhD1,12,13, Felicitas Escher, MD4,6,7, Stefan Kluge, MD10, Tobias B. Huber, MD8, Benjamin Ondruschka, MD3, Heinz‐Peter‐Schultheiss, MD4, Paulus Kirchhof, MD1,2,11, Stefan Blankenberg, MD1,2, Klaus Püschel, MD3, Dirk Westermann, MD1,2 1 Department of Cardiology, University Heart and Vascular Center Hamburg, Germany. 2 DZHK (German Center for Cardiovascular Research), partner site Hamburg/Kiel/Lübeck. 3 Institute of Legal Medicine, University Medical Center Hamburg‐Eppendorf, Germany. 4 Institute for Cardiac Diagnostics and Therapy, Berlin, Germany. 5 Department of Electron Microscopy, Center for Molecular Neurobiology, University Medical Center Hamburg‐Eppendorf, Germany. 6 Department of Cardiology, Charité‐Universitaetsmedizin, Berlin, Germany. 7 DZHK (German Centre for Cardiovascular Research), partner site Berlin, Germany. 8 III. Department of Medicine, University Medical Center Hamburg‐Eppendorf, Germany. 9 GenXPro GmbH, Frankfurter Innovationszentrum, Biotechnologie (FIZ), Frankfurt am Main, Germany. 10 Department of Intensive Care Medicine, University Medical Center Hamburg‐Eppendorf, Germany. 11 Institute of Cardiovascular Sciences, -

Reorganization of the Host Epigenome by a Viral Oncogene
Downloaded from genome.cshlp.org on September 29, 2021 - Published by Cold Spring Harbor Laboratory Press Research Reorganization of the host epigenome by a viral oncogene Roberto Ferrari,1,2 Trent Su,1,3 Bing Li,1 Giancarlo Bonora,1,4 Amit Oberai,1 Yvonne Chan,1 Rajkumar Sasidharan,5 Arnold J. Berk,6,7 Matteo Pellegrini,2,5 and Siavash K. Kurdistani1,2,7,8,9 1Department of Biological Chemistry, David Geffen School of Medicine, University of California, Los Angeles, California 90095, USA; 2Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research, University of California, Los Angeles, California 90095, USA; 3Division of Oral Biology and Medicine, School of Dentistry, David Geffen School of Medicine, University of California, Los Angeles, California 90095, USA; 4UCLA Bioinformatics Interdepartmental Degree Program, David Geffen School of Medicine, University of California, Los Angeles, California 90095, USA; 5Department of Molecular, Cellular, and Developmental Biology, University of California, Los Angeles, California 90095, USA; 6Department of Microbiology, Immunology and Molecular Genetics, University of California, Los Angeles, California 90095, USA; 7Molecular Biology Institute, University of California, Los Angeles, California 90095, USA; 8Department of Pathology and Laboratory of Medicine, David Geffen School of Medicine, University of California, Los Angeles, California 90095, USA Adenovirus small e1a oncoprotein causes ~70% reduction in cellular levels of histone H3 lysine 18 acetylation (H3K18ac). It is unclear, however, where this dramatic reduction occurs genome-wide. ChIP-sequencing revealed that by 24 h after expression, e1a erases 95% of H3K18ac peaks in normal, contact-inhibited fibroblasts and replaces them with one-third as many at new genomic locations. -

The Proteasomal Deubiquitinating Enzyme PSMD14 Regulates Macroautophagy by Controlling Golgi-To-ER Retrograde Transport
Supplementary Materials The proteasomal deubiquitinating enzyme PSMD14 regulates macroautophagy by controlling Golgi-to-ER retrograde transport Bustamante HA., et al. Figure S1. siRNA sequences directed against human PSMD14 used for Validation Stage. Figure S2. Primer pairs sequences used for RT-qPCR. Figure S3. The PSMD14 DUB inhibitor CZM increases the Golgi apparatus area. Immunofluorescence microscopy analysis of the Golgi area in parental H4 cells treated for 4 h either with the vehicle (DMSO; Control) or CZM. The Golgi marker GM130 was used to determine the region of interest in each condition. Statistical significance was determined by Student's t-test. Bars represent the mean ± SEM (n =43 cells). ***P <0.001. Figure S4. CZM causes the accumulation of KDELR1-GFP at the Golgi apparatus. HeLa cells expressing KDELR1-GFP were either left untreated or treated with CZM for 30, 60 or 90 min. Cells were fixed and representative confocal images were acquired. Figure S5. Effect of CZM on proteasome activity. Parental H4 cells were treated either with the vehicle (DMSO; Control), CZM or MG132, for 90 min. Protein extracts were used to measure in vitro the Chymotrypsin-like peptidase activity of the proteasome. The enzymatic activity was quantified according to the cleavage of the fluorogenic substrate Suc-LLVY-AMC to AMC, and normalized to that of control cells. The statistical significance was determined by One-Way ANOVA, followed by Tukey’s test. Bars represent the mean ± SD of biological replicates (n=3). **P <0.01; n.s., not significant. Figure S6. Effect of CZM and MG132 on basal macroautophagy. (A) Immunofluorescence microscopy analysis of the subcellular localization of LC3 in parental H4 cells treated with either with the vehicle (DMSO; Control), CZM for 4 h or MG132 for 6 h. -

