Transcriptional Profiling of Stroma from Inflamed and Resting Lymph Nodes
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Propranolol-Mediated Attenuation of MMP-9 Excretion in Infants with Hemangiomas
Supplementary Online Content Thaivalappil S, Bauman N, Saieg A, Movius E, Brown KJ, Preciado D. Propranolol-mediated attenuation of MMP-9 excretion in infants with hemangiomas. JAMA Otolaryngol Head Neck Surg. doi:10.1001/jamaoto.2013.4773 eTable. List of All of the Proteins Identified by Proteomics This supplementary material has been provided by the authors to give readers additional information about their work. © 2013 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 10/01/2021 eTable. List of All of the Proteins Identified by Proteomics Protein Name Prop 12 mo/4 Pred 12 mo/4 Δ Prop to Pred mo mo Myeloperoxidase OS=Homo sapiens GN=MPO 26.00 143.00 ‐117.00 Lactotransferrin OS=Homo sapiens GN=LTF 114.00 205.50 ‐91.50 Matrix metalloproteinase‐9 OS=Homo sapiens GN=MMP9 5.00 36.00 ‐31.00 Neutrophil elastase OS=Homo sapiens GN=ELANE 24.00 48.00 ‐24.00 Bleomycin hydrolase OS=Homo sapiens GN=BLMH 3.00 25.00 ‐22.00 CAP7_HUMAN Azurocidin OS=Homo sapiens GN=AZU1 PE=1 SV=3 4.00 26.00 ‐22.00 S10A8_HUMAN Protein S100‐A8 OS=Homo sapiens GN=S100A8 PE=1 14.67 30.50 ‐15.83 SV=1 IL1F9_HUMAN Interleukin‐1 family member 9 OS=Homo sapiens 1.00 15.00 ‐14.00 GN=IL1F9 PE=1 SV=1 MUC5B_HUMAN Mucin‐5B OS=Homo sapiens GN=MUC5B PE=1 SV=3 2.00 14.00 ‐12.00 MUC4_HUMAN Mucin‐4 OS=Homo sapiens GN=MUC4 PE=1 SV=3 1.00 12.00 ‐11.00 HRG_HUMAN Histidine‐rich glycoprotein OS=Homo sapiens GN=HRG 1.00 12.00 ‐11.00 PE=1 SV=1 TKT_HUMAN Transketolase OS=Homo sapiens GN=TKT PE=1 SV=3 17.00 28.00 ‐11.00 CATG_HUMAN Cathepsin G OS=Homo -

Supplementary Table 3 Complete List of RNA-Sequencing Analysis of Gene Expression Changed by ≥ Tenfold Between Xenograft and Cells Cultured in 10%O2
Supplementary Table 3 Complete list of RNA-Sequencing analysis of gene expression changed by ≥ tenfold between xenograft and cells cultured in 10%O2 Expr Log2 Ratio Symbol Entrez Gene Name (culture/xenograft) -7.182 PGM5 phosphoglucomutase 5 -6.883 GPBAR1 G protein-coupled bile acid receptor 1 -6.683 CPVL carboxypeptidase, vitellogenic like -6.398 MTMR9LP myotubularin related protein 9-like, pseudogene -6.131 SCN7A sodium voltage-gated channel alpha subunit 7 -6.115 POPDC2 popeye domain containing 2 -6.014 LGI1 leucine rich glioma inactivated 1 -5.86 SCN1A sodium voltage-gated channel alpha subunit 1 -5.713 C6 complement C6 -5.365 ANGPTL1 angiopoietin like 1 -5.327 TNN tenascin N -5.228 DHRS2 dehydrogenase/reductase 2 leucine rich repeat and fibronectin type III domain -5.115 LRFN2 containing 2 -5.076 FOXO6 forkhead box O6 -5.035 ETNPPL ethanolamine-phosphate phospho-lyase -4.993 MYO15A myosin XVA -4.972 IGF1 insulin like growth factor 1 -4.956 DLG2 discs large MAGUK scaffold protein 2 -4.86 SCML4 sex comb on midleg like 4 (Drosophila) Src homology 2 domain containing transforming -4.816 SHD protein D -4.764 PLP1 proteolipid protein 1 -4.764 TSPAN32 tetraspanin 32 -4.713 N4BP3 NEDD4 binding protein 3 -4.705 MYOC myocilin -4.646 CLEC3B C-type lectin domain family 3 member B -4.646 C7 complement C7 -4.62 TGM2 transglutaminase 2 -4.562 COL9A1 collagen type IX alpha 1 chain -4.55 SOSTDC1 sclerostin domain containing 1 -4.55 OGN osteoglycin -4.505 DAPL1 death associated protein like 1 -4.491 C10orf105 chromosome 10 open reading frame 105 -4.491 -

