Dmol Guide to Select a Dmol3 Task 1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Spoken Tutorial Project, IIT Bombay Brochure for Chemistry Department
Spoken Tutorial Project, IIT Bombay Brochure for Chemistry Department Name of FOSS Applications Employability GChemPaint GChemPaint is an editor for 2Dchem- GChemPaint is currently being developed ical structures with a multiple docu- as part of The Chemistry Development ment interface. Kit, and a Standard Widget Tool kit- based GChemPaint application is being developed, as part of Bioclipse. Jmol Jmol applet is used to explore the Jmol is a free, open source molecule viewer structure of molecules. Jmol applet is for students, educators, and researchers used to depict X-ray structures in chemistry and biochemistry. It is cross- platform, running on Windows, Mac OS X, and Linux/Unix systems. For PG Students LaTeX Document markup language and Value addition to academic Skills set. preparation system for Tex typesetting Essential for International paper presentation and scientific journals. For PG student for their project work Scilab Scientific Computation package for Value addition in technical problem numerical computations solving via use of computational methods for engineering problems, Applicable in Chemical, ECE, Electrical, Electronics, Civil, Mechanical, Mathematics etc. For PG student who are taking Physical Chemistry Avogadro Avogadro is a free and open source, Research and Development in Chemistry, advanced molecule editor and Pharmacist and University lecturers. visualizer designed for cross-platform use in computational chemistry, molecular modeling, material science, bioinformatics, etc. Spoken Tutorial Project, IIT Bombay Brochure for Commerce and Commerce IT Name of FOSS Applications / Employability LibreOffice – Writer, Calc, Writing letters, documents, creating spreadsheets, tables, Impress making presentations, desktop publishing LibreOffice – Base, Draw, Managing databases, Drawing, doing simple Mathematical Math operations For Commerce IT Students Drupal Drupal is a free and open source content management system (CMS). -
Biovia Materials Studio Visualizer Datasheet
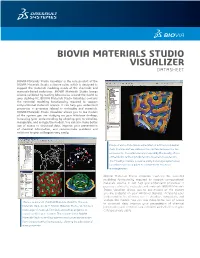
BIOVIA MATERIALS STUDIO VISUALIZER DATASHEET BIOVIA Materials Studio Visualizer is the core product of the BIOVIA Materials Studio software suite, which is designed to support the materials modeling needs of the chemicals and materials-based industries. BIOVIA Materials Studio brings science validated by leading laboratories around the world to your desktop PC. BIOVIA Materials Studio Visualizer contains the essential modeling functionality required to support computational materials science. It can help you understand properties or processes related to molecules and materials. BIOVIA Materials Studio Visualizer allows you to see models of the system you are studying on your Windows desktop, increasing your understanding by allowing you to visualize, manipulate, and analyze the models. You can also make better use of access to structural data, improve your presentation of chemical information, and communicate problems and solutions to your colleagues very easily. Image of early-stage phase segregation in a diblock copolymer melt. The blue surface indicates the interface between the two components. The volume is colormapped by the density of one of the blocks, red being high density, blue being low-density. The MesoDyn module is used to study these large systems over long-times such as required to observe these structural rearrangements. BIOVIA Materials Studio Visualizer contains the essential modeling functionality required to support computational materials science. It can help you understand properties or processes related to molecules and materials. BIOVIA Materials Studio Visualizer allows you to see models of the system you are studying on your Windows desktop, increasing your understanding by allowing you to visualize, manipulate, and analyze the models. -

