Next Generation Sequencing (V10)
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Supplementary Data
Supplementary Data for Quantitative Changes in the Mitochondrial Proteome from Subjects with Mild Cognitive Impairment, Early Stage and Late Stage Alzheimer’s disease Table 1 - 112 unique, non-redundant proteins identified and quantified in at least two of the three analytical replicates for all three disease stages. Table 2 - MCI mitochondrial samples, Protein Summary Table 3 - MCI mitochondrial samples, Experiment 1 Table 4 - MCI mitochondrial samples, Experiment 2 Table 5 - MCI mitochondrial samples, Experiment 3 Table 6 - EAD Mitochondrial Study, Protein Summary Table 7 - EAD Mitochondrial Study, Experiment 1 Table 8 - EAD Mitochondrial Study, Experiment 2 Table 9 - EAD Mitochondrial Study, Experiment 3 Table 10 - LAD Mitochondrial Study, Protein Summary Table 11 - LAD Mitochondrial Study, Experiment 1 Table 12 - LAD Mitochondrial Study, Experiment 2 Table 13 - LAD Mitochondrial Study, Experiment 3 Supplemental Table 1. 112 unique, non-redundant proteins identified and quantified in at least two of the three analytical replicates for all three disease stages. Description Data MCI EAD LAD AATM_HUMAN (P00505) Aspartate aminotransferase, mitochondrial precursor (EC Mean 1.43 1.70 1.31 2.6.1.1) (Transaminase A) (Glutamate oxaloacetate transaminase 2) [MASS=47475] SEM 0.07 0.09 0.09 Count 3.00 3.00 3.00 ACON_HUMAN (Q99798) Aconitate hydratase, mitochondrial precursor (EC 4.2.1.3) Mean 1.24 1.61 1.19 (Citrate hydro-lyase) (Aconitase) [MASS=85425] SEM 0.05 0.17 0.18 Count 3.00 2.00 3.00 ACPM_HUMAN (O14561) Acyl carrier protein, mitochondrial -

The Rise and Fall of the Bovine Corpus Luteum
University of Nebraska Medical Center DigitalCommons@UNMC Theses & Dissertations Graduate Studies Spring 5-6-2017 The Rise and Fall of the Bovine Corpus Luteum Heather Talbott University of Nebraska Medical Center Follow this and additional works at: https://digitalcommons.unmc.edu/etd Part of the Biochemistry Commons, Molecular Biology Commons, and the Obstetrics and Gynecology Commons Recommended Citation Talbott, Heather, "The Rise and Fall of the Bovine Corpus Luteum" (2017). Theses & Dissertations. 207. https://digitalcommons.unmc.edu/etd/207 This Dissertation is brought to you for free and open access by the Graduate Studies at DigitalCommons@UNMC. It has been accepted for inclusion in Theses & Dissertations by an authorized administrator of DigitalCommons@UNMC. For more information, please contact [email protected]. THE RISE AND FALL OF THE BOVINE CORPUS LUTEUM by Heather Talbott A DISSERTATION Presented to the Faculty of the University of Nebraska Graduate College in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy Biochemistry and Molecular Biology Graduate Program Under the Supervision of Professor John S. Davis University of Nebraska Medical Center Omaha, Nebraska May, 2017 Supervisory Committee: Carol A. Casey, Ph.D. Andrea S. Cupp, Ph.D. Parmender P. Mehta, Ph.D. Justin L. Mott, Ph.D. i ACKNOWLEDGEMENTS This dissertation was supported by the Agriculture and Food Research Initiative from the USDA National Institute of Food and Agriculture (NIFA) Pre-doctoral award; University of Nebraska Medical Center Graduate Student Assistantship; University of Nebraska Medical Center Exceptional Incoming Graduate Student Award; the VA Nebraska-Western Iowa Health Care System Department of Veterans Affairs; and The Olson Center for Women’s Health, Department of Obstetrics and Gynecology, Nebraska Medical Center. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Predicting Coupling Probabilities of G-Protein Coupled Receptors Gurdeep Singh1,2,†, Asuka Inoue3,*,†, J
