Aquat Microb Ecol 55: 267-284
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Jpn. J. Protozool. 38(2) 171-183
Jpn. J. Protozool. Vol. 38, No. 2. (2005) 171 Review On the origin of mitochondria and Rickettsia-related eukaryotic endo- symbionts B. Franz Lang1*, Henner Brinkmann1, Liisa B. Koski1, Masahiro Fujishima2, Hans-Dieter Görtz3 and Gertraud Burger1 1Program in Evolutionary Biology, Canadian Institute for Advanced Research; Centre Robert Cedergren, Département de Biochimie, Université de Montréal, 2900 Boulevard Edouard- Montpetit, Montréal, Québec, H3T 1J4, Canada. 2Biological Institute, Faculty of Science, Yamaguchi University, Yamaguchi 753-8512, Japan. 3Abteilung Zoologie, Biologisches Institut, Universität Stuttgart, Pfaffenwaldring 57, 70550 Stuttgart, Germany. SUMMARY sence of genes for oxidative phosphorylation, the TCA cycle, and many other metabolic pathways, Resent insights into the origin and early evo- but the presence of several pathogenesis-related lution of mitochondria come from two approaches: genes and a high number of bacterial IS elements. the investigation of mtDNAs from minimally de- Phylogenetic analyses with multiple protein se- rived (primitive) mitochondriate eukaryotes, in quences place H. obtusa basally to the Rickettsia- particular jakobid flagellates, and of genomes from Ehrlichia-Wolbachia assemblage of bacterial intracellular α-proteobacterial symbionts. Of par- pathogens. This leads us to postulate that H. ob- ticular interest in this context is Holospora obtusa, tusa is the closest bacterial relative of mitochon- an intracellular bacterial endosymbiont that resides dria known to date. and replicates in the somatic nucleus of its eu- karyotic host, the ciliate Paramecium caudatum. Currently we have sequenced close to 50% of the INTRODUCTION ~ 1.7 Mbp H. obtusa genome, revealing the ab- One of the major advancements in under- standing eukaryotic evolution was the discovery that mitochondria evolved from an endosymbiotic α-Proteobacterium, and that mitochondrial DNA *Corresponding author (mtDNA) is a relict bacterial genome. -

De Novo Transcriptome Assembly of Perkinsus Olseni Trophozoite Stimulated in Vitro with Manila Clam (Ruditapes Philippinarum) Plasma
Journal of Invertebrate Pathology 135 (2016) 22–33 Contents lists available at ScienceDirect Journal of Invertebrate Pathology journal homepage: www.elsevier.com/locate/jip De novo transcriptome assembly of Perkinsus olseni trophozoite stimulated in vitro with Manila clam (Ruditapes philippinarum) plasma Abul Farah Md. Hasanuzzaman a,b, Diego Robledo c, Antonio Gómez-Tato d, Jose A. Alvarez-Dios e, ⇑ Peter W. Harrison f, Asunción Cao g, Sergio Fernández-Boo g, Antonio Villalba g, Belén G. Pardo a, , Paulino Martínez a a Departamento de Xenética, Facultade de Veterinaria, Universidade de Santiago de Compostela, Lugo 27002, Spain b Fisheries and Marine Resource Technology Discipline, Khulna University, Khulna 9208, Bangladesh c Departamento de Xenética, Facultade de Bioloxía, Universidade de Santiago de Compostela, Santiago de Compostela 15782, Spain d Departamento de Xeometría e Topoloxía, Facultade de Matemáticas, Universidade de Santiago de Compostela, Santiago de Compostela 15782, Spain e Departamento de Matemática Aplicada, Facultade de Matemáticas, Universidade de Santiago de Compostela, Santiago de Compostela 15782, Spain f Department of Genetics, Evolution and Environment, University College London, London WC1E 6BT, United Kingdom g Centro de Investigacións Mariñas (CIMA), Consellería do Medio Rural e do Mar, Xunta de Galicia, 36620 Vilanova de Arousa, Spain article info abstract Article history: The protistan parasite Perkinsus olseni is a deadly causative agent of perkinsosis, a molluscan disease Received 16 September 2015 affecting Manila clam (Ruditapes philippinarum), having a significant impact on world mollusc production. Revised 18 January 2016 Deciphering the underlying molecular mechanisms in R. philippinarum-P. olseni interaction is crucial for Accepted 24 January 2016 controlling this parasitosis. The present study investigated the transcriptional expression in the parasite Available online 25 January 2016 trophozoite using RNA-seq. -