Transcription Factor and Lncrna Regulatory Networks Identify Key Elements in Lung Adenocarcinoma
G C A T T A C G G C A T genes Article Transcription Factor and lncRNA Regulatory Networks Identify Key Elements in Lung Adenocarcinoma Dan Li 1, William Yang 2, Jialing Zhang 3, Jack Y. Yang 1, Renchu Guan 1 and Mary Qu Yang 1,* 1 Joint Bioinformatics Graduate Program, Department of Information Science, George W. Donaghey College of Engineering and Information Technology, University of Arkansas at Little Rock and University of Arkansas for Medical Sciences, 2801 S. University Ave, Little Rock, AR 72204, USA; [email protected] (D.L.); [email protected] (J.Y.Y.); [email protected] (R.G.) 2 School of Computer Science, Carnegie Mellon University, 5000 Forbes Ave, Pittsburgh, PA 15213, USA; [email protected] 3 Department of Genetics, Yale University, New Haven, CT 06520, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-501-683-2035 Received: 19 September 2017; Accepted: 21 December 2017; Published: 5 January 2018 Abstract: Lung cancer is the second most commonly diagnosed carcinoma and is the leading cause of cancer death. Although significant progress has been made towards its understanding and treatment, unraveling the complexities of lung cancer is still hampered by a lack of comprehensive knowledge on the mechanisms underlying the disease. High-throughput and multidimensional genomic data have shed new light on cancer biology. In this study, we developed a network-based approach integrating somatic mutations, the transcriptome, DNA methylation, and protein-DNA interactions to reveal the key regulators in lung adenocarcinoma (LUAD). By combining Bayesian network analysis with tissue-specific transcription factor (TF) and targeted gene interactions, we inferred 15 disease-related core regulatory networks in co-expression gene modules associated with LUAD. -

Table S1A. Patients and Tumor Characteristics of the Pdxs
Table S1A. Patients and tumor characteristics of the PDXs TUMOR ID Passage Histology Gender Age Smoking Pack/Year Grade Stage TNM PDX1 2 LSCC Male 63 CURRENT 45 G4 (POOR) IIB pT3pN0 PDX2 2 LCNEC Male 71 FORMER 50 G4 (POOR) IIIA pT3pN1 PDX3 2 LSCC Male 76 FORMER 45 G2 (MOD.) IB pT2apN0 PDX4 2 LUAD Male 54 FORMER 85 G4 (POOR) IIB pT3pN0 PDX5 2 LSCC Male 74 CURRENT 50 G3 (POOR) IB pT2apN0 PDX6 2 LSCC Male 70 FORMER 40 G2 (MOD.) IIIA pT4pN1 PDX7 2 LUAD (ACINAR+SOLID) Male 52 CURRENT 37 na IIIA pT3N2 PDX8 2 LSCC Male 74 FORMER na na IIB pT2pN1 PDX9 48 LUAD Female 73 FORMER 20 G3 (POOR) IIIA T1aN2M0 PDX10 17 LUAD Male 74 CURRENT 120 G3 (POOR) IIIA T2bN2M0 PDX11 3 LUAD Male 67 FORMER 30 G3 (POOR) IA T1aN0M0 PDX12 3 LUAD Male 68 CURRENT 50 na IIIA T3N2M0 PDX13 1 LUAD (SOLID+MUCIN) Male 61 FORMER 23 G2 (MOD.) na pT2aNxMx PDX14 1 LSCC Male 61 CURRENT 45 G2 (MOD.) IA pT1aN0Mx LCNEC, lung neuroendocrine carcinoma LUAD, lung adenocarcinoma; LSCC, lung squamous cell carcinoma, na, not available Table 1SB. Exome sequencing depth and coverage report Sample Sample Million Yield % uniquely % Mean Coverage Coverage % of mouse Reference genome barcode name read-pairs (Gb) mapping unmapped coverage at Depth at Depth reads AB0115 N1 27,394 5,533 hsapiens.hs37d5 91,98 0,14 60,17 92,10 87,50 0,66 AB0117 PDX1 24,480 4,945 human.37 92,16 0,48 51,57 90,00 84,00 618N N2 23,458 6,01 hsapiens.hs37d5 92,44 0,16 50,00 88,77 75,35 0,87 AB0106 PDX2 20,858 4,213 human.37 92,64 0,29 48,52 90,60 54,60 AB0107 N3 27,219 5,498 hsapiens.hs37d5 92,81 0,16 60,35 94,90 91,30 1,01 -

