Phd Thesis JM
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

An Animal Model with a Cardiomyocyte-Specific Deletion of Estrogen Receptor Alpha: Functional, Metabolic, and Differential Netwo
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2014 An animal model with a cardiomyocyte-specific deletion of estrogen receptor alpha: Functional, metabolic, and differential network analysis Sriram Devanathan Washington University School of Medicine in St. Louis Timothy Whitehead Washington University School of Medicine in St. Louis George G. Schweitzer Washington University School of Medicine in St. Louis Nicole Fettig Washington University School of Medicine in St. Louis Attila Kovacs Washington University School of Medicine in St. Louis See next page for additional authors Follow this and additional works at: https://digitalcommons.wustl.edu/open_access_pubs Recommended Citation Devanathan, Sriram; Whitehead, Timothy; Schweitzer, George G.; Fettig, Nicole; Kovacs, Attila; Korach, Kenneth S.; Finck, Brian N.; and Shoghi, Kooresh I., ,"An animal model with a cardiomyocyte-specific deletion of estrogen receptor alpha: Functional, metabolic, and differential network analysis." PLoS One.9,7. e101900. (2014). https://digitalcommons.wustl.edu/open_access_pubs/3326 This Open Access Publication is brought to you for free and open access by Digital Commons@Becker. It has been accepted for inclusion in Open Access Publications by an authorized administrator of Digital Commons@Becker. For more information, please contact [email protected]. Authors Sriram Devanathan, Timothy Whitehead, George G. Schweitzer, Nicole Fettig, Attila Kovacs, Kenneth S. Korach, Brian N. Finck, and Kooresh I. Shoghi This open access publication is available at Digital Commons@Becker: https://digitalcommons.wustl.edu/open_access_pubs/3326 An Animal Model with a Cardiomyocyte-Specific Deletion of Estrogen Receptor Alpha: Functional, Metabolic, and Differential Network Analysis Sriram Devanathan1, Timothy Whitehead1, George G. Schweitzer2, Nicole Fettig1, Attila Kovacs3, Kenneth S. -

Pharmacogenomic Characterization in Bipolar Spectrum Disorders
pharmaceutics Review Pharmacogenomic Characterization in Bipolar Spectrum Disorders Stefano Fortinguerra 1,2 , Vincenzo Sorrenti 1,2,3 , Pietro Giusti 2, Morena Zusso 2 and Alessandro Buriani 1,2,* 1 Maria Paola Belloni Center for Personalized Medicine, Data Medica Group (Synlab Limited), 35131 Padova, Italy; [email protected] (S.F.); [email protected] (V.S.) 2 Department of Pharmaceutical & Pharmacological Sciences, University of Padova, 35131 Padova, Italy; [email protected] (P.G.); [email protected] (M.Z.) 3 Bendessere™ Study Center, Solgar Italia Multinutrient S.p.A., 35131 Padova, Italy * Correspondence: [email protected] Received: 25 November 2019; Accepted: 19 December 2019; Published: 21 December 2019 Abstract: The holistic approach of personalized medicine, merging clinical and molecular characteristics to tailor the diagnostic and therapeutic path to each individual, is steadily spreading in clinical practice. Psychiatric disorders represent one of the most difficult diagnostic challenges, given their frequent mixed nature and intrinsic variability, as in bipolar disorders and depression. Patients misdiagnosed as depressed are often initially prescribed serotonergic antidepressants, a treatment that can exacerbate a previously unrecognized bipolar condition. Thanks to the use of the patient’s genomic profile, it is possible to recognize such risk and at the same time characterize specific genetic assets specifically associated with bipolar spectrum disorder, as well as with the individual response to the various therapeutic options. This provides the basis for molecular diagnosis and the definition of pharmacogenomic profiles, thus guiding therapeutic choices and allowing a safer and more effective use of psychotropic drugs. Here, we report the pharmacogenomics state of the art in bipolar disorders and suggest an algorithm for therapeutic regimen choice. -