Published online 30 May 2019 Nucleic Acids Research, 2019, Vol. 47, Web Server issue W395–W401 doi: 10.1093/nar/gkz392 PRECOG: PREdicting COupling probabilities of G-protein coupled receptors Gurdeep Singh1,2,†, Asuka Inoue3,*,†, J. Silvio Gutkind4, Robert B. Russell1,2,* and Francesco Raimondi1,2,* 1CellNetworks, Bioquant, Heidelberg University, Im Neuenheimer Feld 267, 69120 Heidelberg, Germany, 2Biochemie Zentrum Heidelberg (BZH), Heidelberg University, Im Neuenheimer Feld 328, 69120 Heidelberg, Germany, 3Graduate School of Pharmaceutical Sciences, Tohoku University, Sendai, Miyagi 980-8578, Japan and 4Department of Pharmacology and Moores Cancer Center, University of California, San Diego, La Jolla, CA 92093, USA Received February 10, 2019; Revised April 13, 2019; Editorial Decision April 24, 2019; Accepted May 01, 2019 ABSTRACT great use in tinkering with signalling pathways in living sys- tems (5). G-protein coupled receptors (GPCRs) control multi- Ligand binding to GPCRs induces conformational ple physiological states by transducing a multitude changes that lead to binding and activation of G-proteins of extracellular stimuli into the cell via coupling to situated on the inner cell membrane. Most of mammalian intra-cellular heterotrimeric G-proteins. Deciphering GPCRs couple with more than one G-protein giving each which G-proteins couple to each of the hundreds receptor a distinct coupling profile (6) and thus specific of GPCRs present in a typical eukaryotic organism downstream cellular responses. Determining these coupling is therefore critical to understand signalling. Here, profiles is critical to understand GPCR biology and phar- we present PRECOG (precog.russelllab.org): a web- macology. Despite decades of research and hundreds of ob- server for predicting GPCR coupling, which allows served interactions, coupling information is still missing for users to: (i) predict coupling probabilities for GPCRs many receptors and sequence determinants of coupling- specificity are still largely unknown. -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -

Analysis of the Indacaterol-Regulated Transcriptome in Human Airway
Supplemental material to this article can be found at: http://jpet.aspetjournals.org/content/suppl/2018/04/13/jpet.118.249292.DC1 1521-0103/366/1/220–236$35.00 https://doi.org/10.1124/jpet.118.249292 THE JOURNAL OF PHARMACOLOGY AND EXPERIMENTAL THERAPEUTICS J Pharmacol Exp Ther 366:220–236, July 2018 Copyright ª 2018 by The American Society for Pharmacology and Experimental Therapeutics Analysis of the Indacaterol-Regulated Transcriptome in Human Airway Epithelial Cells Implicates Gene Expression Changes in the s Adverse and Therapeutic Effects of b2-Adrenoceptor Agonists Dong Yan, Omar Hamed, Taruna Joshi,1 Mahmoud M. Mostafa, Kyla C. Jamieson, Radhika Joshi, Robert Newton, and Mark A. Giembycz Departments of Physiology and Pharmacology (D.Y., O.H., T.J., K.C.J., R.J., M.A.G.) and Cell Biology and Anatomy (M.M.M., R.N.), Snyder Institute for Chronic Diseases, Cumming School of Medicine, University of Calgary, Calgary, Alberta, Canada Received March 22, 2018; accepted April 11, 2018 Downloaded from ABSTRACT The contribution of gene expression changes to the adverse and activity, and positive regulation of neutrophil chemotaxis. The therapeutic effects of b2-adrenoceptor agonists in asthma was general enriched GO term extracellular space was also associ- investigated using human airway epithelial cells as a therapeu- ated with indacaterol-induced genes, and many of those, in- tically relevant target. Operational model-fitting established that cluding CRISPLD2, DMBT1, GAS1, and SOCS3, have putative jpet.aspetjournals.org the long-acting b2-adrenoceptor agonists (LABA) indacaterol, anti-inflammatory, antibacterial, and/or antiviral activity. Numer- salmeterol, formoterol, and picumeterol were full agonists on ous indacaterol-regulated genes were also induced or repressed BEAS-2B cells transfected with a cAMP-response element in BEAS-2B cells and human primary bronchial epithelial cells by reporter but differed in efficacy (indacaterol $ formoterol . -