SUPPLEMENTARY MATERIAL Pitx3 Directly Regulates Foxe3 During Early Lens Development
www.intjdevbiol.com doi: 10.1387/ijdb.130193jg SUPPLEMENTARY MATERIAL corresponding to: Pitx3 directly regulates Foxe3 during early lens development NAFEES AHMAD, MUHAMMAD ASLAM, DORIS MUENSTER, MARION HORSCH, MUHAMMAD A. KHAN, PETER CARLSSON, JOHANNES BECKERS and JOCHEN GRAW *Address correspondence to: Jochen Graw. Helmholtz Centre Munich – German Research Center for Environmental Health, Institute of Developmental Genetics, Ingolstädter Landstr. 1, D-85764 Neuherberg, Germany. Tel: +49-89/3187-2610. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.130193jg TABLE S1 PROBES FOR EMSA Gene Probe Sequence* Foxe3 Fox3-1-EMSA 5’-Biotin-AATCCCTGGCCATTAATCCCTCCTGCCAGCCC-3’ Fox3-2-EMSA 5’-Biotin-ACGCTGAAAACGCGGATTAGCCCTTGGGCCGC-3’ Prox1 Prox1-EMSA 5’-Biotin-AGGGGGGGCAGTTTAATCCTGTTAAATGTGGT-3’ Tube1 Tube1-3-1-EMSA 5’-Biotin-GACAAGCTGCTAATAAGCTGTTTCTGCCATCT-3’ Tube1-3-2-EMSA 5’-Biotin-TGTAATAACAAACTAAGCTGTATCCTGGCGGC-3’ *Pitx3 putative binding sites are underlined. TABLE S2 PRIMERS FOR GENOTYPING OF APHAKIA MICE Product size (bp) Primer Sequence Annealing (oC) wt ak Pitx3-1/2NF 5’-ATTCGGTGCGGAGAGTAAGG-3’ 63 1,165 399 Pitx3-2R 5’-ATTGGATTTGGCTCTGATGGTT-3’ TABLE S3 PRIMERS FOR RT-QPCR Annealing Product Gene Primer Sequence (oC) size (bp) E4f1 E4FqF 5’-AGTACATTATTGAGGCCACTGC-3’ 60 219 E4FqR 5’-CAATGGTGATCGTGTCTGC-3’ Foxe3 Foxe3-lt 5’-GCCGCCCTACTCATACATC-3’ 60 172 Foxe3-rt 5’-ACAGTCGTTGAGGGTGAGG-3’ Prox1 Prox1qF 5’-ATGCTGTGTCTCCTGTTTCTCT-3’ 60 101 Prox1qR 5’-GCTTATCAGGCTCAAATCAAAC-3’ Tuba* -

(12) Patent Application Publication (10) Pub. No.: US 2009/0269772 A1 Califano Et Al
US 20090269772A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2009/0269772 A1 Califano et al. (43) Pub. Date: Oct. 29, 2009 (54) SYSTEMS AND METHODS FOR Publication Classification IDENTIFYING COMBINATIONS OF (51) Int. Cl. COMPOUNDS OF THERAPEUTIC INTEREST CI2O I/68 (2006.01) CI2O 1/02 (2006.01) (76) Inventors: Andrea Califano, New York, NY G06N 5/02 (2006.01) (US); Riccardo Dalla-Favera, New (52) U.S. Cl. ........... 435/6: 435/29: 706/54; 707/E17.014 York, NY (US); Owen A. (57) ABSTRACT O'Connor, New York, NY (US) Systems, methods, and apparatus for searching for a combi nation of compounds of therapeutic interest are provided. Correspondence Address: Cell-based assays are performed, each cell-based assay JONES DAY exposing a different sample of cells to a different compound 222 EAST 41ST ST in a plurality of compounds. From the cell-based assays, a NEW YORK, NY 10017 (US) Subset of the tested compounds is selected. For each respec tive compound in the Subset, a molecular abundance profile from cells exposed to the respective compound is measured. (21) Appl. No.: 12/432,579 Targets of transcription factors and post-translational modu lators of transcription factor activity are inferred from the (22) Filed: Apr. 29, 2009 molecular abundance profile data using information theoretic measures. This data is used to construct an interaction net Related U.S. Application Data work. Variances in edges in the interaction network are used to determine the drug activity profile of compounds in the (60) Provisional application No. 61/048.875, filed on Apr.