NATURAL KILLER CELLS, HYPOXIA, and EPIGENETIC REGULATION of HEMOCHORIAL PLACENTATION by Damayanti Chakraborty Submitted to the G
NATURAL KILLER CELLS, HYPOXIA, AND EPIGENETIC REGULATION OF HEMOCHORIAL PLACENTATION BY Damayanti Chakraborty Submitted to the graduate degree program in Pathology and Laboratory Medicine and the Graduate Faculty of the University of Kansas in partial fulfillment ofthe requirements for the degree of Doctor of Philosophy. ________________________________ Chair: Michael J. Soares, Ph.D. ________________________________ Jay Vivian, Ph.D. ________________________________ Patrick Fields, Ph.D. ________________________________ Soumen Paul, Ph.D. ________________________________ Michael Wolfe, Ph.D. ________________________________ Adam J. Krieg, Ph.D. Date Defended: 04/01/2013 The Dissertation Committee for Damayanti Chakraborty certifies that this is the approved version of the following dissertation: NATURAL KILLER CELLS, HYPOXIA, AND EPIGENETIC REGULATION OF HEMOCHORIAL PLACENTATION ________________________________ Chair: Michael J. Soares, Ph.D. Date approved: 04/01/2013 ii ABSTRACT During the establishment of pregnancy, uterine stromal cells differentiate into decidual cells and recruit natural killer (NK) cells. These NK cells are characterized by low cytotoxicity and distinct cytokine production. In rodent as well as in human pregnancy, the uterine NK cells peak in number around mid-gestation after which they decline. NK cells associate with uterine spiral arteries and are implicated in pregnancy associated vascular remodeling processes and potentially in modulating trophoblast invasion. Failure of trophoblast invasion and vascular remodeling has been shown to be associated with pathological conditions like preeclampsia syndrome, hypertension in mother and/or fetal growth restriction. We hypothesize that NK cells fundamentally contribute to the organization of the placentation site. In order to study the in vivo role of NK cells during pregnancy, gestation stage- specific NK cell depletion was performed in rats using anti asialo GM1 antibodies. -

140503 IPF Signatures Supplement Withfigs Thorax
Supplementary material for Heterogeneous gene expression signatures correspond to distinct lung pathologies and biomarkers of disease severity in idiopathic pulmonary fibrosis Daryle J. DePianto1*, Sanjay Chandriani1⌘*, Alexander R. Abbas1, Guiquan Jia1, Elsa N. N’Diaye1, Patrick Caplazi1, Steven E. Kauder1, Sabyasachi Biswas1, Satyajit K. Karnik1#, Connie Ha1, Zora Modrusan1, Michael A. Matthay2, Jasleen Kukreja3, Harold R. Collard2, Jackson G. Egen1, Paul J. Wolters2§, and Joseph R. Arron1§ 1Genentech Research and Early Development, South San Francisco, CA 2Department of Medicine, University of California, San Francisco, CA 3Department of Surgery, University of California, San Francisco, CA ⌘Current address: Novartis Institutes for Biomedical Research, Emeryville, CA. #Current address: Gilead Sciences, Foster City, CA. *DJD and SC contributed equally to this manuscript §PJW and JRA co-directed this project Address correspondence to Paul J. Wolters, MD University of California, San Francisco Department of Medicine Box 0111 San Francisco, CA 94143-0111 [email protected] or Joseph R. Arron, MD, PhD Genentech, Inc. MS 231C 1 DNA Way South San Francisco, CA 94080 [email protected] 1 METHODS Human lung tissue samples Tissues were obtained at UCSF from clinical samples from IPF patients at the time of biopsy or lung transplantation. All patients were seen at UCSF and the diagnosis of IPF was established through multidisciplinary review of clinical, radiological, and pathological data according to criteria established by the consensus classification of the American Thoracic Society (ATS) and European Respiratory Society (ERS), Japanese Respiratory Society (JRS), and the Latin American Thoracic Association (ALAT) (ref. 5 in main text). Non-diseased normal lung tissues were procured from lungs not used by the Northern California Transplant Donor Network. -