Neurodegeneration Induces a Developmental RNA Processing Program by Calpain
bioRxiv preprint doi: https://doi.org/10.1101/2020.07.24.219121; this version posted July 25, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Neurodegeneration induces a developmental RNA processing program by calpain- mediated MBNL2 degradation Authors/Affiliations: Lee-Hsin Wanga,b, Yu-Mei Linb, Chien-Yu Linb, Yijuang Cherna,b,c and Guey-Shin Wanga,b,c,* aTaiwan International Graduate Program in Interdisciplinary Neuroscience, National Yang- Ming University and Academia Sinica, Taipei, Taiwan bInstitute of Biomedical Sciences, Academia Sinica, Taipei 11529, Taiwan cProgram in Molecular Medicine, National Yang-Ming University and Academia Sinica, Taipei, Taiwan *To whom correspondence should be addressed: Guey-Shin Wang, PhD Institute of Biomedical Sciences Academia Sinica 128, Section 2, Academia Rd. Taipei 11529, Taiwan. Tel: (886)-226-523-051 Fax: (886)-227-829-142 Email: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.07.24.219121; this version posted July 25, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The Muscleblind-like (MBNL) protein family plays an important role in regulating developmental RNA processing transition. Loss of MBNL2 function has been implicated in the neurodegeneration of myotonic dystrophy type 1 (DM1). However, the causal mechanism of neurodegeneration-induced MBNL2 loss of function remains elusive. Here, we show that neurodegenerative conditions including NMDAR-mediated excitotoxicity and dysregulated calcium homeostasis triggered nuclear translocation of calpain-2 resulting in MBNL2 degradation and reversion of MBNL2-regulated RNA processing to developmental patterns. -

Molecular Diagnostic Requisition
BAYLOR MIRACA GENETICS LABORATORIES SHIP TO: Baylor Miraca Genetics Laboratories 2450 Holcombe, Grand Blvd. -Receiving Dock PHONE: 800-411-GENE | FAX: 713-798-2787 | www.bmgl.com Houston, TX 77021-2024 Phone: 713-798-6555 MOLECULAR DIAGNOSTIC REQUISITION PATIENT INFORMATION SAMPLE INFORMATION NAME: DATE OF COLLECTION: / / LAST NAME FIRST NAME MI MM DD YY HOSPITAL#: ACCESSION#: DATE OF BIRTH: / / GENDER (Please select one): FEMALE MALE MM DD YY SAMPLE TYPE (Please select one): ETHNIC BACKGROUND (Select all that apply): UNKNOWN BLOOD AFRICAN AMERICAN CORD BLOOD ASIAN SKELETAL MUSCLE ASHKENAZIC JEWISH MUSCLE EUROPEAN CAUCASIAN -OR- DNA (Specify Source): HISPANIC NATIVE AMERICAN INDIAN PLACE PATIENT STICKER HERE OTHER JEWISH OTHER (Specify): OTHER (Please specify): REPORTING INFORMATION ADDITIONAL PROFESSIONAL REPORT RECIPIENTS PHYSICIAN: NAME: INSTITUTION: PHONE: FAX: PHONE: FAX: NAME: EMAIL (INTERNATIONAL CLIENT REQUIREMENT): PHONE: FAX: INDICATION FOR STUDY SYMPTOMATIC (Summarize below.): *FAMILIAL MUTATION/VARIANT ANALYSIS: COMPLETE ALL FIELDS BELOW AND ATTACH THE PROBAND'S REPORT. GENE NAME: ASYMPTOMATIC/POSITIVE FAMILY HISTORY: (ATTACH FAMILY HISTORY) MUTATION/UNCLASSIFIED VARIANT: RELATIONSHIP TO PROBAND: THIS INDIVIDUAL IS CURRENTLY: SYMPTOMATIC ASYMPTOMATIC *If family mutation is known, complete the FAMILIAL MUTATION/ VARIANT ANALYSIS section. NAME OF PROBAND: ASYMPTOMATIC/POPULATION SCREENING RELATIONSHIP TO PROBAND: OTHER (Specify clinical findings below): BMGL LAB#: A COPY OF ORIGINAL RESULTS ATTACHED IF PROBAND TESTING WAS PERFORMED AT ANOTHER LAB, CALL TO DISCUSS PRIOR TO SENDING SAMPLE. A POSITIVE CONTROL MAY BE REQUIRED IN SOME CASES. REQUIRED: NEW YORK STATE PHYSICIAN SIGNATURE OF CONSENT I certify that the patient specified above and/or their legal guardian has been informed of the benefits, risks, and limitations of the laboratory test(s) requested. -

Multi-Functionality of Proteins Involved in GPCR and G Protein Signaling: Making Sense of Structure–Function Continuum with In