The Planktonic Protist Interactome: Where Do We Stand After a Century of Research?
bioRxiv preprint doi: https://doi.org/10.1101/587352; this version posted May 2, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Bjorbækmo et al., 23.03.2019 – preprint copy - BioRxiv The planktonic protist interactome: where do we stand after a century of research? Marit F. Markussen Bjorbækmo1*, Andreas Evenstad1* and Line Lieblein Røsæg1*, Anders K. Krabberød1**, and Ramiro Logares2,1** 1 University of Oslo, Department of Biosciences, Section for Genetics and Evolutionary Biology (Evogene), Blindernv. 31, N- 0316 Oslo, Norway 2 Institut de Ciències del Mar (CSIC), Passeig Marítim de la Barceloneta, 37-49, ES-08003, Barcelona, Catalonia, Spain * The three authors contributed equally ** Corresponding authors: Ramiro Logares: Institute of Marine Sciences (ICM-CSIC), Passeig Marítim de la Barceloneta 37-49, 08003, Barcelona, Catalonia, Spain. Phone: 34-93-2309500; Fax: 34-93-2309555. [email protected] Anders K. Krabberød: University of Oslo, Department of Biosciences, Section for Genetics and Evolutionary Biology (Evogene), Blindernv. 31, N-0316 Oslo, Norway. Phone +47 22845986, Fax: +47 22854726. [email protected] Abstract Microbial interactions are crucial for Earth ecosystem function, yet our knowledge about them is limited and has so far mainly existed as scattered records. Here, we have surveyed the literature involving planktonic protist interactions and gathered the information in a manually curated Protist Interaction DAtabase (PIDA). In total, we have registered ~2,500 ecological interactions from ~500 publications, spanning the last 150 years. -

Sex Is a Ubiquitous, Ancient, and Inherent Attribute of Eukaryotic Life
PAPER Sex is a ubiquitous, ancient, and inherent attribute of COLLOQUIUM eukaryotic life Dave Speijera,1, Julius Lukešb,c, and Marek Eliášd,1 aDepartment of Medical Biochemistry, Academic Medical Center, University of Amsterdam, 1105 AZ, Amsterdam, The Netherlands; bInstitute of Parasitology, Biology Centre, Czech Academy of Sciences, and Faculty of Sciences, University of South Bohemia, 370 05 Ceské Budejovice, Czech Republic; cCanadian Institute for Advanced Research, Toronto, ON, Canada M5G 1Z8; and dDepartment of Biology and Ecology, University of Ostrava, 710 00 Ostrava, Czech Republic Edited by John C. Avise, University of California, Irvine, CA, and approved April 8, 2015 (received for review February 14, 2015) Sexual reproduction and clonality in eukaryotes are mostly Sex in Eukaryotic Microorganisms: More Voyeurs Needed seen as exclusive, the latter being rather exceptional. This view Whereas absence of sex is considered as something scandalous for might be biased by focusing almost exclusively on metazoans. a zoologist, scientists studying protists, which represent the ma- We analyze and discuss reproduction in the context of extant jority of extant eukaryotic diversity (2), are much more ready to eukaryotic diversity, paying special attention to protists. We accept that a particular eukaryotic group has not shown any evi- present results of phylogenetically extended searches for ho- dence of sexual processes. Although sex is very well documented mologs of two proteins functioning in cell and nuclear fusion, in many protist groups, and members of some taxa, such as ciliates respectively (HAP2 and GEX1), providing indirect evidence for (Alveolata), diatoms (Stramenopiles), or green algae (Chlor- these processes in several eukaryotic lineages where sex has oplastida), even serve as models to study various aspects of sex- – not been observed yet. -

