RIT2, a Neuron-Specific Small Guanosine Triphosphatase, Is Expressed in Retinal Neuronal Cells and Its Promoter Is Modulated by the POU4 Transcription Factors
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Ph-Eqtl Interaction at the RIT2-SYT4 Parkinson's Disease Risk
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.16.423140; this version posted June 4, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. A pH-eQTL interaction at the RIT2-SYT4 Parkinson’s disease risk locus in the substantia nigra Authors and affiliations: Sejal Patel1*, Derek Howard1, Leon French1,2,3,4* 1. Krembil Centre for Neuroinformatics, Centre for Addiction and Mental Health, Toronto, ON, Canada 2. Campbell Family Mental Health Research Institute, Centre for Addiction and Mental Health, Toronto, Canada 3. Department of Psychiatry, University of Toronto, Toronto 4. Institute for Medical Science, University of Toronto, Toronto, Canada Abstract Parkinson's disease (PD) causes severe motor and cognitive disabilities that result from the progressive loss of dopamine neurons in the substantia nigra. The rs12456492 variant in the RIT2 gene has been repeatedly associated with increased risk for Parkinson's disease. From a transcriptomic perspective, a meta-analysis found that RIT2 gene expression is correlated with pH in the human brain. To assess these pH associations in relation to PD risk, we examined the two datasets that assayed rs12456492, gene expression, and pH in the postmortem human brain. Using the BrainEAC dataset, we replicate the positive correlation between RIT2 gene expression and pH in the human brain (n=100). Furthermore, we found that the relationship between expression and pH is influenced by rs12456492. -

Zbtb16 Regulates Social Cognitive Behaviors and Neocortical
Usui et al. Translational Psychiatry (2021) 11:242 https://doi.org/10.1038/s41398-021-01358-y Translational Psychiatry ARTICLE Open Access Zbtb16 regulates social cognitive behaviors and neocortical development Noriyoshi Usui 1,2,3,4, Stefano Berto5,AmiKonishi1, Makoto Kondo1,4, Genevieve Konopka5,HideoMatsuzaki 2,6,7 and Shoichi Shimada1,2,4 Abstract Zinc finger and BTB domain containing 16 (ZBTB16) play the roles in the neural progenitor cell proliferation and neuronal differentiation during development, however, how the function of ZBTB16 is involved in brain function and behaviors unknown. Here we show the deletion of Zbtb16 in mice leads to social impairment, repetitive behaviors, risk- taking behaviors, and cognitive impairment. To elucidate the mechanism underlying the behavioral phenotypes, we conducted histological analyses and observed impairments in thinning of neocortical layer 6 (L6) and a reduction of TBR1+ neurons in Zbtb16 KO mice. Furthermore, we found increased dendritic spines and microglia, as well as developmental defects in oligodendrocytes and neocortical myelination in the prefrontal cortex (PFC) of Zbtb16 KO mice. Using genomics approaches, we identified the Zbtb16 transcriptome that includes genes involved in neocortical maturation such as neurogenesis and myelination, and both autism spectrum disorder (ASD) and schizophrenia (SCZ) pathobiology. Co-expression networks further identified Zbtb16-correlated modules that are unique to ASD or SCZ, respectively. Our study provides insight into the novel roles of ZBTB16 in behaviors and neocortical development related to the disorders. 1234567890():,; 1234567890():,; 1234567890():,; 1234567890():,; Introduction identified as a causative mutation for skeletal defects, ZBTB16 (PLZF) encodes a transcription factor, which genital hypoplasia, and mental retardation (SGYMR)6,7. -

Identification of Potential Key Genes and Pathway Linked with Sporadic Creutzfeldt-Jakob Disease Based on Integrated Bioinformatics Analyses
medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Identification of potential key genes and pathway linked with sporadic Creutzfeldt-Jakob disease based on integrated bioinformatics analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Abstract Sporadic Creutzfeldt-Jakob disease (sCJD) is neurodegenerative disease also called prion disease linked with poor prognosis. The aim of the current study was to illuminate the underlying molecular mechanisms of sCJD. The mRNA microarray dataset GSE124571 was downloaded from the Gene Expression Omnibus database. Differentially expressed genes (DEGs) were screened. -