Cellular and Molecular Life Sciences (2019) 76:4461–4492 https://doi.org/10.1007/s00018-019-03276-1 Cellular andMolecular Life Sciences REVIEW Multi‑functionality of proteins involved in GPCR and G protein signaling: making sense of structure–function continuum with intrinsic disorder‑based proteoforms Alexander V. Fonin1 · April L. Darling2 · Irina M. Kuznetsova1 · Konstantin K. Turoverov1,3 · Vladimir N. Uversky2,4 Received: 5 August 2019 / Revised: 5 August 2019 / Accepted: 12 August 2019 / Published online: 19 August 2019 © Springer Nature Switzerland AG 2019 Abstract GPCR–G protein signaling system recognizes a multitude of extracellular ligands and triggers a variety of intracellular signal- ing cascades in response. In humans, this system includes more than 800 various GPCRs and a large set of heterotrimeric G proteins. Complexity of this system goes far beyond a multitude of pair-wise ligand–GPCR and GPCR–G protein interactions. In fact, one GPCR can recognize more than one extracellular signal and interact with more than one G protein. Furthermore, one ligand can activate more than one GPCR, and multiple GPCRs can couple to the same G protein. This defnes an intricate multifunctionality of this important signaling system. Here, we show that the multifunctionality of GPCR–G protein system represents an illustrative example of the protein structure–function continuum, where structures of the involved proteins represent a complex mosaic of diferently folded regions (foldons, non-foldons, unfoldons, semi-foldons, and inducible foldons). The functionality of resulting highly dynamic conformational ensembles is fne-tuned by various post-translational modifcations and alternative splicing, and such ensembles can undergo dramatic changes at interaction with their specifc partners. -

New Tests 2 Test Updates 4 CPT Code Updates 11 Deleted Tests 11 Diagnostics Update Volume XXI, No
Volume XXI, No. 6 JUNE 2021 MONTHLY TEST UPDATES Diagnostics Update New Tests 2 Test Updates 4 CPT Code Updates 11 Deleted Tests 11 Diagnostics Update Volume XXI, No. 6 | JUNE 2021 New Tests Use Anti-DFS70 antibodies may help identify individuals who do not have an Anti-Carbamylated Protein (CarP) Antibody 520311 ANA-associated Autoimmune Rheumatic Disease (AARD) especially in the absence of significant clinical findings.1 Anti-DFS70 Ab, especially when positive CPT 83516 in isolation (i.e. in the absence of AARD-associated autoantibodies), may Synonyms Anti-CarP antigen antibody; RA marker prevent unnecessary referrals and examinations of ANA-positive individuals.2 Special Instructions This test has not been approved for NY state clients. Limitations This test should be used with clinical findings and other Specimen Serum autoimmune testing; it cannot be used alone to rule out autoimmune disease. Volume 1 mL This test was developed and its performance characteristics determined Minimum Volume 0.5 mL by Labcorp. It has not been cleared or approved by the Food and Drug Container Red-top tube; serum from red-top tube; serum from a gel tube; or Administration. serum gel tube Methodology Enzyme-linked immunosorbent assay (ELISA) Collection Separate serum from cells within one hour of collection. Transfer to a Additional Information Anti-DFS70 antibodies target the dense fine speckled plastic transport tube before shipping. protein of 70 kDa which is identical to Lens Epithelium-Derived Growth Factor Storage Instructions Refrigerate or freeze. or transcription co-activator p75 (LEDGFp75). They are detectable in 2% to 22% Stability of healthy individuals and in less than 1% of patients with AARD are of unknown Temperature Period clinical significance. -

High Constitutive Cytokine Release by Primary Human Acute Myeloid Leukemia Cells Is Associated with a Specific Intercellular Communication Phenotype