Methotrexate Induces Hyperplasia of Enterochromaffin
112 BPB Reports 4, 112-115 (2021) Vol. 4, No. 4 Report Methotrexate Induces Hyperplasia of Enterochromaffin Cells in Mouse Jejunum Takuji Machida,*,a Airi Tanaka,a Megumi Kobayashi,a Kohei Imai,a Masahiko Hirafuji,a,b and Kenji Iizukaa aDepartment of Pharmacological Sciences, School of Pharmaceutical Sciences, Health Sciences University of Hokkaido, 1757 Kanazawa, Ishikari-Tobetsu, Hokkaido 061-0293, Japan; bPresent address: School of Health Sciences, Iryo Sosei University, 5-5-1 Chuodai Iino, Iwaki, Fukushima 970-8551, Japan Received May 12, 2021; Accepted July 26, 2021 5-Hydroxytryptamine (5-HT) is synthesized by L-tryptophan hydroxylase (TPH) and is stored mainly in enterochromaffin cells of the mucosal epithelium. We previously reported that administration of methotrex- ate, an anticancer agent, to rats caused hyperplasia of enterochromaffin cells, and nitric oxide (NO) might be involved in the underlying mechanism. The aim of this study was to clarify the effect of methotrexate on hyper- plasia of enterochromaffin cells in mice. C57BL/6J mice were intraperitoneally injected with methotrexate or saline as a control. Methotrexate caused an increase in the number of TPH-expressing cells (i.e., enterochromaf- fin cells) in the jejunum. Methotrexate also increased inducible, but not constitutive, NOS mRNA expression. Our results indicate that methotrexate potentiates 5-HT synthesis in mice, as we previously found in rats. Key words methotrexate, intestine, 5-hydroxytriptamine, inducible nitric oxide synthase INTRODUCTION the effects of methotrexate are similar to those previously found in rats.4,6) The use of high doses of methotrexate, an anti-cancer drug, is associated with gastrointestinal damage, which leads to mal- MATERIALS AND METHODS absorption, diarrhea, and weight loss.1,2) In addition, metho- trexate causes moderate emesis in humans and experimental Materials Methotrexate was obtained from Pfizer Co., Ltd. -

Electronic Supplementary Material (ESI) for Metallomics
Electronic Supplementary Material (ESI) for Metallomics. This journal is © The Royal Society of Chemistry 2018 Uniprot Entry name Gene names Protein names Predicted Pattern Number of Iron role EC number Subcellular Membrane Involvement in disease Gene ontology (biological process) Id iron ions location associated 1 P46952 3HAO_HUMAN HAAO 3-hydroxyanthranilate 3,4- H47-E53-H91 1 Fe cation Catalytic 1.13.11.6 Cytoplasm No NAD biosynthetic process [GO:0009435]; neuron cellular homeostasis dioxygenase (EC 1.13.11.6) (3- [GO:0070050]; quinolinate biosynthetic process [GO:0019805]; response to hydroxyanthranilate oxygenase) cadmium ion [GO:0046686]; response to zinc ion [GO:0010043]; tryptophan (3-HAO) (3-hydroxyanthranilic catabolic process [GO:0006569] acid dioxygenase) (HAD) 2 O00767 ACOD_HUMAN SCD Acyl-CoA desaturase (EC H120-H125-H157-H161; 2 Fe cations Catalytic 1.14.19.1 Endoplasmic Yes long-chain fatty-acyl-CoA biosynthetic process [GO:0035338]; unsaturated fatty 1.14.19.1) (Delta(9)-desaturase) H160-H269-H298-H302 reticulum acid biosynthetic process [GO:0006636] (Delta-9 desaturase) (Fatty acid desaturase) (Stearoyl-CoA desaturase) (hSCD1) 3 Q6ZNF0 ACP7_HUMAN ACP7 PAPL PAPL1 Acid phosphatase type 7 (EC D141-D170-Y173-H335 1 Fe cation Catalytic 3.1.3.2 Extracellular No 3.1.3.2) (Purple acid space phosphatase long form) 4 Q96SZ5 AEDO_HUMAN ADO C10orf22 2-aminoethanethiol dioxygenase H112-H114-H193 1 Fe cation Catalytic 1.13.11.19 Unknown No oxidation-reduction process [GO:0055114]; sulfur amino acid catabolic process (EC 1.13.11.19) (Cysteamine -
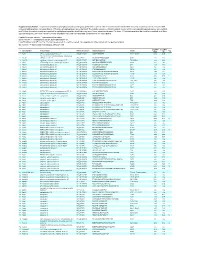
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

