Mouse Chromosome 10
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Construction of Stable Mouse Arti Cial Chromosome from Native Mouse
Construction of Stable Mouse Articial Chromosome from Native Mouse Chromosome 10 for Generation of Transchromosomic Mice Satoshi Abe Tottori University Kazuhisa Honma Trans Chromosomics, Inc Akane Okada Tottori University Kanako Kazuki Tottori University Hiroshi Tanaka Trans Chromosomics, Inc Takeshi Endo Trans Chromosomics, Inc Kayoko Morimoto Trans Chromosomics, Inc Takashi Moriwaki Tottori University Shusei Hamamichi Tottori University Yuji Nakayama Tottori University Teruhiko Suzuki Tokyo Metropolitan Institute of Medical Science Shoko Takehara Trans Chromosomics, Inc Mitsuo Oshimura Tottori University Yasuhiro Kazuki ( [email protected] ) Tottori University Research Article Page 1/21 Keywords: mouse articial chromosome (MAC), microcell-mediated chromosome transfer (MMCT), chromosome engineering, transchromosomic (Tc) mouse, humanized model mouse Posted Date: July 9th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-675300/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 2/21 Abstract Mammalian articial chromosomes derived from native chromosomes have been applied to biomedical research and development by generating cell sources and transchromosomic (Tc) animals. Human articial chromosome (HAC) is a precedent chromosomal vector which achieved generation of valuable humanized animal models for fully human antibody production and human pharmacokinetics. While humanized Tc animals created by HAC vector have attained signicant contributions, there was a potential issue to be addressed regarding stability in mouse tissues, especially highly proliferating hematopoietic cells. Mouse articial chromosome (MAC) vectors derived from native mouse chromosome 11 demonstrated improved stability, and they were utilized for humanized Tc mouse production as a standard vector. In mouse, however, stability of MAC vector derived from native mouse chromosome other than mouse chromosome 11 remains to be evaluated. -

134 Mb (Almost the Same As the Size of Chromosome 10). It Is ~4–4.5% of the Total Human Genome
Chromosome 11 ©Chromosome Disorder Outreach Inc. (CDO) Technical genetic content provided by Dr. Iosif Lurie, M.D. Ph.D Medical Geneticist and CDO Medical Consultant/Advisor. Ideogram courtesy of the University of Washington Department of Pathology: ©1994 David Adler.hum_11.gif Introduction The genetic size of chromosome 11 is ~134 Mb (almost the same as the size of chromosome 10). It is ~4–4.5% of the total human genome. The length of its short arm is ~50 Mb; the length of its long arm in ~84 Mb. Chromosome 11 is a very gene–rich area. It contains ~1,500 genes. Mutations of ~200 of these genes are known to cause birth defects or some functional abnormalities. The short arm of chromosome 11 contains a region which is known to be imprinted. As a result duplications of this region will have different manifestations depending on the sex of the parent responsible for this defect. Phenotypes of persons with duplications of the maternal origin will be different from the phenotypes of the persons with a paternal duplication of the same area. There are ~1,400 patients with different structural abnormalities of chromosome 11 as the only abnormality or in association with abnormalities for other chromosomes. At least 800 of these patients had different deletions of chromosome 11. Deletions of the short arm have been reported in ~250 patients (including those with an additional imbalance); deletions of the long arm have been described in ~550 patients. There are two syndromes caused by deletions of the short arm (both of these syndromes have been known for several years) and one well–known syndrome caused by distal deletions of the long arm (Jacobsen syndrome). -