Gene Expression Responses to DNA Damage Are Altered in Human Aging and in Werner Syndrome
Oncogene (2005) 24, 5026–5042 & 2005 Nature Publishing Group All rights reserved 0950-9232/05 $30.00 www.nature.com/onc Gene expression responses to DNA damage are altered in human aging and in Werner Syndrome Kasper J Kyng1,2, Alfred May1, Tinna Stevnsner2, Kevin G Becker3, Steen Klvra˚ 4 and Vilhelm A Bohr*,1 1Laboratory of Molecular Gerontology, National Institute on Aging, National Institutes of Health, 5600 Nathan Shock Drive, Baltimore, MD 21224, USA; 2Danish Center for Molecular Gerontology, Department of Molecular Biology, University of Aarhus, DK-8000 Aarhus C, Denmark; 3Gene Expression and Genomics Unit, National Institute on Aging, National Institutes of Health, Baltimore, MD 21224, USA; 4Institute for Human Genetics, University of Aarhus, Denmark The accumulation of DNA damage and mutations is syndromes, caused by heritable mutations inactivating considered a major cause of cancer and aging. While it is proteins that sense or repair DNA damage, which known that DNA damage can affect changes in gene accelerate some but not all signs of normal aging (Hasty expression, transcriptional regulation after DNA damage et al., 2003). Age is associated withan increase in is poorly understood. We characterized the expression of susceptibility to various forms of stress, and sporadic 6912 genes in human primary fibroblasts after exposure to reports suggest that an age-related decrease in DNA three different kinds of cellular stress that introduces repair may increase the susceptibility of cells to agents DNA damage: 4-nitroquinoline-1-oxide (4NQO), c-irra- causing DNA damage. Reduced base excision repair has diation, or UV-irradiation. Each type of stress elicited been demonstrated in nuclear extracts from aged human damage specific gene expression changes of up to 10-fold. -

Experimental Eye Research 129 (2014) 93E106
Experimental Eye Research 129 (2014) 93e106 Contents lists available at ScienceDirect Experimental Eye Research journal homepage: www.elsevier.com/locate/yexer Transcriptomic analysis across nasal, temporal, and macular regions of human neural retina and RPE/choroid by RNA-Seq S. Scott Whitmore a, b, Alex H. Wagner a, c, Adam P. DeLuca a, b, Arlene V. Drack a, b, Edwin M. Stone a, b, Budd A. Tucker a, b, Shemin Zeng a, b, Terry A. Braun a, b, c, * Robert F. Mullins a, b, Todd E. Scheetz a, b, c, a Stephen A. Wynn Institute for Vision Research, The University of Iowa, Iowa City, IA, USA b Department of Ophthalmology and Visual Sciences, Carver College of Medicine, The University of Iowa, Iowa City, IA, USA c Department of Biomedical Engineering, College of Engineering, The University of Iowa, Iowa City, IA, USA article info abstract Article history: Proper spatial differentiation of retinal cell types is necessary for normal human vision. Many retinal Received 14 September 2014 diseases, such as Best disease and male germ cell associated kinase (MAK)-associated retinitis pigmen- Received in revised form tosa, preferentially affect distinct topographic regions of the retina. While much is known about the 31 October 2014 distribution of cell types in the retina, the distribution of molecular components across the posterior pole Accepted in revised form 4 November 2014 of the eye has not been well-studied. To investigate regional difference in molecular composition of Available online 5 November 2014 ocular tissues, we assessed differential gene expression across the temporal, macular, and nasal retina and retinal pigment epithelium (RPE)/choroid of human eyes using RNA-Seq. -

Mouse Rit2 Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Rit2 Knockout Project (CRISPR/Cas9) Objective: To create a Rit2 knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Rit2 gene (NCBI Reference Sequence: NM_009065 ; Ensembl: ENSMUSG00000057455 ) is located on Mouse chromosome 18. 5 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 5 (Transcript: ENSMUST00000153060). Exon 3 will be selected as target site. Cas9 and gRNA will be co-injected into fertilized eggs for KO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Exon 3 starts from about 24.73% of the coding region. Exon 3 covers 11.37% of the coding region. The size of effective KO region: ~74 bp. The KO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 3 5 Legends Exon of mouse Rit2 Knockout region Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot (up) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section upstream of Exon 3 is aligned with itself to determine if there are tandem repeats. No significant tandem repeat is found in the dot plot matrix. So this region is suitable for PCR screening or sequencing analysis. Overview of the Dot Plot (down) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section downstream of Exon 3 is aligned with itself to determine if there are tandem repeats. -