A Clinicopathological and Molecular Genetic Analysis of Low-Grade Glioma in Adults
A CLINICOPATHOLOGICAL AND MOLECULAR GENETIC ANALYSIS OF LOW-GRADE GLIOMA IN ADULTS Presented by ANUSHREE SINGH MSc A thesis submitted in partial fulfilment of the requirements of the University of Wolverhampton for the degree of Doctor of Philosophy Brain Tumour Research Centre Research Institute in Healthcare Sciences Faculty of Science and Engineering University of Wolverhampton November 2014 i DECLARATION This work or any part thereof has not previously been presented in any form to the University or to any other body whether for the purposes of assessment, publication or for any other purpose (unless otherwise indicated). Save for any express acknowledgments, references and/or bibliographies cited in the work, I confirm that the intellectual content of the work is the result of my own efforts and of no other person. The right of Anushree Singh to be identified as author of this work is asserted in accordance with ss.77 and 78 of the Copyright, Designs and Patents Act 1988. At this date copyright is owned by the author. Signature: Anushree Date: 30th November 2014 ii ABSTRACT The aim of the study was to identify molecular markers that can determine progression of low grade glioma. This was done using various approaches such as IDH1 and IDH2 mutation analysis, MGMT methylation analysis, copy number analysis using array comparative genomic hybridisation and identification of differentially expressed miRNAs using miRNA microarray analysis. IDH1 mutation was present at a frequency of 71% in low grade glioma and was identified as an independent marker for improved OS in a multivariate analysis, which confirms the previous findings in low grade glioma studies. -

BD Biosciences New RUO Reagents - November 2020
BD Biosciences New RUO reagents - November 2020 Reactivity Description Format Clone Size Cat. number Hu CD133 FITC W6B3C1 100µg 567029 Hu CD133 FITC W6B3C1 25µg 567033 Hu CD39 PE A1/CD39 100Tst 567156 Hu CD39 PE A1/CD39 25Tst 567157 Hu KIR2DL1/S1/S3/S5 PE HP-MA4 100Tst 567158 Hu KIR2DL1/S1/S3/S5 PE HP-MA4 25Tst 567159 Hu IL-22 Alexa Fluor® 647 MH22B2 100µg 567160 Hu IL-22 Alexa Fluor® 647 MH22B2 25µg 567161 Hu CD99 R718 TU12 50µg 751651 Hu CD161 R718 DX12 50µg 751652 Hu CD116 R718 HGMCSFR-M1 50µg 751653 Hu HLA-G R718 87G 50µg 751670 Hu CD27 R718 O323 50µg 751686 Hu CD80 (B7-1) R718 2D10.4 50µg 751737 Hu Integrin αvβ5 R718 ALULA 50µg 751738 Hu CD266 (Tweak-R) R718 ITEM-4 50µg 751739 Hu ErbB3 (HER-3) R718 SGP1 50µg 751799 Hu TCR Vβ5.1 R718 LC4 50µg 751816 Hu CD123 (IL-3Ra) R718 6H6 50µg 751844 Hu CD1a R718 SK9 50µg 751847 Hu CD20 R718 L27 50µg 751849 Hu Disial GD2 R718 14.G2A 50µg 751851 Reactivity Description Format Clone Size Cat. number Hu CD71 R718 L01.1 50µg 751853 Hu CD278 (ICOS) R718 DX29 50µg 751854 Hu B7-H4 R718 MIH43 50µg 751857 Hu CD53 R718 HI29 50µg 751858 Hu CD197 (CCR7) R718 2-L1-A 50µg 751859 Hu CD197 (CCR7) R718 3D12 50µg 751861 Hu CD31 R718 L133.1 50µg 751863 Hu EGF Receptor R718 EMAB-134 50µg 751864 Hu CD8b R718 2ST8.5H7 50µg 751867 Hu CD31 R718 MBC 78.2 50µg 751869 Hu CD162 R718 KPL-1 50µg 751873 Hu CD24 R718 ML5 50µg 751874 Hu CD159C (NKG2C) R718 134591 50µg 751876 Hu CD169 (Siglec-1) R718 7-239 50µg 751877 Hu CD16b R718 CLB-GRAN11.5 50µg 751880 Hu IgM R718 UCH-B1 50µg 751881 Hu CD275 R718 2D3/B7-H2 50µg 751883 Hu CD307e -