WNT16 Is a New Marker of Senescence
Table S1. A. Complete list of 177 genes overexpressed in replicative senescence Value Gene Description UniGene RefSeq 2.440 WNT16 wingless-type MMTV integration site family, member 16 (WNT16), transcript variant 2, mRNA. Hs.272375 NM_016087 2.355 MMP10 matrix metallopeptidase 10 (stromelysin 2) (MMP10), mRNA. Hs.2258 NM_002425 2.344 MMP3 matrix metallopeptidase 3 (stromelysin 1, progelatinase) (MMP3), mRNA. Hs.375129 NM_002422 2.300 HIST1H2AC Histone cluster 1, H2ac Hs.484950 2.134 CLDN1 claudin 1 (CLDN1), mRNA. Hs.439060 NM_021101 2.119 TSPAN13 tetraspanin 13 (TSPAN13), mRNA. Hs.364544 NM_014399 2.112 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 2.070 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 2.026 DCBLD2 discoidin, CUB and LCCL domain containing 2 (DCBLD2), mRNA. Hs.203691 NM_080927 2.007 SERPINB2 serpin peptidase inhibitor, clade B (ovalbumin), member 2 (SERPINB2), mRNA. Hs.594481 NM_002575 2.004 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.989 OBFC2A Oligonucleotide/oligosaccharide-binding fold containing 2A Hs.591610 1.962 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.947 PLCB4 phospholipase C, beta 4 (PLCB4), transcript variant 2, mRNA. Hs.472101 NM_182797 1.934 PLCB4 phospholipase C, beta 4 (PLCB4), transcript variant 1, mRNA. Hs.472101 NM_000933 1.933 KRTAP1-5 keratin associated protein 1-5 (KRTAP1-5), mRNA. Hs.534499 NM_031957 1.894 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.884 CYTL1 cytokine-like 1 (CYTL1), mRNA. Hs.13872 NM_018659 tumor necrosis factor receptor superfamily, member 10d, decoy with truncated death domain (TNFRSF10D), 1.848 TNFRSF10D Hs.213467 NM_003840 mRNA. -

Gene Mapping of Familial Autosomal Dominant Dilated Cardiomyopathy to Chromosome 10Q21-23
Gene mapping of familial autosomal dominant dilated cardiomyopathy to chromosome 10q21-23. K R Bowles, … , R Pignatelli, J A Towbin J Clin Invest. 1996;98(6):1355-1360. https://doi.org/10.1172/JCI118922. Research Article Dilated cardiomyopathy (DCM) is the most common form of primary myocardial disorder, accounting for 60% of all cardiomyopathies. In 20-30% of cases, familial inheritance can be demonstrated; an autosomal dominant transmission is the usual type of inheritance pattern identified. Previously, genetic heterogeneity was demonstrated in familial autosomal dominant dilated cardiomyopathy (FDCM). Gene localization to chromosome 1 (1p1-1q1 and 1q32), chromosome 3 (3p25-3p22), and chromosome 9 (9q13-9q22) has recently been identified. We report one family with 26 members (12 affected) with familial autosomal dominant dilated cardiomyopathy in which linkage to chromosome 10 at the 10q21-q23 locus is identified. Using short tandem repeat polymorphism (STR) markers with heterozygosity > 70%, 169 markers (50% of the genome) were used before linkage was found to markers D10S605 and D10S201 with a pairwise LOD score = 3.91, theta = 0, penetrance = 100% for both markers. Linkage to 1p1-1q1, 1q32, 3p25-3p22, and 9q13-9q22 was excluded. We conclude that a new locus for pure autosomal dominant FDCM exists, and that this gene is localized to a 9 cM region of 10q21-10q23. The search for the disease causing gene and the responsible mutation(s) is ongoing. Find the latest version: https://jci.me/118922/pdf Gene Mapping of Familial Autosomal Dominant Dilated Cardiomyopathy to Chromosome 10q21-23 Karla R. Bowles,* Robert Gajarski,‡ Patrick Porter,‡ Veronica Goytia,‡ Linda Bachinski,ʈ Robert Roberts,ʈ Ricardo Pignatelli,‡ and Jeffrey A. -

