Supplementary Material Identification of Novel Differentially Methylated Sites with Potential As Clinical Predictors of Impaired Respiratory Function and COPD
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Epigenetic Alterations in Human Papillomavirus-Associated Cancers
viruses Review Epigenetic Alterations in Human Papillomavirus-Associated Cancers David Soto ID , Christine Song and Margaret E. McLaughlin-Drubin * Division of Infectious Disease, Department of Medicine, Brigham & Women’s Hospital, Harvard Medical School, 181 Longwood Avenue, Boston, MA 02115, USA; [email protected] (D.S.); [email protected] (C.S.) * Correspondence: [email protected]; Tel.: +1-617-525-4262 Academic Editors: Alison A. McBride and Karl Munger Received: 14 August 2017; Accepted: 25 August 2017; Published: 1 September 2017 Abstract: Approximately 15–20% of human cancers are caused by viruses, including human papillomaviruses (HPVs). Viruses are obligatory intracellular parasites and encode proteins that reprogram the regulatory networks governing host cellular signaling pathways that control recognition by the immune system, proliferation, differentiation, genomic integrity, and cell death. Given that key proteins in these regulatory networks are also subject to mutation in non-virally associated diseases and cancers, the study of oncogenic viruses has also been instrumental to the discovery and analysis of many fundamental cellular processes, including messenger RNA (mRNA) splicing, transcriptional enhancers, oncogenes and tumor suppressors, signal transduction, immune regulation, and cell cycle control. More recently, tumor viruses, in particular HPV, have proven themselves invaluable in the study of the cancer epigenome. Epigenetic silencing or de-silencing of genes can have cellular consequences that are akin to genetic mutations, i.e., the loss and gain of expression of genes that are not usually expressed in a certain cell type and/or genes that have tumor suppressive or oncogenic activities, respectively. Unlike genetic mutations, the reversible nature of epigenetic modifications affords an opportunity of epigenetic therapy for cancer. -

Ethz-A-005708685
Research Collection Doctoral Thesis Molecular mechanisms controlling lymphatic vascular function Author(s): Shin, Jae (Jay) Woo Publication Date: 2008 Permanent Link: https://doi.org/10.3929/ethz-a-005708685 Rights / License: In Copyright - Non-Commercial Use Permitted This page was generated automatically upon download from the ETH Zurich Research Collection. For more information please consult the Terms of use. ETH Library DISS. ETH NO. 17752 MOLECULAR MECHANISMS CONTROLLING LYMPHATIC VASCULAR FUNCTION A dissertation submitted to ETH Zurich for the degree of Doctor of Sciences presented by JAE (JAY) WOO SHIN April 4th, 1981 citizen of United States of America accepted on the recommendation of Prof. Michael Detmar Prof. Dario Neri 2008 TABLE OF CONTENTS 1 SUMMARY........................................................................................................6 1.1 Summary ..............................................................................................................................................7 1.2 Zusammenfassung...............................................................................................................................9 2 INTRODUCTION............................................................................................12 2.1 CHARACTERISTICS OF THE LYMPHATIC VASCULATURE.........................................13 2.1.1 Anatomy and physiology of the lymphatic vasculature......................................................13 2.1.2 Genes and mechanisms in lymphatic development .............................................................15 -

Supp Table 6.Pdf
Supplementary Table 6. Processes associated to the 2037 SCL candidate target genes ID Symbol Entrez Gene Name Process NM_178114 AMIGO2 adhesion molecule with Ig-like domain 2 adhesion NM_033474 ARVCF armadillo repeat gene deletes in velocardiofacial syndrome adhesion NM_027060 BTBD9 BTB (POZ) domain containing 9 adhesion NM_001039149 CD226 CD226 molecule adhesion NM_010581 CD47 CD47 molecule adhesion NM_023370 CDH23 cadherin-like 23 adhesion NM_207298 CERCAM cerebral endothelial cell adhesion molecule adhesion NM_021719 CLDN15 claudin 15 adhesion NM_009902 CLDN3 claudin 3 adhesion NM_008779 CNTN3 contactin 3 (plasmacytoma associated) adhesion NM_015734 COL5A1 collagen, type V, alpha 1 adhesion NM_007803 CTTN cortactin adhesion NM_009142 CX3CL1 chemokine (C-X3-C motif) ligand 1 adhesion NM_031174 DSCAM Down syndrome cell adhesion molecule adhesion NM_145158 EMILIN2 elastin microfibril interfacer 2 adhesion NM_001081286 FAT1 FAT tumor suppressor homolog 1 (Drosophila) adhesion NM_001080814 FAT3 FAT tumor suppressor homolog 3 (Drosophila) adhesion NM_153795 FERMT3 fermitin family homolog 3 (Drosophila) adhesion NM_010494 ICAM2 intercellular adhesion molecule 2 adhesion NM_023892 ICAM4 (includes EG:3386) intercellular adhesion molecule 4 (Landsteiner-Wiener blood group)adhesion NM_001001979 MEGF10 multiple EGF-like-domains 10 adhesion NM_172522 MEGF11 multiple EGF-like-domains 11 adhesion NM_010739 MUC13 mucin 13, cell surface associated adhesion NM_013610 NINJ1 ninjurin 1 adhesion NM_016718 NINJ2 ninjurin 2 adhesion NM_172932 NLGN3 neuroligin -