Differential Gene Expression Profile of Human Neutrophils Cultured with Plasmodium Falciparum-Parasitized Title Erythrocytes
Differential Gene Expression Profile of Human Neutrophils Cultured with Plasmodium falciparum-Parasitized Title Erythrocytes Author(s) Terkawi, Mohamad Alaa; Takano, Ryo; Kato, Kentaro Journal of immunology research, 2018, 6709424 Citation https://doi.org/10.1155/2018/6709424 Issue Date 2018-07-05 Doc URL http://hdl.handle.net/2115/71366 Rights(URL) http://creativecommons.org/licenses/by/4.0/ Type article File Information 6709424.pdf Instructions for use Hokkaido University Collection of Scholarly and Academic Papers : HUSCAP Hindawi Journal of Immunology Research Volume 2018, Article ID 6709424, 8 pages https://doi.org/10.1155/2018/6709424 Research Article Differential Gene Expression Profile of Human Neutrophils Cultured with Plasmodium falciparum-Parasitized Erythrocytes 1,2 1 1,3 Mohamad Alaa Terkawi, Ryo Takano, and Kentaro Kato 1National Research Center for Protozoan Diseases, Obihiro University of Agriculture and Veterinary Medicine, Inada-cho, Obihiro, Hokkaido 080-8555, Japan 2Department of Orthopedic Surgery, Faculty of Medicine and Graduate School of Medicine, Hokkaido University, Kita-15, Nish-7, Kita-ku, Sapporo 060-8638, Japan 3Research Center for Global Agromedicine, Obihiro University of Agriculture and Veterinary Medicine, Inada-cho, Obihiro, Hokkaido 080-8555, Japan Correspondence should be addressed to Kentaro Kato; [email protected] Received 31 January 2018; Revised 8 May 2018; Accepted 15 May 2018; Published 5 July 2018 Academic Editor: Baohui Xu Copyright © 2018 Mohamad Alaa Terkawi et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Neutrophils (PMNs) are the most abundant cellular component of our innate immune system, where they play central roles in the pathogenesis of and resistance to a broad range of diseases. -

Differential Gene Expression Profile of Human Neutrophils Cultured with Plasmodium Falciparum-Parasitized Erythrocytes
Hindawi Journal of Immunology Research Volume 2018, Article ID 6709424, 8 pages https://doi.org/10.1155/2018/6709424 Research Article Differential Gene Expression Profile of Human Neutrophils Cultured with Plasmodium falciparum-Parasitized Erythrocytes 1,2 1 1,3 Mohamad Alaa Terkawi, Ryo Takano, and Kentaro Kato 1National Research Center for Protozoan Diseases, Obihiro University of Agriculture and Veterinary Medicine, Inada-cho, Obihiro, Hokkaido 080-8555, Japan 2Department of Orthopedic Surgery, Faculty of Medicine and Graduate School of Medicine, Hokkaido University, Kita-15, Nish-7, Kita-ku, Sapporo 060-8638, Japan 3Research Center for Global Agromedicine, Obihiro University of Agriculture and Veterinary Medicine, Inada-cho, Obihiro, Hokkaido 080-8555, Japan Correspondence should be addressed to Kentaro Kato; [email protected] Received 31 January 2018; Revised 8 May 2018; Accepted 15 May 2018; Published 5 July 2018 Academic Editor: Baohui Xu Copyright © 2018 Mohamad Alaa Terkawi et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Neutrophils (PMNs) are the most abundant cellular component of our innate immune system, where they play central roles in the pathogenesis of and resistance to a broad range of diseases. However, their roles in malarial infection remain poorly understood. Therefore, we examined the transcriptional gene profile of human PMNs in response to Plasmodium falciparum-parasitized erythrocytes (iRBCs) by using oligonucleotide microarrays. Results revealed that PMNs induced a broad and vigorous set of changes in gene expression in response to malarial parasites, represented by 118 upregulated and 216 downregulated genes. -
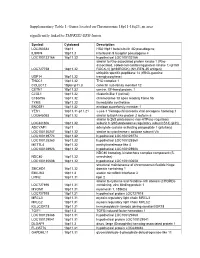
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

The Genetics of Parkinson's Disease and Implications for Clinical Practice
G C A T T A C G G C A T genes Review The Genetics of Parkinson’s Disease and Implications for Clinical Practice Jacob Oliver Day 1 and Stephen Mullin 1,2,* 1 Faculty of Health, University of Plymouth, Plymouth PL4 8AA, UK; [email protected] 2 Department of Clinical and Movement Neurosciences, University College London Institute of Neurology, London WC1N 3BG, UK * Correspondence: [email protected] Abstract: The genetic landscape of Parkinson’s disease (PD) is characterised by rare high penetrance pathogenic variants causing familial disease, genetic risk factor variants driving PD risk in a signif- icant minority in PD cases and high frequency, low penetrance variants, which contribute a small increase of the risk of developing sporadic PD. This knowledge has the potential to have a major impact in the clinical care of people with PD. We summarise these genetic influences and discuss the implications for therapeutics and clinical trial design. Keywords: Parkinson’s disease; genetics; precision medicine; clinical trials; monogenic; polygenic 1. Introduction Parkinson’s disease (PD) is a neurodegenerative condition affecting over 6 million people worldwide that is expected to double in prevalence by 2040 [1]. It is characterised by a core set of movement (motor) abnormalities - slowness of movement, muscle rigidity Citation: Day, J.O.; Mullin, S. The and tremor – as well as a number of non-motor features such as constipation, anxiety and Genetics of Parkinson’s Disease and dementia [2]. There is often a prodromal phase of non-motor symptoms which precede Implications for Clinical Practice. motor symptoms by many years [3]. -