Role of Metopus Es in the Anaerobic Degradation of Organic Matter and Biomethanation Process
Role of Metopus es in the anaerobic degradation of organic matter and biomethanation process Thesis submitted to the University of Kerala for award of the degree of Doctor of Philosophy in Microbiology By Nimi Narayanan Under the Supervision of Dr. V. B. Manilal Process Engineering and Environmental Technology Division, National Institute for Interdisciplinary Science and Technology (NIIST), CSIR, Thiruvananthapuram, Kerala, INDIA - 695019 2011 To my Family DECLARATION I hereby declare that the work presented in this thesis is based on the original work done by me under the guidance of Dr. V. B. Manilal, Principal Scientist, Process Engineering and Environmental Technology Division, National Institute for Interdisciplinary Science and Technology, and that no part of this has been included in any other thesis submitted previously for the award of any degree. Nimi Narayanan Acknowledgement It is a great pleasure to express my sincere gratitude and sense of appreciation to my research guide, Dr. V.B. Manilal, Principal Scientist, Environmental Technology, NIIST, Trivandrum, for his constant encouragement, enthusiastic support and valuable guidance throughout the period of study. I am indebted to him for giving me the ample freedom to do the work and express my ideas during this period. I am grateful to Dr. Ajit Haridas, Scientist-in-charge, Environmental Technology, NIIST for his valuable suggestions and constructive criticism during my tenure. It is an honor for me to thank the present Director, Dr. Suresh Das and the former directors of NIIST, Trivandrum for providing the necessary infrastructural facilities for the successful completion of work. I would like to thank Council of Scientific and Industrial Research, New Delhi, for the research fellowship. -

University of Oklahoma
UNIVERSITY OF OKLAHOMA GRADUATE COLLEGE MACRONUTRIENTS SHAPE MICROBIAL COMMUNITIES, GENE EXPRESSION AND PROTEIN EVOLUTION A DISSERTATION SUBMITTED TO THE GRADUATE FACULTY in partial fulfillment of the requirements for the Degree of DOCTOR OF PHILOSOPHY By JOSHUA THOMAS COOPER Norman, Oklahoma 2017 MACRONUTRIENTS SHAPE MICROBIAL COMMUNITIES, GENE EXPRESSION AND PROTEIN EVOLUTION A DISSERTATION APPROVED FOR THE DEPARTMENT OF MICROBIOLOGY AND PLANT BIOLOGY BY ______________________________ Dr. Boris Wawrik, Chair ______________________________ Dr. J. Phil Gibson ______________________________ Dr. Anne K. Dunn ______________________________ Dr. John Paul Masly ______________________________ Dr. K. David Hambright ii © Copyright by JOSHUA THOMAS COOPER 2017 All Rights Reserved. iii Acknowledgments I would like to thank my two advisors Dr. Boris Wawrik and Dr. J. Phil Gibson for helping me become a better scientist and better educator. I would also like to thank my committee members Dr. Anne K. Dunn, Dr. K. David Hambright, and Dr. J.P. Masly for providing valuable inputs that lead me to carefully consider my research questions. I would also like to thank Dr. J.P. Masly for the opportunity to coauthor a book chapter on the speciation of diatoms. It is still such a privilege that you believed in me and my crazy diatom ideas to form a concise chapter in addition to learn your style of writing has been a benefit to my professional development. I’m also thankful for my first undergraduate research mentor, Dr. Miriam Steinitz-Kannan, now retired from Northern Kentucky University, who was the first to show the amazing wonders of pond scum. Who knew that studying diatoms and algae as an undergraduate would lead me all the way to a Ph.D. -

Morphology and Phylogeny of the Soil Ciliate Metopus Yantaiensis N. Sp
Journal of Eukaryotic Microbiology ISSN 1066-5234 ORIGINAL ARTICLE Morphology and Phylogeny of the Soil Ciliate Metopus yantaiensis n. sp. (Ciliophora, Metopida), with Identification of the Intracellular Bacteria Atef Omara,b, Qianqian Zhanga, Songbao Zoua,c & Jun Gonga,c a Laboratory of Microbial Ecology and Matter Cycles, Yantai Institute of Coastal Zone Research, Chinese Academy of Sciences, Yantai 264003, China b Department of Zoology, Al-Azhar University, Assiut 71524, Egypt c University of Chinese Academy of Sciences, Beijing 100049, China Keywords ABSTRACT Anaerobic ciliates; Armophorea; digestion- resistant bacteria; Metopus contortus; The morphology and infraciliature of a new ciliate, Metopus yantaiensis n. sp., Metopus hasei. discovered in coastal soil of northern China, were investigated. It is distin- guished from its congeners by a combination of the following features: nuclear Correspondence apparatus situated in the preoral dome; 18–21 somatic ciliary rows, of which J. Gong, Yantai Institute of Coastal Zone three extend onto the preoral dome (dome kineties); three to five distinctly Research, Chinese Academy of Sciences, elongated caudal cilia, and 21–29 adoral polykinetids. The 18S rRNA genes of Yantai 264003, China this new species and two congeners, Metopus contortus and Metopus hasei, Telephone number: +86-535-2109123; were sequenced and phylogenetically analyzed. The new species is more clo- FAX number: +86-535-2109000; sely related to M. hasei and the clevelandellids than to other congeners; both e-mail: [email protected] the genus Metopus and the order Metopida are not monophyletic. In addition, the digestion-resistant bacteria in the cytoplasm of M. yantaiensis were identi- Received: 13 January 2017; revised 2 March fied, using a 16S rRNA gene clone library, sequencing, and fluorescence 2017; accepted March 8, 2017. -