Profound Treg Perturbations Correlate with COVID-19 Severity
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.11.416180; this version posted December 15, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Profound Treg perturbations correlate with COVID-19 severity Silvia Galván-Peña1*, Juliette Leon1,2*, Kaitavjeet Chowdhary1, Daniel A. Michelson1, Brinda Vijaykumar1, Liang Yang1, Angela Magnuson1, Zachary Manickas-Hill3,4, Alicja Piechocka- Trocha3,4, Daniel P. Worrall3,4, Kathryn E. Hall3,5, Musie Ghebremichael3,4, Bruce D. Walker3,4,6, Jonathan Z. Li3,7, Xu G. Yu3,4, MGH COVID-19 Collection & Processing Team, Diane Mathis1 and Christophe Benoist1,8,+ 1Department of Immunology, Harvard Medical School, Boston, MA, USA 2 INSERM UMR 1163, University of Paris, Imagine Institute, Paris, France 3Massachusetts Consortium on Pathogen Readiness, Boston, MA, USA 4Ragon Institute of MGH, MIT and Harvard, Cambridge, MA, USA 5Department of Medicine, Massachusetts General Hospital, Boston, MA, USA 6Howard Hughes Medical Institute, Center for the AIDS Programme of Research in South Africa. 7Brigham and Women’s Hospital, Harvard Medical School, Boston, MA, USA 8Lead contact * Equal contribution +Address correspondence to: Christophe Benoist Department of Immunology Harvard Medical School 77 Avenue Louis Pasteur, Boston, MA 02115 e-mail: [email protected] Phone: (617) 432-7741 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.12.11.416180; this version posted December 15, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. -

Regulation Of
Sede Amministrativa: Università degli Studi di Padova Dipartimento di Istologia, Microbiologia e Biotecnologie Mediche SCUOLA DI DOTTORATO DI RICERCA IN BIOSCIENZE INDIRIZZO GENETICA CICLO XXIII EXPRESSION AND FUNCTIONAL ROLE OF EMILIN-3, A PECULIAR MEMBER OF THE EMILIN/MULTIMERIN FAMILY Direttore della Scuola : Ch.mo Prof. Giuseppe Zanotti Coordinatore d’indirizzo: Ch.mo Prof. Paolo Bonaldo Supervisore :Ch.mo Prof. Paolo Bonaldo Dottorando : Alvise Schiavinato 1 2 THESIS CONTENTS ABSTRACT ________________________________________________________________________________________ 5 ABSTRACT (ITALIANO)__________________________________________________________________________ 6 INTRODUCTION__________________________________________________________________________________ 7 THE EXTRACELLULAR MATRIX_____________________________________________________________________ 7 REGULATION OF TGF−Β SIGNALING BY THE EXTRACELLULAR MATRIX______________________________ 9 EMILIN-1________________________________________________________________________________________11 THE EMILIN/MULTIMERIN FAMILY ______________________________________________________________12 THE EMILIN/MULTIMERIN FAMILY IN ZEBRAFISH ________________________________________________13 STRUCTURE AND FUNCTION OF THE NOTOCHORD _________________________________________________14 AIM OF THE RESEARCH ______________________________________________________________________16 MATERIALS AND METHODS__________________________________________________________________ 17 ANIMALS ________________________________________________________________________________________17 -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -
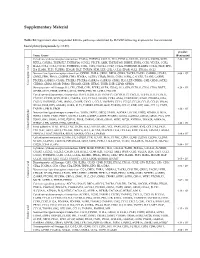
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