Supplementary Information High Constitutive Cytokine Release by Primary Human Acute Myeloid Leukemia Cells Is Associated with a Specific Intercellular Communication Phenotype Håkon Reikvam 1,2,*, Elise Aasebø 1, Annette K. Brenner 2, Sushma Bartaula-Brevik 1, Ida Sofie Grønningsæter 2, Rakel Brendsdal Forthun 2, Randi Hovland 3,4 and Øystein Bruserud 1,2 1 Department of Clinical Science, University of Bergen, 5020, Bergen, Norway 2 Department of Medicine, Haukeland University Hospital, 5021, Bergen, Norway 3 Department of Medical Genetics, Haukeland University Hospital, 5021, Bergen, Norway 4 Institute of Biomedicine, University of Bergen, 5020, Bergen, Norway * Correspondence: [email protected]; Tel.: +55-97-50-00 J. Clin. Med. 2019, 8, x 2 of 36 Figure S1. Mutational studies in a cohort of 71 AML patients. The figure shows the number of patients with the various mutations (upper), the number of mutations in for each patient (middle) and the number of main classes with mutation(s) in each patient (lower). 2 J. Clin. Med. 2019, 8, x; doi: www.mdpi.com/journal/jcm J. Clin. Med. 2019, 8, x 3 of 36 Figure S2. The immunophenotype of primary human AML cells derived from 62 unselected patients. The expression of the eight differentiation markers CD13, CD14, CD15, CD33, CD34, CD45, CD117 and HLA-DR was investigated for 62 of the 71 patients included in our present study. We performed an unsupervised hierarchical cluster analysis and identified four patient main clusters/patient subsets. The mutational profile for each f the 62 patients is also given (middle), no individual mutation of main class of mutations showed any significant association with any of the for differentiation marker clusters (middle). -
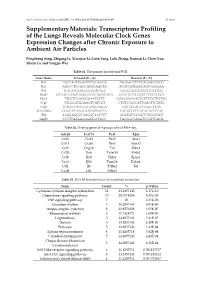
Transcriptome Profiling of the Lungs Reveals Molecular Clock Genes Expression Changes After Chronic Exposure to Ambient Air Particles
Int. J. Environ. Res. Public Health 2017, 14, 0090; doi:10.3390/ijerph14010090 S1 of S6 Supplementary Materials: Transcriptome Profiling of the Lungs Reveals Molecular Clock Genes Expression Changes after Chronic Exposure to Ambient Air Particles Pengcheng Song, Zhigang Li, Xiaoqian Li, Lixin Yang, Lulu Zhang, Nannan Li, Chen Guo, Shuyu Lu and Yongjie Wei Table S1. The primers for real-time PCR. Gene Name Forward (5'—3') Reverse (5'—3') Per1 CAGCAGTGGAGTCTGGAGGA TAGGAGCTCTGAGAAGCGGG Per2 AGCCCTGCAGCATGGAAGTA ACGTCATGAGGAGCCAGGAA Per3 TGTGTTCAAGGGTCCACTGC GGTGCTGGCAACTTCTTTCG Bmal1 CCAAGAAAGTATGGACACAGACAAA GCATTCTTGATCCTTCCTTGGT Clock TTGCTCCACGGGAATCCTT GGAGGGAAAGTGCTCTGTTGTAG Cry1 CTGGCGTGGAAGTCATCGT CTGTCCGCCATTGAGTTCTATG Cry2 TGTCCCTTCCTGTGTGGAAGA GCTCCCAGCTTGGCTTGA REV-ERBα GGGCACAAGCAACATTACCA CACGTCCCCACACACCTTAC Dbp AAGGAGCGCAAGGCAACTCT AGATGTCAAGCCTGCGCGGT Gapdh CCCTTAAGAGGGATGCTGCC TACGGCCAAATCCGTTCACA Table S2. Overlap genes of 4 groups data of RNA-Seq. Adcy9 Cxcl14 Per1 A2m Arntl Cxcl2 Per2 Alas1 Ccl11 Cxcl6 Per3 Alox15 Ccl2 Gng10 Tnf Ddit4 Ccl20 Ifnk Tnfsf10 Efnb2 Ccl9 Il10 Hif3a Epas1 Cry1 Il1b Trim16 Fabp4 Ctf1 Il6 Prl8a2 Tef Cxcl1 Lifr Prl4a1 Table S3. DAVID bioinformatics for functional annotation. Term Count % p-Value Cytokine-cytokine receptor interaction 12 34.2857143 1.27E-10 Chemokine signaling pathway 10 28.5714286 8.32E-09 TNF signaling pathway 7 20 2.31E-06 Circadian rhythm 5 14.2857143 4.03E-06 Herpes simplex infection 8 22.8571429 1.05E-05 Rheumatoid arthritis 6 17.1428571 1.60E-05 Legionellosis 5 14.2857143 5.41E-05 Malaria 5 14.2857143 6.20E-05 Pertussis 5 14.2857143 1.43E-04 African trypanosomiasis 4 11.4285714 3.62E-04 Circadian entrainment 5 14.2857143 4.28E-04 Chagas disease (American 5 14.2857143 6.21E-04 trypanosomiasis) NOD-like receptor signaling pathway 4 11.4285714 0.001137577 Jak-STAT signaling pathway 5 14.2857143 0.001482411 Inflammatory bowel disease (IBD) 4 11.4285714 0.001752567 Int.