Download Author Version (PDF)
PCCP Accepted Manuscript This is an Accepted Manuscript, which has been through the Royal Society of Chemistry peer review process and has been accepted for publication. Accepted Manuscripts are published online shortly after acceptance, before technical editing, formatting and proof reading. Using this free service, authors can make their results available to the community, in citable form, before we publish the edited article. We will replace this Accepted Manuscript with the edited and formatted Advance Article as soon as it is available. You can find more information about Accepted Manuscripts in the Information for Authors. Please note that technical editing may introduce minor changes to the text and/or graphics, which may alter content. The journal’s standard Terms & Conditions and the Ethical guidelines still apply. In no event shall the Royal Society of Chemistry be held responsible for any errors or omissions in this Accepted Manuscript or any consequences arising from the use of any information it contains. www.rsc.org/pccp Page 1 of 11 PhysicalPlease Chemistry do not adjust Chemical margins Physics PCCP PAPER Effect of nanosize on surface properties of NiO nanoparticles for adsorption of Quinolin-65 ab a a Received 00th January 20xx, Nedal N. Marei, Nashaat N. Nassar* and Gerardo Vitale Accepted 00th January 20xx Using Quinolin-65 (Q-65) as a model-adsorbing compound for polar heavy hydrocarbons, the nanosize effect of NiO Manuscript DOI: 10.1039/x0xx00000x nanoparticles on adsorption of Q-65 was investigated. Different-sized NiO nanoparticles with sizes between 5 and 80 nm were prepared by controlled thermal dehydroxylation of Ni(OH)2. -

Supporting Information
Electronic Supplementary Material (ESI) for Chemical Science. This journal is © The Royal Society of Chemistry 2015 Supporting Information A Single Crystalline Porphyrinic Titanium Metal−Organic Framework Shuai Yuana†, Tian-Fu Liua†, Dawei Fenga, Jian Tiana, Kecheng Wanga, Junsheng Qina, a a a a b Qiang Zhang , Ying-Pin Chen , Mathieu Bosch , Lanfang Zou , Simon J. Teat, Scott J. c a Dalgarno and Hong-Cai Zhou * a Department of Chemistry, Texas A&M University, College Station, Texas 77842-3012, USA b Advanced Light Source, Lawrence Berkeley National Laboratory Berkeley, CA 94720, USA c Institute of Chemical Sciences, Heriot-Watt University Riccarton, Edinburgh EH14 4AS, U.K. † Equal contribution to this work *To whom correspondence should be addressed. Email: [email protected] Tel: +1 (979) 845-4034; Fax: +1 (979) 845-1595 S1 Contents S1. Ligand Synthesis..............................................................................................................3 S2. Syntheses of PCN-22.......................................................................................................5 S3. X-ray Crystallography .....................................................................................................6 S4. Topological Analyses ......................................................................................................9 S5. N2 Sorption Isotherm .....................................................................................................10 S6. Simulation of the Accessible Surface Area ...................................................................12 -

PDF Hosted at the Radboud Repository of the Radboud University Nijmegen
PDF hosted at the Radboud Repository of the Radboud University Nijmegen The following full text is a publisher's version. For additional information about this publication click this link. http://hdl.handle.net/2066/19078 Please be advised that this information was generated on 2021-09-23 and may be subject to change. Computational Chemistry Metho ds Applications to Racemate Resolution and Radical Cation Chemistry ISBN Computational Chemistry Metho ds Applications to Racemate Resolution and Radical Cation Chemistry een wetenschapp elijkeproeve op het gebied van de Natuurwetenschapp en Wiskunde en Informatica Pro efschrift ter verkrijging van de graad van do ctor aan de KatholiekeUniversiteit Nijmegen volgens b esluit van het College van Decanen in het op enbaar te verdedigen op dinsdag januari des namiddags om uur precies do or Gijsb ert Schaftenaar geb oren op augustus te Harderwijk Promotores Prof dr ir A van der Avoird Prof dr E Vlieg Copromotor Prof dr RJ Meier Leden manuscriptcommissie Prof dr G Vriend Prof dr RA de Gro ot Dr ir PES Wormer The research rep orted in this thesis was nancially supp orted by the Dutch Or ganization for the Advancement of Science NWO and DSM Contents Preface Intro duction Intro duction Chirality Metho ds for obtaining pure enantiomers Racemate Resolution via diastereomeric salt formation Rationalization of diastereomeric salt formation Computational metho ds for mo deling the lattice energy Molecular Mechanics Quantum Chemical -