BMC Research Notes Biomed Central
BMC Research Notes BioMed Central Short Report Open Access A split and rearranged nuclear gene encoding the iron-sulfur subunit of mitochondrial succinate dehydrogenase in Euglenozoa Ryan MR Gawryluk and Michael W Gray* Address: Department of Biochemistry and Molecular Biology, Dalhousie University, Halifax, Nova Scotia B3H 1X5, Canada Email: Ryan MR Gawryluk - [email protected]; Michael W Gray* - [email protected] * Corresponding author Published: 3 February 2009 Received: 18 December 2008 Accepted: 3 February 2009 BMC Research Notes 2009, 2:16 doi:10.1186/1756-0500-2-16 This article is available from: http://www.biomedcentral.com/1756-0500/2/16 © 2009 Gray et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: Analyses based on phylogenetic and ultrastructural data have suggested that euglenids (such as Euglena gracilis), trypanosomatids and diplonemids are members of a monophyletic lineage termed Euglenozoa. However, many uncertainties are associated with phylogenetic reconstructions for ancient and rapidly evolving groups; thus, rare genomic characters become increasingly important in reinforcing inferred phylogenetic relationships. Findings: We discovered that the iron-sulfur subunit (SdhB) of mitochondrial succinate dehydrogenase is encoded by a split and rearranged nuclear gene in Euglena gracilis and trypanosomatids, an example of a rare genomic character. The two subgenic modules are transcribed independently and the resulting mRNAs appear to be independently translated, with the two protein products imported into mitochondria, based on the presence of predicted mitochondrial targeting peptides. -

Multigene Eukaryote Phylogeny Reveals the Likely Protozoan Ancestors of Opis- Thokonts (Animals, Fungi, Choanozoans) and Amoebozoa
Accepted Manuscript Multigene eukaryote phylogeny reveals the likely protozoan ancestors of opis- thokonts (animals, fungi, choanozoans) and Amoebozoa Thomas Cavalier-Smith, Ema E. Chao, Elizabeth A. Snell, Cédric Berney, Anna Maria Fiore-Donno, Rhodri Lewis PII: S1055-7903(14)00279-6 DOI: http://dx.doi.org/10.1016/j.ympev.2014.08.012 Reference: YMPEV 4996 To appear in: Molecular Phylogenetics and Evolution Received Date: 24 January 2014 Revised Date: 2 August 2014 Accepted Date: 11 August 2014 Please cite this article as: Cavalier-Smith, T., Chao, E.E., Snell, E.A., Berney, C., Fiore-Donno, A.M., Lewis, R., Multigene eukaryote phylogeny reveals the likely protozoan ancestors of opisthokonts (animals, fungi, choanozoans) and Amoebozoa, Molecular Phylogenetics and Evolution (2014), doi: http://dx.doi.org/10.1016/ j.ympev.2014.08.012 This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain. 1 1 Multigene eukaryote phylogeny reveals the likely protozoan ancestors of opisthokonts 2 (animals, fungi, choanozoans) and Amoebozoa 3 4 Thomas Cavalier-Smith1, Ema E. Chao1, Elizabeth A. Snell1, Cédric Berney1,2, Anna Maria 5 Fiore-Donno1,3, and Rhodri Lewis1 6 7 1Department of Zoology, University of Oxford, South Parks Road, Oxford OX1 3PS, UK. -