TMHS Is an Integral Component of the Mechanotransduction Machinery of Cochlear Hair Cells
TMHS Is an Integral Component of the Mechanotransduction Machinery of Cochlear Hair Cells Wei Xiong,1 Nicolas Grillet,1 Heather M. Elledge,1 Thomas F.J. Wagner,1 Bo Zhao,1 Kenneth R. Johnson,2 Piotr Kazmierczak,1 and Ulrich Mu¨ller1,* 1The Dorris Neuroscience Center, Department of Cell Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA 2The Jackson Laboratory, Bar Harbor, ME 04609, USA *Correspondence: [email protected] http://dx.doi.org/10.1016/j.cell.2012.10.041 SUMMARY deflection of the stereociliary bundles, which directly control the activity of the mechanotransduction channels in stereocilia. Hair cells are mechanosensors for the perception of It is thought that tip links, fine extracellular filaments that connect sound, acceleration, and fluid motion. Mechano- the tips of neighboring stereocilia, transmit tension force onto transduction channels in hair cells are gated by tip the transduction channels (Gillespie and Mu¨ ller, 2009). links, which connect the stereocilia of a hair cell in In recent years, significant progress has been made in the direction of their mechanical sensitivity. The the identification of components of the mechanotransduction molecular constituents of the mechanotransduction machinery of hair cells (Figure 1A). These studies have shown that tip links are formed by CDH23 homodimers that interact channels of hair cells are not known. Here, we show with PCDH15 homodimers to form the upper and lower parts that mechanotransduction is impaired in mice lack- of tip links (Ahmed et al., 2006; Kazmierczak et al., 2007; ing the tetraspan TMHS. TMHS binds to the tip-link Siemens et al., 2004; So¨ llner et al., 2004). -

Local Genetic and Environmental Factors in Asthma Disease Pathogenesis: Chronicity and Persistence Mechanisms
Eur Respir J 2007; 29: 793–803 DOI: 10.1183/09031936.00087506 CopyrightßERS Journals Ltd 2007 SERIES ‘‘GENETICS OF ASTHMA AND COPD IN THE POSTGENOME ERA’’ Edited by E. von Mutius, M. Kabesch and F. Kauffmann Number 4 in this Series Local genetic and environmental factors in asthma disease pathogenesis: chronicity and persistence mechanisms S.T. Holgate*, D.E. Davies*, R.M. Powell*, P.H. Howarth*, H.M. Haitchi# and J.W. Holloway" ABSTRACT: While asthma is an inflammatory disorder of the airways usually associated with AFFILIATIONS atopy, an important additional component is involvement of the epithelium and underlying *Allergy and Inflammation Research, Division of Infection, Inflammation mesenchyme acting as a trophic unit (EMTU). and Repair, In addition to allergens, a wide range of environmental factors interact with the EMTU, such as #IIR Division and virus infections, environmental tobacco smoke and pollutants, to initiate tissue damage and "Division of Human Genetics, School aberrant repair responses that are translated into remodelling of the airways. While candidate of Medicine, Southampton General Hospital, Southampton, UK. gene association studies have revealed polymorphic variants that influence asthmatic inflamma- tion, positional cloning of previously unknown genes is identifying a high proportion of novel CORRESPONDENCE genes in the EMTU. S.T. Holgate Dipeptidyl peptidase (DPP) 10 and disintegrin and metalloproteinase (ADAM)33 are newly Allergy and Inflammation Research MP810 identified genes strongly associated with asthma that are preferentially expressed in the airway Level D epithelium and underlying mesenchyme, respectively. Centre Block Also of increasing importance is the recognition that genes associated with asthma and atopy Southampton General Hospital have important interactions with the environment through epigenetic mechanisms that influence Southampton SO16 6YD UK their expression. -