Influence of CYP2D6 and CYP2C19 Gene Variants on Antidepressant
The Pharmacogenomics Journal (2014) 14, 176–181 & 2014 Macmillan Publishers Limited All rights reserved 1470-269X/14 www.nature.com/tpj ORIGINAL ARTICLE Influence of CYP2D6 and CYP2C19 gene variants on antidepressant response in obsessive-compulsive disorder EJ Brandl1,2, AK Tiwari1,2, X Zhou2,3, J Deluce2,3, JL Kennedy1,2,DJMu¨ ller1,2,4 and MA Richter1,2,3,4 Numerous studies have reported on pharmacogenetics of antidepressant response in depression. In contrast, little is known of response predictors in obsessive-compulsive disorder (OCD), a disorder with among the lowest proportion of responders to medication (40–60%). Our study is the largest investigation to date (N ¼ 184) of treatment response and side effects to antidepressants in OCD based on metabolizer status for CYP2D6 and CYP2C19. We observed significantly more failed medication trials in CYP2D6 non-extensive compared with extensive metabolizers (P ¼ 0.007). CYP2D6 metabolizer status was associated with side effects to venlafaxine (P ¼ 0.022). There were nonsignificant trends for association of CYP2D6 metabolizer status with response to fluoxetine (P ¼ 0.056) and of CYP2C19 metabolizer status with response to sertraline (P ¼ 0.064). Our study is the first to indicate that CYP genes may have a role in antidepressant response in OCD. More research is required for a future clinical application of genetic testing, which could lead to improved treatment outcomes. The Pharmacogenomics Journal (2014) 14, 176–181; doi:10.1038/tpj.2013.12; published online 2 April 2013 Keywords: obsessive-compulsive disorder (OCD); pharmacogenetics; CYP2D6; CYP2C19; treatment response INTRODUCTION OCD is widely accepted as having strong genetic influence, with Obsessive-compulsive disorder (OCD) occurs in approximately two replicated findings reported for the glutamate transporter gene percent of the population1 and is characterized by a broad variety SLC1A1 and several serotonergic, dopaminergic and other 11 of obsessions and/or compulsions causing significant distress for candidate genes. -

The Genetics of Bipolar Disorder
Molecular Psychiatry (2008) 13, 742–771 & 2008 Nature Publishing Group All rights reserved 1359-4184/08 $30.00 www.nature.com/mp FEATURE REVIEW The genetics of bipolar disorder: genome ‘hot regions,’ genes, new potential candidates and future directions A Serretti and L Mandelli Institute of Psychiatry, University of Bologna, Bologna, Italy Bipolar disorder (BP) is a complex disorder caused by a number of liability genes interacting with the environment. In recent years, a large number of linkage and association studies have been conducted producing an extremely large number of findings often not replicated or partially replicated. Further, results from linkage and association studies are not always easily comparable. Unfortunately, at present a comprehensive coverage of available evidence is still lacking. In the present paper, we summarized results obtained from both linkage and association studies in BP. Further, we indicated new potential interesting genes, located in genome ‘hot regions’ for BP and being expressed in the brain. We reviewed published studies on the subject till December 2007. We precisely localized regions where positive linkage has been found, by the NCBI Map viewer (http://www.ncbi.nlm.nih.gov/mapview/); further, we identified genes located in interesting areas and expressed in the brain, by the Entrez gene, Unigene databases (http://www.ncbi.nlm.nih.gov/entrez/) and Human Protein Reference Database (http://www.hprd.org); these genes could be of interest in future investigations. The review of association studies gave interesting results, as a number of genes seem to be definitively involved in BP, such as SLC6A4, TPH2, DRD4, SLC6A3, DAOA, DTNBP1, NRG1, DISC1 and BDNF. -