Molecular Identity of Strains of Heterotrophic Flagellates Isolated from Surface Waters and Deep-Sea Sediments of the South Atlantic Based on SSU Rdna
AQUATIC MICROBIAL ECOLOGY Vol. 38: 239–247, 2005 Published March 18 Aquat Microb Ecol Molecular identity of strains of heterotrophic flagellates isolated from surface waters and deep-sea sediments of the South Atlantic based on SSU rDNA Frank Scheckenbach1, Claudia Wylezich1, Markus Weitere1, Klaus Hausmann2, Hartmut Arndt1,* 1Department of General Ecology and Limnology, Zoological Institute, University of Cologne, 50923 Cologne, Germany 2Institute of Biology/Zoology, Free University of Berlin, Research Group Protozoology, 14195 Berlin, Germany ABSTRACT: Whereas much is known about the biodiversity of prokaryotes and macroorganisms in the deep sea, knowledge on the biodiversity of protists remains very limited. Molecular studies have changed our view of marine environments and have revealed an astonishing number of previously unknown eukaryotic organisms. Morphological findings have shown that at least some widely dis- tributed nanoflagellates can also be found in the deep sea. Whether these flagellates have contact with populations from other habitats is still uncertain. We performed a molecular comparison of strains isolated from deep-sea sediments (>5000 m depth) and surface waters on the basis of their small subunit ribosomal DNA (SSU rDNA). Sequences of Rhynchomonas nasuta, Amastigomonas debruynei, Ancyromonas sigmoides, Cafeteria roenbergensis and Caecitellus parvulus were analysed, and 2 contrasting results obtained. Firstly, we found nearly identical genotypes within 1 morphospecies (C. roenbergensis), and secondly, quite different genotypes within certain morpho- species (R. nasuta, A. sigmoides and C. parvulus). In addition, high genetic distances between the dif- ferent strains of A. sigmoides and C. parvulus indicate that these morphospecies should be divided into different at least genetically distinguishable species. In contrast, some heterotrophic nanoflagel- lates must indeed be regarded as being cosmopolitan. -
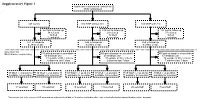
Supplementary Figure 1 Multicenter Randomised Control Trial 2746 Randomised
Supplementary Figure 1 Multicenter randomised control trial 2746 randomised 947 control 910 MNP without zinc 889 MNP with zinc 223 lost to follow up 219 lost to follow up 183 lost to follow up 34 refused 29 refused 37 refused 16 died 12 died 9 died 3 excluded 4 excluded 1 excluded 671 in follow-up 646 in follow-up 659 in follow-up at 24mo of age at 24mo of age at 24mo of age Selection for Microbiome sequencing 516 paired samples unavailable 469 paired samples unavailable 497 paired samples unavailable 69 antibiotic use 63 antibiotic use 67 antibiotic use 31 outside of WLZ criteria 37 outside of WLZ criteria 34 outside of WLZ criteria 6 diarrhea last 7 days 2 diarrhea last 7 days 7 diarrhea last 7 days 39 WLZ > -1 at 24 mo 10 WLZ < -2 at 24mo 58 WLZ > -1 at 24 mo 17 WLZ < -2 at 24mo 48 WLZ > -1 at 24 mo 8 WLZ < -2 at 24mo available for selection available for selection available for selection available for selection available for selection1 available for selection1 14 selected 10 selected 15 selected 14 selected 20 selected1 7 selected1 1 Two subjects (one in the reference WLZ group and one undernourished) had, at 12 months, no diarrhea within 1 day of stool collection but reported diarrhea within 7 days prior. Length, cm kg Weight, Supplementary Figure 2. Length (left) and weight (right) z-scores of children recruited into clinical trial NCT00705445 during the first 24 months of life. Median and quantile values are shown, with medians for participants profiled in current study indicated by red (undernourished) and black (reference WLZ) lines. -

Why the –Omic Future of Apicomplexa Should Include Gregarines Julie Boisard, Isabelle Florent
Why the –omic future of Apicomplexa should include Gregarines Julie Boisard, Isabelle Florent To cite this version: Julie Boisard, Isabelle Florent. Why the –omic future of Apicomplexa should include Gregarines. Biology of the Cell, Wiley, 2020, 10.1111/boc.202000006. hal-02553206 HAL Id: hal-02553206 https://hal.archives-ouvertes.fr/hal-02553206 Submitted on 24 Apr 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Article title: Why the –omic future of Apicomplexa should include Gregarines. Names of authors: Julie BOISARD1,2 and Isabelle FLORENT1 Authors affiliations: 1. Molécules de Communication et Adaptation des Microorganismes (MCAM, UMR 7245), Département Adaptations du Vivant (AVIV), Muséum National d’Histoire Naturelle, CNRS, CP52, 57 rue Cuvier 75231 Paris Cedex 05, France. 2. Structure et instabilité des génomes (STRING UMR 7196 CNRS / INSERM U1154), Département Adaptations du vivant (AVIV), Muséum National d'Histoire Naturelle, CP 26, 57 rue Cuvier 75231 Paris Cedex 05, France. Short Title: Gregarines –omics for Apicomplexa studies