Reactive Molecular Dynamics Study of the Thermal Decomposition of Phenolic Resins
Article Reactive Molecular Dynamics Study of the Thermal Decomposition of Phenolic Resins Marcus Purse 1, Grace Edmund 1, Stephen Hall 1, Brendan Howlin 1,* , Ian Hamerton 2 and Stephen Till 3 1 Department of Chemistry, Faculty of Engineering and Physical Sciences, University of Surrey, Guilford, Surrey GU2 7XH, UK; [email protected] (M.P.); [email protected] (G.E.); [email protected] (S.H.) 2 Bristol Composites Institute (ACCIS), Department of Aerospace Engineering, School of Civil, Aerospace, and Mechanical Engineering, University of Bristol, Bristol BS8 1TR, UK; [email protected] 3 Defence Science and Technology Laboratory, Porton Down, Salisbury SP4 0JQ, UK; [email protected] * Correspondence: [email protected]; Tel.: +44-1483-686-248 Received: 6 March 2019; Accepted: 23 March 2019; Published: 28 March 2019 Abstract: The thermal decomposition of polyphenolic resins was studied by reactive molecular dynamics (RMD) simulation at elevated temperatures. Atomistic models of the polyphenolic resins to be used in the RMD were constructed using an automatic method which calls routines from the software package Materials Studio. In order to validate the models, simulated densities and heat capacities were compared with experimental values. The most suitable combination of force field and thermostat for this system was the Forcite force field with the Nosé–Hoover thermostat, which gave values of heat capacity closest to those of the experimental values. Simulated densities approached a final density of 1.05–1.08 g/cm3 which compared favorably with the experimental values of 1.16–1.21 g/cm3 for phenol-formaldehyde resins. -

Pymol Modelling Workshop
PyMOL Modelling Workshop My website: http://pldserver1.biochem.queensu.ca/~rlc/work/teaching/BCHM442/ There you will find links to download the latest educational version of PyMOL as well as a link to my “Introduction to PyMOL”, which in turn has links to other people's PyMOL tutorials. Note also the PyMOL Wiki: http://pymolwiki.org. Structure files can be found by searching the Protein Data Bank (PDB) for structure: easy to remember website http://www.pdb.org. There is also the PDBe (PDB Europe, http://www.pdbe.org) that contains the same databank of structures, but with a different web interface for searching for structures and a different set of tools for analyzing structures. What is in a PDB file? Lots of information in the “header” (the section of the file preceding the actual atomic coordinates) as well as the coordinates for the atoms. When assessing a structure, one needs to take account of the resolution and R-factor, error estimates and missing residues. There is information about the sequence that was used to determine the structure with a sequence database reference. There is also information about the biological unit. In the case of crystal structures the biological unit may need to be generated by applying crystallographic symmetry operators, although there are web sites that also try to provide that information. (e.g. http://www.ebi.ac.uk/msd-srv/pisa/). Warning: One cannot blindly trust a crystal structure to be providing you with a completely accurate picture of reality. Reading the paper that describes the structure is a good start! Outline of PyMOL usage PyMOL is a very powerful, scriptable (customizable) tool for making publication-quality figures and performing analyses on structures. -