Chemical Agent and Antibodies B-Raf Inhibitor RAF265
Supplemental Materials and Methods: Chemical agent and antibodies B-Raf inhibitor RAF265 [5-(2-(5-(trifluromethyl)-1H-imidazol-2-yl)pyridin-4-yloxy)-N-(4-trifluoromethyl)phenyl-1-methyl-1H-benzp{D, }imidazol-2- amine] was kindly provided by Novartis Pharma AG and dissolved in solvent ethanol:propylene glycol:2.5% tween-80 (percentage 6:23:71) for oral delivery to mice by gavage. Antibodies to phospho-ERK1/2 Thr202/Tyr204(4370), phosphoMEK1/2(2338 and 9121)), phospho-cyclin D1(3300), cyclin D1 (2978), PLK1 (4513) BIM (2933), BAX (2772), BCL2 (2876) were from Cell Signaling Technology. Additional antibodies for phospho-ERK1,2 detection for western blot were from Promega (V803A), and Santa Cruz (E-Y, SC7383). Total ERK antibody for western blot analysis was K-23 from Santa Cruz (SC-94). Ki67 antibody (ab833) was from ABCAM, Mcl1 antibody (559027) was from BD Biosciences, Factor VIII antibody was from Dako (A082), CD31 antibody was from Dianova, (DIA310), and Cot antibody was from Santa Cruz Biotechnology (sc-373677). For the cyclin D1 second antibody staining was with an Alexa Fluor 568 donkey anti-rabbit IgG (Invitrogen, A10042) (1:200 dilution). The pMEK1 fluorescence was developed using the Alexa Fluor 488 chicken anti-rabbit IgG second antibody (1:200 dilution).TUNEL staining kits were from Promega (G2350). Mouse Implant Studies: Biopsy tissues were delivered to research laboratory in ice-cold Dulbecco's Modified Eagle Medium (DMEM) buffer solution. As the tissue mass available from each biopsy was limited, we first passaged the biopsy tissue in Balb/c nu/Foxn1 athymic nude mice (6-8 weeks of age and weighing 22-25g, purchased from Harlan Sprague Dawley, USA) to increase the volume of tumor for further implantation. -

Application of Microrna Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease
Article Application of microRNA Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease Jennifer L. Major, Rushita A. Bagchi * and Julie Pires da Silva * Department of Medicine, Division of Cardiology, University of Colorado Anschutz Medical Campus, Aurora, CO 80045, USA; [email protected] * Correspondence: [email protected] (R.A.B.); [email protected] (J.P.d.S.) Supplementary Tables Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 www.mdpi.com/journal/mps Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 2 of 25 Table 1. List of all hsa-miRs identified by Human microRNA Disease Database (HMDD; v3.2) analysis. hsa-miRs were identified using the term “genetics” and “circulating” as input in HMDD. Targets CAD hsa-miR-1 Targets IR injury hsa-miR-423 Targets Obesity hsa-miR-499 hsa-miR-146a Circulating Obesity Genetics CAD hsa-miR-423 hsa-miR-146a Circulating CAD hsa-miR-149 hsa-miR-499 Circulating IR Injury hsa-miR-146a Circulating Obesity hsa-miR-122 Genetics Stroke Circulating CAD hsa-miR-122 Circulating Stroke hsa-miR-122 Genetics Obesity Circulating Stroke hsa-miR-26b hsa-miR-17 hsa-miR-223 Targets CAD hsa-miR-340 hsa-miR-34a hsa-miR-92a hsa-miR-126 Circulating Obesity Targets IR injury hsa-miR-21 hsa-miR-423 hsa-miR-126 hsa-miR-143 Targets Obesity hsa-miR-21 hsa-miR-223 hsa-miR-34a hsa-miR-17 Targets CAD hsa-miR-223 hsa-miR-92a hsa-miR-126 Targets IR injury hsa-miR-155 hsa-miR-21 Circulating CAD hsa-miR-126 hsa-miR-145 hsa-miR-21 Targets Obesity hsa-mir-223 hsa-mir-499 hsa-mir-574 Targets IR injury hsa-mir-21 Circulating IR injury Targets Obesity hsa-mir-21 Targets CAD hsa-mir-22 hsa-mir-133a Targets IR injury hsa-mir-155 hsa-mir-21 Circulating Stroke hsa-mir-145 hsa-mir-146b Targets Obesity hsa-mir-21 hsa-mir-29b Methods Protoc. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in