Genetic Regulation of Tmem106b in the Pathogenesis of Frontotemporal Lobar Degeneration
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2017 Genetic Regulation Of Tmem106b In The Pathogenesis Of Frontotemporal Lobar Degeneration Michael Gallagher University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Genetics Commons, Molecular Biology Commons, and the Neuroscience and Neurobiology Commons Recommended Citation Gallagher, Michael, "Genetic Regulation Of Tmem106b In The Pathogenesis Of Frontotemporal Lobar Degeneration" (2017). Publicly Accessible Penn Dissertations. 2294. https://repository.upenn.edu/edissertations/2294 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/2294 For more information, please contact [email protected]. Genetic Regulation Of Tmem106b In The Pathogenesis Of Frontotemporal Lobar Degeneration Abstract Neurodegenerative diseases are an emerging global health crisis, with the projected global cost of dementia alone expected to exceed $1 trillion, or >1% of world GDP, by 2018. However, there are no disease-modifying treatments for the major neurodegenerative diseases, such as Alzheimer’s disease, Parkinson’s disease, frontotemporal lobar degeneration (FTLD), and amyotrophic lateral sclerosis. Therefore, there is an urgent need for a better understanding of the pathophysiology underlying these diseases. While genome-wide association studies (GWAS) have identified ~200 genetic ariantsv that are associated with risk of developing neurodegenerative disease, the biological mechanisms underlying these associations are largely unknown. This dissertation investigates the mechanisms by which common genetic variation at TMEM106B, a GWAS-identified risk locus for FTLD, influences disease risk. First, using genetic and clinical data from thirty American and European medical centers, I demonstrate that the TMEM106B locus acts as a genetic modifier of a common Mendelian form of FTLD. -

Uniprot Acceprotiens 121 113 Ratio(113/12 114 Ratio
Uniprot Acceprotiens 121 113 ratio(113/12 114 ratio(114/12 115 ratio(115/12 116 ratio(116/12 117 ratio(117/12 118 ratio(118/12 119 ratio(119/121) P02768 Serum albumin OS=Homo s666397.2 862466.6 1.29 593482.1 0.89 2220420.5 3.33 846469.3 1.27 634302.5 0.95 736961.1 1.11 842297.5 1.26 P02760 Protein AMBP OS=Homo s381627.7 294812.3 0.77 474165.8 1.24 203377.3 0.53 349197.6 0.92 346271.7 0.91 328356.1 0.86 411229.3 1.08 B4E1B2 cDNA FLJ53691, highly sim78511.8 107560.1 1.37 85218.8 1.09 199640.4 2.54 90022.3 1.15 73427.3 0.94 82722 1.05 102491.8 1.31 A0A0K0K1HEpididymis secretory sperm 3358.1 4584.8 1.37 4234.8 1.26 8496.1 2.53 4193.7 1.25 3507.1 1.04 3632.2 1.08 4873.3 1.45 D3DNU8 Kininogen 1, isoform CRA_302648.3 294936.6 0.97 257956.9 0.85 193831.3 0.64 290406.7 0.96 313453.3 1.04 279805.5 0.92 228883.9 0.76 B4E1C2 Kininogen 1, isoform CRA_167.9 229.7 1.37 263.2 1.57 278 1.66 326 1.94 265.5 1.58 290.3 1.73 341.5 2.03 O60494 Cubilin OS=Homo sapiens G40132.6 45037.5 1.12 38654.5 0.96 34055.8 0.85 39708.6 0.99 44702.9 1.11 45025.7 1.12 32701.3 0.81 P98164 Low-density lipoprotein rece40915.4 45344.8 1.11 35817.7 0.88 35721.8 0.87 42157.7 1.03 46693.4 1.14 48624 1.19 38847.7 0.95 A0A024RABHeparan sulfate proteoglyca46985.3 43536.1 0.93 49827.7 1.06 33964.3 0.72 44780.9 0.95 46858.6 1.00 47703.5 1.02 37785.7 0.80 P01133 Pro-epidermal growth factor 75270.8 73109.5 0.97 66336.1 0.88 56680.9 0.75 70877.8 0.94 76444.3 1.02 81110.3 1.08 65749.7 0.87 Q6N093 Putative uncharacterized pro47825.3 55632.5 1.16 48428.3 1.01 63601.5 1.33 65204.2 1.36 59384.5 -
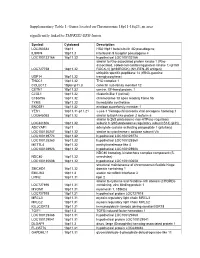
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