Chromosome 10
Chromosome 10 Description Humans normally have 46 chromosomes in each cell, divided into 23 pairs. Two copies of chromosome 10, one copy inherited from each parent, form one of the pairs. Chromosome 10 spans more than 133 million DNA building blocks (base pairs) and represents between 4 and 4.5 percent of the total DNA in cells. Identifying genes on each chromosome is an active area of genetic research. Because researchers use different approaches to predict the number of genes on each chromosome, the estimated number of genes varies. Chromosome 10 likely contains 700 to 800 genes that provide instructions for making proteins. These proteins perform a variety of different roles in the body. Health Conditions Related to Chromosomal Changes The following chromosomal conditions are associated with changes in the structure or number of copies of chromosome 10. 10q26 deletion syndrome 10q26 deletion syndrome is a condition that results from the loss (deletion) of a small piece of chromosome 10 in each cell. The deletion occurs on the long (q) arm of the chromosome at a position designated 10q26. The signs and symptoms of 10q26 deletion syndrome vary widely, even among affected members of the same family. Affected individuals may have distinctive facial features, growth problems, mild to moderate intellectual disability, developmental delay, genital abnormalities in males, or skeletal or heart defects. People with 10q26 deletion syndrome are missing between 3.5 million and 17 million DNA building blocks (base pairs), also written as 3.5 and 17 megabases (Mb), at position q26 on chromosome 10. The exact size of the deletion varies, and it is unclear what exact region needs to be deleted to cause the condition. -

Chromosome 10 Introduction the Genetic Length of Chromosome 10 Is ~135 Mb. It Is ~4–4.5% of the Whole Human Genome. the Long A
Chromosome 10 ©Chromosome Disorder Outreach Inc. (CDO) Technical genetic content provided by Dr. Iosif Lurie, M.D. Ph.D Medical Geneticist and CDO Medical Consultant/Advisor. Ideogram courtesy of the University of Washington Department of Pathology: ©1994 David Adler.hum_10.gif Introduction The genetic length of chromosome 10 is ~135 Mb. It is ~4–4.5% of the whole human genome. The long arm of this chromosome (90 Mb) is twice as large as the short arm (45 Mb). This chromosome has between 800 and 1,200 genes. Almost 15% of these genes are known to be related to the development of body organs or to cause other types of genetic abnormalities. Different forms of structural abnormalities of this chromosome are reported in ~1,400 patients (including the persons where abnormalities of chromosome 10 were associated with abnormalities of other chromosomes). Various forms of deletions were found in ~750–800 patients. The ratio between persons with deletions of the short arm (~200) and the persons with deletions of the long arm (~550– 600) is close to the ratio between the genetic size of the short and long arms. There are two syndromes caused by deletions of the short arm (Di George syndrome type II and HDR syndrome) and several syndromes caused by deletions of the long arm. Two of these syndromes, which have small deletions of the proximal part of 10q and are associated with Hirschsprung’s disease and Cockayne’s syndrome, became known only after the invention of molecular cytogenetics. Two other syndromes (deletion 10q23 and distal deletions of 10q) have been well known for a long time. -