Meta-Analysis of Parkinson Disease: Identification of a Novel Locus, RIT2 Nathan Pankratz, Indiana University Gary W
Meta-analysis of Parkinson disease: Identification of a novel locus, RIT2 Nathan Pankratz, Indiana University Gary W. Beecham, University of Miami Anita L. DeStefano, Boston University Ted M. Dawson, Johns Hopkins University Kimberly F. Doheny, Johns Hopkins University Stewart Factor, Emory University Taye H. Hamza, New York State Department of Health Albert Y. Hung, Massachusetts General Hospital Bradley T. Hyman, Massachusetts General Hospital Adrian J. Ivinson, Harvard University Only first 10 authors above; see publication for full author list. Journal Title: Annals of Neurology Volume: Volume 71, Number 3 Publisher: Wiley: 12 months | 2012-03-01, Pages 370-384 Type of Work: Article | Post-print: After Peer Review Publisher DOI: 10.1002/ana.22687 Permanent URL: https://pid.emory.edu/ark:/25593/sq7bb Final published version: http://dx.doi.org/10.1002/ana.22687 Copyright information: © 2012 American Neurological Association. Accessed September 26, 2021 9:07 PM EDT NIH Public Access Author Manuscript Ann Neurol. Author manuscript; available in PMC 2013 March 01. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Ann Neurol. 2012 March ; 71(3): 370–384. doi:10.1002/ana.22687. Meta-analysis of Parkinson disease: Identification of a novel locus, RIT2 Nathan Pankratz, Ph.D.1, Gary W. Beecham, Ph.D.2, Anita L. DeStefano, Ph.D.3,4, Ted M. Dawson, M.D., Ph.D.5, Kimberly F. Doheny, Ph.D.5, Stewart A. Factor, D.O.6, Taye H. Hamza, Ph.D.7, Albert Y. Hung, M.D., Ph.D.8,9, Bradley T. Hyman, M.D., Ph.D.8,10, Adrian J. -

Rnai Screening Reveals Requirement for Host Cell Secretory Pathway in Infection by Diverse Families of Negative-Strand RNA Viruses
RNAi screening reveals requirement for host cell secretory pathway in infection by diverse families of negative-strand RNA viruses Debasis Pandaa,b, Anshuman Dasa,b, Phat X. Dinha,b, Sakthivel Subramaniama,b, Debasis Nayakc, Nicholas J. Barrowsd, James L. Pearsond, Jesse Thompsonb, David L. Kellye, Istvan Ladungaf, and Asit K. Pattnaika,b,1 aSchool of Veterinary Medicine and Biomedical Sciences and bNebraska Center for Virology, University of Nebraska, Lincoln, NE 68583; cNational Institute of Neurological Disorders and Stroke, National Institutes of Health, Bethesda, MD 20892; dDuke RNAi Screening Facility, Duke University Medical Center, Durham, NC 27710; eUniversity of Nebraska Medical Center, Omaha, NE 68198; and fDepartment of Statistics, University of Nebraska, Lincoln, NE 68588 Edited by Peter Palese, Mount Sinai School of Medicine, New York, NY, and approved October 17, 2011 (received for review August 19, 2011) Negative-strand (NS) RNA viruses comprise many pathogens that and other NS RNA virus infections. Identifying the cellular factors cause serious diseases in humans and animals. Despite their clinical and studying the mechanisms of their involvement in these viral importance, little is known about the host factors required for their infections is important not only for understanding the biology of infection. Using vesicular stomatitis virus (VSV), a prototypic NS RNA these pathogens, but also for development of antiviral therapeutics. virus in the family Rhabdoviridae, we conducted a human genome- The advent of siRNA technology and the availability of ge- wide siRNA screen and identified 72 host genes required for viral nome-wide siRNA libraries have been useful in identifying host fl infection. Many of these identified genes were also required for factors required for in uenza virus, an NS RNA virus, and several – infection by two other NS RNA viruses, the lymphocytic choriome- positive-strand RNA viruses, as well as HIV (10 19).