Supplementary Data
Supplemental Material Materials and Methods Immunohistochemistry Primary antibodies used for validation studies include: mouse anti-desmoglein-3 (Cat. # 32-6300, Invitrogen, CA, USA; 1:25), rabbit anti-cytokeratin 4 (Cat. # ab11215, Abcam, Cambridge, MA, USA; 1:100), mouse anti-cytokeratin 16 (Cat. # ab8741, Abcam; 1:25), rabbit anti-desmoplakin antibody (Cat. # ab14418, Abcam; 1:200), mouse anti-vimentin (Cat. # M7020, Dako, Carpinteria, CA, USA; 1:100). Secondary antibodies conjugated with biotin (Vector, Burlingame, CA, USA) were used, diluted to 1:400. Tissues slides containing archival FFPE sections, or tissue micro arrays (TMA) consisting of 508 HNSCC and controls, were dewaxed in SafeClear II (Fisher Scientific, Pittsburgh, PA, USA) hydrated through graded alcohols, immersed in 3% hydrogen peroxide in PBS for 30 min to quench the endogenous peroxidase, and processed for antigen retrieval and immunostaining with the appropriate primary antibodies and biotinylated secondary antibodies as described (1), followed by the avidin-biotin complex method (Vector Stain Elite, ABC kit; Vector). Slides were washed and developed in 3,3'- diaminobenzidine (Sigma FASTDAB tablet; Sigma Chemical) under microscopic control, and counterstained with Mayer's hematoxylin. For each stained TMA the number of positive cells in each core was visually evaluated and the results expressed as a percentage of stained cells/ total number of cells. According to their immunoreactivity the tissues array cores were divided according to tumor differentiation, where the percentage of stained cells in the three tumor classes were scored as more than 5% and less than 25% of the cells stained, 26 to 50%, 51 to 75% or, 75 to 100%. -

Agricultural University of Athens
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΩΝ ΖΩΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΖΩΙΚΗΣ ΠΑΡΑΓΩΓΗΣ ΕΡΓΑΣΤΗΡΙΟ ΓΕΝΙΚΗΣ ΚΑΙ ΕΙΔΙΚΗΣ ΖΩΟΤΕΧΝΙΑΣ ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ ΑΘΗΝΑ 2020 ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Genome-wide association analysis and gene network analysis for (re)production traits in commercial broilers ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ Τριμελής Επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Επταμελής εξεταστική επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Πηνελόπη Μπεμπέλη (Καθ. ΓΠΑ) Δημήτριος Βλαχάκης (Επ. Καθ. ΓΠΑ) Ευάγγελος Ζωίδης (Επ.Καθ. ΓΠΑ) Γεώργιος Θεοδώρου (Επ.Καθ. ΓΠΑ) 2 Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Περίληψη Σκοπός της παρούσας διδακτορικής διατριβής ήταν ο εντοπισμός γενετικών δεικτών και υποψηφίων γονιδίων που εμπλέκονται στο γενετικό έλεγχο δύο τυπικών πολυγονιδιακών ιδιοτήτων σε κρεοπαραγωγικά ορνίθια. Μία ιδιότητα σχετίζεται με την ανάπτυξη (σωματικό βάρος στις 35 ημέρες, ΣΒ) και η άλλη με την αναπαραγωγική -