Partial Monosomy of Chromosome 10 Short Arms
J Med Genet: first published as 10.1136/jmg.20.2.107 on 1 April 1983. Downloaded from Journal of Medical Genetics, 1983, 20, 107-111 Partial monosomy of chromosome 10 short arms A GENCIK*, U BRONNIMANt, R TOBLERt, AND P AUJF DER MAUR* From *Zytogenetisches Labor, Pathologisches Institut der Universitdt, Bern; and tSduglingsspital Elfenau, Bern, Switzerland. SUMMARY Two children with monosmy IOp13 are reported. In the first case the monosomy was the result of a maternal balanced translocation t(3;10) (p27;pl3) while the second case was a de novo mutation. We reviewed clinical details of cases reported so far and found that certain symptoms are typical of the deletion of a comparatively large segment of chromosome 10 short arms. These symptoms include mental and growth retardation, skull abnormalities, antimongoloid slant of the eyes, ear abnormalities, anteverted nostrils, abnormalities of the hands and feet, cryptorchidism in boys, and, primarily, hypoplasia or aplasia of the olfactory bulbs and olfactory tracts as well as narrow palpebral fissures or eyelid ptosis. Since the first description by Elliott et al1 of a Family A probable partial deletion of lOp in a child with many clinical symptoms, a number of reports of patients II 1 12 with lOp partial monosomy have been published,2-7 copyright. thus enlarging the clinical picture. Deletion of part of the chromosome 10 short arms associated with deletion of the long arm terminal segment is also found in cases with ring chromosome 1 2 3 4 5 6 7 8 9 10 11 12 13 10.8-12 As a rule, the deletion of both telomeric Farmily B W Congenital anomalies segments is small and the clinical picture is not as I E} 4 ETranslocation carrier severe as in cases of the above mentioned iOp partial 1 2 E1Normal karyotype http://jmg.bmj.com/ monosomy. -

Chromosome 10 Abnormality Predicts Prognosis of Neuroblastoma Patients with Bone Marrow Metastasis
Jiang et al. Italian Journal of Pediatrics (2021) 47:134 https://doi.org/10.1186/s13052-021-01085-6 RESEARCH Open Access Chromosome 10 abnormality predicts prognosis of neuroblastoma patients with bone marrow metastasis Chi-yi Jiang1, Xiao Xu1, Bing-lin Jian1, Xue Zhang1, Zhi-xia Yue1, Wei Guo2 and Xiao-li Ma1* Abstract Background: Neuroblastoma (NB) is the most common extracranial solid tumor in children. It is known for high heterogeneity and concealed onset. In recent years, the mechanism of its occurrence and development has been gradually revealed. The purpose of this study is to summarize the clinical characteristics of children with NB and abnormal chromosome 10, and to investigate the relationship between the number and structure of chromosome 10 abnormalities and NB prognosis. Methods: Chromosome G-banding was used at the time of diagnosis to evaluate the genetics of chromosomes in patients with NB and track their clinical characteristics and prognosis. All participants were diagnosed with NB in the Medical Oncology Department of the Beijing Children’s Hospital from May 2015 to December 2018 and were followed up with for at least 1 year. Results: Of all 150 patients with bone marrow metastases, 42 were clearly diagnosed with chromosomal abnormalities. Thirteen patients showed abnormalities in chromosome 10, and chromosome 10 was the most commonly missing chromosome. These 13 patients had higher LDH and lower OS and EFS than children with chromosomal abnormalities who did not have an abnormality in chromosome 10. Eight patients had both MYCN amplification and 1p36 deletion. Two patients had optic nerve damage and no vision, and one patient had left supraorbital metastases 5 months after treatment. -

Terminal Deletion of the Long Arm of Chromosome 10 on September 30, 2021 by Guest
J Med Genet: first published as 10.1136/jmg.23.5.478 on 1 October 1986. Downloaded from 478 Case reports using routine methods and analysed after trypsin G abnormalities in balanced X;autosome trans- banding. In addition, lymphocytes were cultured locations, due to dissociation of genes from regulator with BrdU in order to demonstrate the late repli- sites or the spread of inactivation from an area of cating X chromosome. heterochromatin to a portion of the neighbouring Both parents had normal karyotypes, but the translocated chromosome which would not normally proband's karyotype revealed an apparently be inactivated. balanced X;13 translocation: 46,X,t(X;13) (Xpter-- Five of seven previously reported patients with X; Xql3: :13pll->13pter;13qter--13pll: :Xql3--Xqter) autosome translocations with breakpoints at Xq132 (figure). BrdU incorporation showed the normal X to were infertile, but none had other dysmorphic be late labelling in all of 100 cells examined. As the features. Of seven patients reported with deletions of normal X was late replicating in each cell, it was not Xq with breakpoints at Xql3, none had phenotypic possible to detect any spread of inactivation from the abnormalities other than the Turner stigmata.5 Our X to the 13 in the 100 cells examined. patient is thus the first reported case of a balanced X; autosome'translocation with a breakpoint at Xql3 and an abnormal phenotype, other than gonadal Discussion dysgenesis. Although it is easy to see why unbalanced X; autosome translocations cause phenotypic abnorma- References lities due to deletion or duplication of chromosomal Scheres JMJC, Merkx GFM, Hustinx TWJ. -