Density Functional Theory (DFT)
Herramientas mecano-cuánticas basadas en DFT para el estudio de moléculas y materiales en Materials Studio 7.0 Javier Ramos Biophysics of Macromolecular Systems group (BIOPHYM) Departamento de Física Macromolecular Instituto de Estructura de la Materia – CSIC [email protected] Webinar, 26 de Junio 2014 Anteriores webinars Como conseguir los videos y las presentaciones de anteriores webminars: Linkedin: Grupo de Química Computacional http://www.linkedin.com/groups/Química-computacional-7487634 Índice Density Functional Theory (DFT) The Jacob’s ladder DFT modules in Maretials Studio DMOL3, CASTEP and ONETEP XC functionals Basis functions Interfaces in Materials Studio Tasks Properties Example: n-butane conformations Density Functional Theory (DFT) DFT is built around the premise that the energy of an electronic system can be defined in terms of its electron probability density (ρ). (Hohenberg-Kohn Theorem) E 0 [ 0 ] Te [ 0 ] E ne [ 0 ] E ee [ 0 ] (easy) Kinetic Energy for ????? noninteracting (r )v (r ) dr electrons(easy) 1 E[]()()[]1 r r d r d r E e e2 1 2 1 2 X C r12 Classic Term(Coulomb) Non-classic Kohn-Sham orbitals Exchange & By minimizing the total energy functional applying the variational principle it is Correlation possible to get the SCF equations (Kohn-Sham) The Jacob’s Ladder Accurate form of XC potential Meta GGA Empirical (Fitting to Non-Empirical Generalized Gradient Approx. atomic properties) (physics rules) Local Density Approximation DFT modules in Materials Studio DMol3: Combine computational speed with the accuracy of quantum mechanical methods to predict materials properties reliably and quickly CASTEP: CASTEP offers simulation capabilities not found elsewhere, such as accurate prediction of phonon spectra, dielectric constants, and optical properties. -

Atomistic Molecular Dynamics Simulation of Graphene- Isoprene Nanocomposites
Atomistic molecular dynamics simulation of graphene- isoprene nanocomposites P Chanlert1, J Wong-Ekkabut2, W Liewrian3,4,5 and T Sutthibutpong3,4,5* 1 Program of Physics, Faculty of Science and Technology, Songkhla Rajabhat University, Songkhla, 90000, Thailand 2 Department of Physics, Faculty of Science, Kasetsart University, Bangkok 10900, Thailand 3 Theoretical and Computational Physics Group, Department of Physics, Faculty of Science, King Mongkut’s University of Technology Thonburi (KMUTT), Bangkok 10140, Thailand 4 Theoretical and Computational Science Center (TaCS), Science Laboratory Building, Faculty of Science, King Mongkut’s University of Technology Thonburi (KMUTT), Bangkok 10140, Thailand 5 Thailand Center of Excellence in Physics (ThEP Center), Commission on Higher Education, Bangkok 10400, Thailand *email: [email protected] Abstract. A series of atomistic molecular dynamics simulations were performed for 4-mer isoprene molecules confined between graphene sheets with varied graphene separations in order to observe the finite size effects arose from the van der Waals interactions between the isoprene oligomer, forming high density shells at the graphene interfaces. For the small confinement regions, 1.24 nm and 2.21 nm, local density of isoprene at the graphene interface becomes higher than 10 times the bulk density. These results provided the further insights towards the rational design of nanostructures based on graphene sheets and natural rubber polymer. 1. Introduction Composite material is the combination between two materials or more where chemical reaction between these materials is absent. In composites, one with higher quantity is called matrix while those with lesser quantity are called fillers. In general, adding fillers into matrix material enhances both physical and mechanical properties in some ways. -

Prediction of Solubility of Amino Acids Based on Cosmo Calculaition
PREDICTION OF SOLUBILITY OF AMINO ACIDS BASED ON COSMO CALCULAITION by Kaiyu Li A dissertation submitted to Johns Hopkins University in conformity with the requirement for the degree of Master of Science in Engineering Baltimore, Maryland October 2019 Abstract In order to maximize the concentration of amino acids in the culture, we need to obtain solubility of amino acid as a function of concentration of other components in the solution. This function can be obtained by calculating the activity coefficient along with solubility model. The activity coefficient of the amino acid can be calculated by UNIFAC. Due to the wide range of applications of UNIFAC, the prediction of the activity coefficient of amino acids is not very accurate. So we want to fit the parameters specific to amino acids based on the UNIFAC framework and existing solubility data. Due to the lack of solubility of amino acids in the multi-system, some interaction parameters are not available. COSMO is a widely used way to describe pairwise interactions in the solutions in the chemical industry. After suitable assumptions COSMO can calculate the pairwise interactions in the solutions, and largely reduce the complexion of quantum chemical calculation. In this paper, a method combining quantum chemistry and COSMO calculation is designed to accurately predict the solubility of amino acids in multi-component solutions in the ii absence of parameters, as a supplement to experimental data. Primary Reader and Advisor: Marc D. Donohue Secondary Reader: Gregory Aranovich iii Contents -

Molecular Dynamics Simulations of Molecular Diffusion Equilibrium and Breakdown Mechanism of Oil-Impregnated Pressboard with Water Impurity
polymers Article Molecular Dynamics Simulations of Molecular Diffusion Equilibrium and Breakdown Mechanism of Oil-Impregnated Pressboard with Water Impurity Yi Guan 1,2, Ming-He Chi 1,2, Wei-Feng Sun 1,2,*, Qing-Guo Chen 1,2 and Xin-Lao Wei 1,2 1 Heilongjiang Provincial Key Laboratory of Dielectric Engineering, School of Electrical and Electronic Engineering, Harbin University of Science and Technology, Harbin 150080, China; [email protected] (Y.G.); [email protected] (M.-H.C.); [email protected] (Q.-G.C.); [email protected] (X.-L.W.) 2 Key Laboratory of Engineering Dielectrics and Its Application, Ministry of Education, Harbin University of Science and Technology, Harbin 150080, China * Correspondence: [email protected]; Tel.: +86-15846592798 Received: 30 October 2018; Accepted: 15 November 2018; Published: 16 November 2018 Abstract: The water molecule migration and aggregation behaviors in oil-impregnated pressboard are investigated by molecular dynamics simulations in combination with Monte Carlo molecular simulation technique. The free energy and phase diagram of H2O-dodecylbenzene (DDB) and H2O-cellulose mixtures are calculated by Monte Carlo technique combined with the modified Flory-Huggins model, demonstrating that H2O molecules can hardly dissolved with infinitesimal content in cellulose system at temperature lower than 650 K, based on which the oil/cellulose layered structure with water impurity representing three-phase coexistence in oil-impregnated pressboard are modeled and performed for molecular dynamics. The molecular dynamics of H2O/DDB/cellulose three-phase mixture simulating oil-paper insulating system with H2O impurity indicates that DDB molecules can thermally intrude into the cellulose-water interface so as to separate the water phase and cellulose fiber. -

What's New in Biovia Materials Studio 2020
WHAT’S NEW IN BIOVIA MATERIALS STUDIO 2020 Datasheet BIOVIA Materials Studio 2020 is the latest release of BIOVIA’s predictive science tools for chemistry and materials science research. Materials Studio empowers researchers to understand the relationships between a material’s mo- lecular or crystal structure and its properties in order to make more informed decisions about materials research and development. More often than not materials performance is influenced by phenomena at multiple scales. Scientists using Materials Studio 2020 have an extensive suite of world class solvers and parameter sets operating from atoms to microscale for simulating more materials and more properties than ever before. Furthermore, the predicted properties can now be used in multi-physics modeling and in systems modeling software such as SIMULIA Abaqus and CATIA Dymola to predict macroscopic behaviors. In this way multiscale simulations can be used to solve some of the toughest pro- blems in materials design and product optimization. BETTER MATERIALS - BETTER BATTERIES Safe, fast charging batteries with high energy density and long life are urgently needed for a host of applications - not least for the electrification of all modes of transportation as an alternative to fossil fuel energy sources. Battery design relies on a complex interplay between thermal, mechanical and chemical processes from the smallest scales of the material (electronic structure) through to the geometry of the battery cell and pack design. Improvements to the component materials used in batteries and capacitors are fundamental to providing the advances in performance needed. Materials Studio provides new functionality to enable the simula- tion of key materials parameters for both liquid electrolytes and electrode components.