Plant & Animal Genome V
January 9-13, 2016 PLANT & ANIMAL GENOME XXIV Town & Country Hotel THE INTERNATIONAL CONFERENCE San Diego, CA ON THE STATUS OF PLANT & ANIMAL GENOME RESEARCH FINAL PROGRAM & EXHIBIT GUIDE Organizing Committee Chairman: Stephen R. Heller, NIST (USA) Ī PLANT COORGANIZERS Juan F. Medrano, University of California, Dave Clements, Johns Hopkins University, Davis, USA USA Huaijun Zhou, University of California, Catherine Feuillet, Bayer CropScience, USA Davis, USA J. Perry Gustafson, University of Missouri, (Retired ), USA Ī ABSTRACT & WEBSITE Jerome P. Miksche, Emeritus Director, COORDINATORS USDA, Plant Genome Program, USA David Grant, USDA/ARS/CICGR, USA Graham Moore, John Innes Centre, UK Gerard Lazo, USDA/ARS/WRRC, USA Susan R. Wessler, University of California, Victoria Carollo Blake, USA Riverside, USA Rod A. Wing, University of Arizona, USA; Ī TRAVEL GRANTS COORDINATOR International Rice Research Institute, Tom Blake, Professor Emeritus, Philippines Montana State University, USA Ī ANIMAL COORGANIZERS Ī SPECIAL DUTY COORDINATORS Daniel Ciobanu, University of Nebraska – Hans Cheng, USDA/ARS, USA Lincoln, USA Max Rothschild, Iowa State University, USA Kwan-Suk Kim, Chungbuk National University, South Korea Sponsors and Supporters ORGANIZER Scherago International Ī USDA, Agricultural Research Service 111 Town Square Place Ī USDA, National Agricultural Library Suite 1208 USDA, National Institute of Food and Agriculture Ī Jersey City, NJ 07310 Ī John Innes Centre Phone: (201) 653-4777 Fax: (201) 653-5705 Cover artwork provided by Applied Biosystems. Originally developed for the company’s “Genetic Harvest” Agriculture Seminars, this image Email: [email protected] represents the importance of molecular genetic approaches in plant and animal research. Website: www.intlpag.org About Frasergen Frasergen is an innovative leader in cutting-edge bioinformatics and overseas in high-throughput genome sequencing, big genome data genomics. -

Mechanism and in Vitro Pharmacology of TAK1 Inhibition by 5Z-7-Oxozeaenol Jiaquan Wu 1, Francoise Powell 2, Nicholas A
Mechanism and In Vitro Pharmacology of TAK1 Inhibition by 5Z-7-Oxozeaenol Jiaquan Wu 1, Francoise Powell 2, Nicholas A. Larsen 1, Zhongwu Lai 2, Kate F. Byth 2, Jon Read 3, Rong-Fang Gu 4, Mark Roth 2, Dorin Toader 2, Jamal Carlos Saeh 2, Huawei Chen 2 1Discovery Sciences, AstraZeneca R&D Boston, Waltham, MA 02451 2Oncology Innovative Medicine Unit, AstraZeneca R&D Boston, Waltham, MA 02451 3Discovery Sciences, AstraZeneca R&D Alderley Park, Cheshire, England SK10 4TG 4Infection Innovative Medicine Unit, AstraZeneca R&D Boston, Waltham, MA 02451 Supplemental Table 1. Growth inhibition of mixed liquid tumour cell lines by TAK1 inhibitor 5Z -7-Oxozeaenol. GI50 is the concentration of 5Z -7-Oxozeaenol needed to inhibit the cell growth by 50% comparing to untreated controls. TGI is the concentration required to completely inhibit the growth of treated cells. Cell line ID Liquid Tumour Type GI50 (µM) pGI50 TGI(µM) KRAS NRAS WSU -DLCL2 DLBCL 0.02 1.70 0.035 ND ND IM -9 MM 0.02 1.69 0.137 WT MUT OCI -AML2 AML 0.03 1.52 0.074 WT MUT JeKo -1 Mantle Cell Lymphoma 0.08 1.11 0.482 ND ND OCI -AML5 AML 0.09 1.07 0.548 ND MUT KG -1 AML 0.17 0.78 0.391 WT MUT OCI -Ly19 DLBCL 0.23 0.63 0.669 ND MUT THP -1 AML 0.28 0.55 2.037 WT MUT MOLM -16 AML 0.40 0.39 1.553 ND ND JVM -3 Chronic B Cell 0.42 0.38 1.598 WT MUT CMK AML 0.51 0.29 1.757 WT WT AMO -1 MM 0.57 0.24 ND ND NOMO -1 AML 0.57 0.24 5.683 MUT WT Ramos Burkitt's Lymphoma 0.73 0.14 2.077 ND ND MEC -1 B-CLL 0.98 0.01 2.676 ND ND U-937 Histiocytic Lym phoma 1.03 -0.01 2.816 WT WT Jurkat T Cell Leukemia -

Exosomal Signaling During Hypoxia Mediates Microvascular Endothelial Cell Migration and Vasculogenesis
Exosomal Signaling during Hypoxia Mediates Microvascular Endothelial Cell Migration and Vasculogenesis Carlos Salomon1*, Jennifer Ryan1, Luis Sobrevia1,2, Miharu Kobayashi1, Keith Ashman1, Murray Mitchell1, Gregory E. Rice1 1 University of Queensland Centre for Clinical Research, Herston, Queensland, Australia, 2 Cellular and Molecular Physiology Laboratory (CMPL), Division of Obstetrics and Gynaecology, School of Medicine, Faculty of Medicine, Pontificia Universidad Cato´lica de Chile, Santiago, Chile Abstract Vasculogenesis and angiogenesis are critical processes in fetal circulation and placental vasculature development. Placental mesenchymal stem cells (pMSC) are known to release paracrine factors (some of which are contained within exosomes) that promote angiogenesis and cell migration. The aims of this study were: to determine the effects of oxygen tension on the release of exosomes from pMSC; and to establish the effects of pMSC-derived exosomes on the migration and angiogenic tube formation of placental microvascular endothelial cells (hPMEC). pMSC were isolated from placental villi (8–12 weeks of gestation, n = 6) and cultured under an atmosphere of 1%, 3% or 8% O2. Cell-conditioned media were collected and exosomes (exo-pMSC) isolated by differential and buoyant density centrifugation. The dose effect (5–20 mg exosomal protein/ml) of pMSC-derived exosomes on hPMEC migration and tube formation were established using a real-time, live-cell imaging system (IncucyteTM). The exosome pellet was resuspended in PBS and protein content was established by mass spectrometry (MS). Protein function and canonical pathways were identified using the PANTHER program and Ingenuity Pathway Analysis, respectively. Exo-pMSC were identified, by electron microscopy, as spherical vesicles, with a typical cup- shape and diameters around of 100 nm and positive for exosome markers: CD63, CD9 and CD81. -

Zinc Signaling in Physiology and Pathogenesis
International Journal of Molecular Sciences Books Zinc Signaling in Physiology and Pathogenesis Edited by Toshiyuki Fukada and Taiho Kambe Printed Edition of the Special Issue Published in IJMS www.mdpi.com/journal/ijms MDPI Zinc Signaling in Physiology and Pathogenesis Special Issue Editors Toshiyuki Fukada Books Taiho Kambe MDPI • Basel • Beijing • Wuhan • Barcelona • Belgrade MDPI Special Issue Editors ToshiyukiFukada TokushimaBunri University Japan TaihoKambe Kyoto University Japan Editorial Office MDPI St. Alban-Anlage 66 Basel, Switzerland This edition is a reprint of the Special Issue published online in the open access journal IJMS (ISSN 1422-0067) from 2017-2018(available at: http:/ /www.mdpi.com/journal/ijms/special...issues/ zinc...signaling). Books For citation purposes, cite each article independently as indicated on the article page online and as indicated below: Lastname, F.M.; Lastname, F.M. Article title. Journal Name Year,A rticle number, page range. First Edition 2018 ISBN 978-3-03842-821-3 (Pbk) ISBN 978-3-03842-822-0 (PDF) Cover photo courtesy ofKoh-ei Toyoshimaand TakashiTsuji. It depicts mouse epidermal hair follicles which express zinc transporter ZIPlO (Bum-Ho Bin et al. PNAS 114, 12243-12248, 2017) Articles in this volume are Open Access and distributed under the Creative Commons Attribution (CC BY) license, which allows users to download, copy and build upon published articles even for commercial purposes, as long as the author and publisher are properly credited, which ensures maximum dissemination and a wider impact of our publications. The book taken as a whole is©2018 MDPI, Basel, Switzerland, distributed under the terms and conditions of the Creative Commons license CC BY-NC-ND(http://creativecommons.org/licenses/by-nc-nd/ 4.0/).