Gene List HTG Edgeseq Immuno-Oncology Assay
Gene List HTG EdgeSeq Immuno-Oncology Assay Adhesion ADGRE5 CLEC4A CLEC7A IBSP ICAM4 ITGA5 ITGB1 L1CAM MBL2 SELE ALCAM CLEC4C DST ICAM1 ITGA1 ITGA6 ITGB2 LGALS1 MUC1 SVIL CDH1 CLEC5A EPCAM ICAM2 ITGA2 ITGAL ITGB3 LGALS3 NCAM1 THBS1 CDH5 CLEC6A FN1 ICAM3 ITGA4 ITGAM ITGB4 LGALS9 PVR THY1 Apoptosis APAF1 BCL2 BID CARD11 CASP10 CASP8 FADD NOD1 SSX1 TP53 TRAF3 BCL10 BCL2L1 BIRC5 CASP1 CASP3 DDX58 NLRP3 NOD2 TIMP1 TRAF2 TRAF6 B-Cell Function BLNK BTLA CD22 CD79A FAS FCER2 IKBKG PAX5 SLAMF1 SLAMF7 SPN BTK CD19 CD24 EBF4 FASLG IKBKB MS4A1 RAG1 SLAMF6 SPI1 Cell Cycle ABL1 ATF1 ATM BATF CCND1 CDK1 CDKN1B NCL RELA SSX1 TBX21 TP53 ABL2 ATF2 AXL BAX CCND3 CDKN1A EGR1 REL RELB TBK1 TIMP1 TTK Cell Signaling ADORA2A DUSP4 HES1 IGF2R LYN MAPK1 MUC1 NOTCH1 RIPK2 SMAD3 STAT5B AKT3 DUSP6 HES5 IKZF1 MAF MAPK11 MYC PIK3CD RNF4 SOCS1 STAT6 BCL6 ELK1 HEY1 IKZF2 MAP2K1 MAPK14 NFATC1 PIK3CG RORC SOCS3 SYK CEBPB EP300 HEY2 IKZF3 MAP2K2 MAPK3 NFATC3 POU2F2 RUNX1 SPINK5 TAL1 CIITA ETS1 HEYL JAK1 MAP2K4 MAPK8 NFATC4 PRKCD RUNX3 STAT1 TCF7 CREB1 FLT3 HMGB1 JAK2 MAP2K7 MAPKAPK2 NFKB1 PRKCE S100B STAT2 TYK2 CREB5 FOS HRAS JAK3 MAP3K1 MEF2C NFKB2 PTEN SEMA4D STAT3 CREBBP GATA3 IGF1R KIT MAP3K5 MTDH NFKBIA PYCARD SMAD2 STAT4 Chemokine CCL1 CCL16 CCL20 CCL25 CCL4 CCR2 CCR7 CX3CL1 CXCL12 CXCL3 CXCR1 CXCR6 CCL11 CCL17 CCL21 CCL26 CCL5 CCR3 CCR9 CX3CR1 CXCL13 CXCL5 CXCR2 MST1R CCL13 CCL18 CCL22 CCL27 CCL7 CCR4 CCRL2 CXCL1 CXCL14 CXCL6 CXCR3 PPBP CCL14 CCL19 CCL23 CCL28 CCL8 CCR5 CKLF CXCL10 CXCL16 CXCL8 CXCR4 XCL2 CCL15 CCL2 CCL24 CCL3 CCR1 CCR6 CMKLR1 CXCL11 CXCL2 CXCL9 CXCR5 -

The Expression and Regulation of Chemerin in the Epidermis
RESEARCH ARTICLE The Expression and Regulation of Chemerin in the Epidermis Magdalena Banas1, Aneta Zegar1, Mateusz Kwitniewski1, Katarzyna Zabieglo1, Joanna Marczynska1, Monika Kapinska-Mrowiecka2, Melissa LaJevic3,4, Brian A. Zabel4, Joanna Cichy1* 1 Department of Immunology, Faculty of Biochemistry, Biophysics and Biotechnology, Jagiellonian University, Kraków, Poland, 2 Department of Dermatology, Zeromski Hospital, Kraków, Poland, 3 Stanford University School of Medicine, Department of Pathology, Stanford, California, United States of America, 4 Palo Alto Veterans Institute for Research, VA Palo Alto Health Care System, Palo Alto, California, United States of America * [email protected] Abstract OPEN ACCESS Chemerin is a protein ligand for the G protein-coupled receptor CMKLR1 and also binds to two atypical heptahelical receptors, CCRL2 and GPR1. Chemerin is a leukocyte attractant, Citation: Banas M, Zegar A, Kwitniewski M, Zabieglo K, Marczynska J, Kapinska-Mrowiecka M, et al. adipokine, and antimicrobial protein. Although chemerin was initially identified as a highly (2015) The Expression and Regulation of Chemerin expressed gene in healthy skin keratinocytes that was downregulated during psoriasis, the in the Epidermis. PLoS ONE 10(2): e0117830. regulation of chemerin and its receptors in the skin by specific cytokines and microbial fac- doi:10.1371/journal.pone.0117830 tors remains unexplored. Here we show that chemerin, CMKLR1, CCRL2 and GPR1 are Academic Editor: Bernhard Ryffel, French National expressed in human and mouse epidermis, suggesting that this tissue may be both a Centre for Scientific Research, FRANCE source and target for chemerin mediated effects. In human skin cultures, chemerin is signifi- Received: October 1, 2014 cantly downregulated by IL-17 and IL-22, key cytokines implicated in psoriasis, whereas it is Accepted: December 31, 2014 upregulated by acute phase cytokines oncostatin M and IL-1β.