Chicken Skeletal Muscle-Associated Macroarray for Gene Discovery
Chicken skeletal muscle-associated macroarray for gene discovery E.C. Jorge1, C.M.R. Melo2, M.F. Rosário1, J.R.S. Rossi1, M.C. Ledur3, A.S.A.M.T. Moura4 and L.L. Coutinho1 1Departamento de Zootecnia, Escola Superior de Agricultura “Luiz de Queiroz”, Universidade de São Paulo, Piracicaba, SP, Brasil 2Departamento de Aquicultura, Universidade Federal de Santa Catarina, Florianópolis, SC, Brasil 3Embrapa Suínos e Aves, Genética e Melhoramento Animal, Vila Tamanduá, Concórdia, SC, Brasil 4Departamento de Produção Animal, Faculdade de Medicina Veterinária e Zootecnia de Botucatu, Universidade Estadual Paulista Júlio de Mesquita Filho, Botucatu, SP, Brasil Corresponding author: L.L. Coutinho E-mail: [email protected] Genet. Mol. Res. 9 (1): 188-207 (2010) Received November 4, 2009 Accepted December 2, 2009 Published February 2, 2010 ABSTRACT. Macro- and microarrays are well-established technologies to determine gene functions through repeated measurements of transcript abundance. We constructed a chicken skeletal muscle-associated array based on a muscle-specific EST database, which was used to generate a tissue expression dataset of ~4500 chicken genes across 5 adult tissues (skeletal muscle, heart, liver, brain, and skin). Only a small number of ESTs were sufficiently well characterized by BLAST searches to determine their probable cellular functions. Evidence of a particular tissue-characteristic expression can be considered an indication that the transcript is likely to be functionally significant. The skeletal muscle macroarray platform was first used to search for evidence of tissue- specific expression, focusing on the biological function of genes/ transcripts, since gene expression profiles generated across tissues were found to be reliable and consistent. -

Duplications of 10Q FTNW
Duplications of 10q rarechromo.org Duplications of 10q A duplication of 10q is a chromosome disorder . A chromosome disorder is a change in chromosome number or structure which results in a set of features or symptoms. People with a duplication of 10q have some extra genetic material on one of their 46 chromosomes. A duplication of 10q is sometimes also called a 10q trisomy or a partial 10q trisomy . Chromosomes are made up of DNA held together by proteins. They are rod-like structures in the nucleus of the body’s cells that carry genetic information (known as genes), telling the body how to develop and function. They come in 23 pairs, one from each parent, and 22 of the pairs are numbered 1-22 according to size, from the largest to the smallest. In addition to these 44 chromosomes, each person has another pair of chromosomes, called the sex chromosomes. Girls have two Xs (XX), whereas boys have an X and a Y chromosome (XY). Each chromosome has a short (p) arm (shown at the top in the diagram on the next page) and a long (q) arm (the bottom part of the chromosome). For healthy development, chromosomes should contain just the right amount of material – not too much and not too little. People with a duplication of 10q have one intact chromosome 10, but there is an extra piece from the other and this is likely to affect their learning and physical development. Most of the clinical difficulties are probably caused by the presence of an extra copy (instead of the usual two) of a number of genes. -
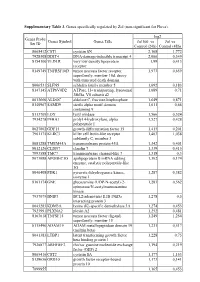
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis