Chromosome Preference of Disease Genes and Vectorization for the Prediction of Non-Coding Disease Genes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Nucleus Size and DNA Accessibility Are Linked to the Regulation Of
Grosch et al. BMC Biology (2020) 18:42 https://doi.org/10.1186/s12915-020-00770-y RESEARCH ARTICLE Open Access Nucleus size and DNA accessibility are linked to the regulation of paraspeckle formation in cellular differentiation Markus Grosch1, Sebastian Ittermann1, Ejona Rusha2, Tobias Greisle1, Chaido Ori1,3, Dong-Jiunn Jeffery Truong4, Adam C. O’Neill1, Anna Pertek2, Gil Gregor Westmeyer4 and Micha Drukker1,2* Abstract Background: Many long noncoding RNAs (lncRNAs) have been implicated in general and cell type-specific molecular regulation. Here, we asked what underlies the fundamental basis for the seemingly random appearance of nuclear lncRNA condensates in cells, and we sought compounds that can promote the disintegration of lncRNA condensates in vivo. Results: As a basis for comparing lncRNAs and cellular properties among different cell types, we screened lncRNAs in human pluripotent stem cells (hPSCs) that were differentiated to an atlas of cell lineages. We found that paraspeckles, which form by aggregation of the lncRNA NEAT1, are scaled by the size of the nucleus, and that small DNA-binding molecules promote the disintegration of paraspeckles and other lncRNA condensates. Furthermore, we found that paraspeckles regulate the differentiation of hPSCs. Conclusions: Positive correlation between the size of the nucleus and the number of paraspeckles exist in numerous types of human cells. The tethering and structure of paraspeckles, as well as other lncRNAs, to the genome can be disrupted by small molecules that intercalate in DNA. The structure-function relationship of lncRNAs that regulates stem cell differentiation is likely to be determined by the dynamics of nucleus size and binding site accessibility. -

Long Non-Coding RNA NEAT1 Promotes Proliferation, Migration
Int. J. Med. Sci. 2018, Vol. 15 1227 Ivyspring International Publisher International Journal of Medical Sciences 2018; 15(11): 1227-1234. doi: 10.7150/ijms.25662 Research Paper Long non-coding RNA NEAT1 promotes proliferation, migration and invasion of human osteosarcoma cells Pengcheng Li1, Rui Huang2, Tao Huang1, Shuo Cheng1, Yao Chen1, Zhihang Wang1 1. Department of Orthopedics, The First Affiliated Hospital of China Medical University. Shenyang 110001, Liaoning, P.R. China 2. Department of Clinical Medicine, Da Lian Medical University. Dalian 116000, Liaoning, P.R. China. Corresponding author: Tao Huang, Department of Orthopedics, The First Affiliated Hospital of China Medical University. Shenyang 110001, Liaoning, P.R. China. Email: [email protected] © Ivyspring International Publisher. This is an open access article distributed under the terms of the Creative Commons Attribution (CC BY-NC) license (https://creativecommons.org/licenses/by-nc/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2018.02.22; Accepted: 2018.05.27; Published: 2018.07.30 Abstract Aim: Long non-coding RNAs (LncRNAs) have been identified to play a crucial role in tumorigenesis and the progression of many types of tumors. However, the clinical significance and biological function of lncRNA nuclear-enriched abundant transcript 1(NEAT1) in human osteosarcoma remains unknown. Here, we investigated the role of NEAT1 in human osteosarcoma cell lines and clinical tumor samples. Methods: In this study, expression of NEAT1 was analyzed in 19 osteosarcoma tissues and paired adjacent non-tumor tissues by using quantitative real-time PCR. Additionally, knockdown of NEAT1 expression using Lentivirus-mediated siRNA was performed in order to explore the biological function of NEAT1 on osteosarcoma cell proliferation and metastasis through MTT, colony formation assay and transwell assay. -

Lncrna NEAT1 in Paraspeckles: a Structural Scaffold for Cellular DNA Damage Response Systems?
non-coding RNA Review LncRNA NEAT1 in Paraspeckles: A Structural Scaffold for Cellular DNA Damage Response Systems? Elisa Taiana 1,2,* , Domenica Ronchetti 1,2 , Katia Todoerti 2, Lucia Nobili 1 , Pierfrancesco Tassone 3, Nicola Amodio 3 and Antonino Neri 1,2,* 1 Department of Oncology and Hemato-oncology, University of Milan, 20122 Milan, Italy; [email protected] (D.R.); [email protected] (L.N.) 2 Hematology, Fondazione Cà Granda IRCCS Policlinico, 20122 Milan, Italy; [email protected] 3 Department of Experimental and Clinical Medicine, Magna Graecia University of Catanzaro, 88100 Catanzaro, Italy; [email protected] (P.T.); [email protected] (N.A.) * Correspondence: [email protected] (E.T.); [email protected] (A.N.); Tel.: +39-02-5032-0420 (E.T. & A.N.) Received: 5 June 2020; Accepted: 28 June 2020; Published: 1 July 2020 Abstract: Nuclear paraspeckle assembly transcript 1 (NEAT1) is a long non-coding RNA (lncRNA) reported to be frequently deregulated in various types of cancers and neurodegenerative processes. NEAT1 is an indispensable structural component of paraspeckles (PSs), which are dynamic and membraneless nuclear bodies that affect different cellular functions, including stress response. Furthermore, increasing evidence supports the crucial role of NEAT1 and essential structural proteins of PSs (PSPs) in the regulation of the DNA damage repair (DDR) system. This review aims to provide an overview of the current knowledge on the involvement of NEAT1 and PSPs in DDR, which might strengthen the rationale underlying future NEAT1-based therapeutic options in tumor and neurodegenerative diseases. Keywords: lncRNA; NEAT1; paraspeckle; DNA damage repair; cancer; neurodegenerative disease 1. -

Inhibition of Lncrna NEAT1 Suppresses the Inflammatory
INTERNATIONAL JOURNAL OF MOleCular meDICine 42: 2903-2913, 2018 Inhibition of lncRNA NEAT1 suppresses the inflammatory response in IBD by modulating the intestinal epithelial barrier and by exosome-mediated polarization of macrophages RUI LIU1-3, ANLIU TANG1-3, XIAOYAN WANG1-3, XIONG CHEN1-3, LIAN ZHAO1-3, ZHIMING XIAO1-3 and SHOURONG SHEN1-3 1Department of Gastroenterology, The Third Xiangya Hospital of Central South University; 2Hunan Key Laboratory of Nonresolving Inflammation and Cancer; 3National Key Clinical Specialty, Changsha, Hunan 410013, P.R. China Received February 10, 2018; Accepted July 24, 2018 DOI: 10.3892/ijmm.2018.3829 Abstract. Inflammatory bowel disease (IBD) is a multifac- of NEAT1 was high in IBD and was involved in the inflam- torial inflammatory disease, and increasing evidence has matory response by regulating the intestinal epithelial barrier demonstrated that the mechanism of the pathogenesis of IBD and through exosome‑mediated polarization of macrophages. is associated with intestinal epithelial barrier injury. Long The downregulation of NEAT1 suppressed the inflamma- non‑coding RNAs (lncRNAs) are a class of transcripts >200 tory response by modulating the intestinal epithelial barrier nucleotides in length with limited protein‑coding capability. and through exosome‑mediated polarization of macrophages Nuclear paraspeckle assembly transcript 1 (NEAT1) is a in IBD. The results of the present study revealed a potential recently identified nuclear‑restricted lncRNA, which localizes strategy of targeting NEAT1 -

Molecular Anatomy of the Architectural NEAT1 Noncoding
Molecular anatomy of the architectural NEAT1 noncoding RNA : The domains, interactors, and biogenesis pathway Title required to build phase-separated nuclear paraspeckles Author(s) Hirose, Tetsuro; Yamazaki, Tomohiro; Nakagawa, Shinichi Wiley interdisciplinary reviews, RNA, 10(6), e1545 Citation https://doi.org/10.1002/wrna.1545 Issue Date 2019-11 Doc URL http://hdl.handle.net/2115/79649 This is the peer reviewed version of the following article: Tetsuro Hirose Tomohiro Yamazaki Shinichi Nakagawa. (2019) Molecular anatomy of the architectural NEAT1 noncoding RNA: The domains, interactors, and biogenesis Rights pathway required to build phase-separated nuclear paraspeckles. Wiley interdisciplinary reviews. RNA: 10(6): e1545., which has been published in final form at https://doi.org/10.1002/wrna.1545. This article may be used for non- commercial purposes in accordance with Wiley Terms and Conditions for Use of Self-Archived Versions. Type article (author version) File Information Wiley Interdiscip Rev RNA_10(6)_e1545.pdf Instructions for use Hokkaido University Collection of Scholarly and Academic Papers : HUSCAP Article Title: Molecular anatomy of the architectural NEAT1 noncoding RNA: the domains, interactors, and biogenesis pathway required to build phase-separated nuclear paraspeckles Article Type: Authors: First author Tetsuro Hirose*, ORCID iD: https://orcid.org/0000-0003-1068-5464, Affiliation: Institute for Genetic Medicine, Hokkaido University, E-mail address: [email protected], no conflict of interest Second author Tomohiro Yamazaki, ORCID iD: https://orcid.org/0000-0003-0411-3590, Affiliation: Institute for Genetic Medicine, Hokkaido University, E-mail address: [email protected], no conflict of interest Third author Shinichi Nakagawa, ORCID iD: https://orcid.org/0000-0002-6806-7493, Faculty of Pharmaceutical Sciences, Hokkaido University, E-mail address: [email protected], no conflict of interest Abstract Long noncoding RNAs (lncRNAs) are extremely diverse and have various significant physiological functions. -

NEAT1 Long Isoform Is Highly Expressed in Chronic Lymphocytic Leukemia Irrespectively of Cytogenetic Groups Or Clinical Outcome
non-coding RNA Communication NEAT1 Long Isoform Is Highly Expressed in Chronic Lymphocytic Leukemia Irrespectively of Cytogenetic Groups or Clinical Outcome Domenica Ronchetti 1,2 , Vanessa Favasuli 1 , Paola Monti 3, Giovanna Cutrona 4 , Sonia Fabris 2, Ilaria Silvestris 1, Luca Agnelli 1 , Monica Colombo 4, Paola Menichini 3 , Serena Matis 4, Massimo Gentile 5, Ramil Nurtdinov 6 , Roderic Guigó 6 , Luca Baldini 1,2, Gilberto Fronza 3 , Manlio Ferrarini 7, Fortunato Morabito 8,9 , Antonino Neri 1,2,* and Elisa Taiana 1,2 1 Department of Oncology and Hemato-oncology, University of Milan, 20122 Milan, Italy; [email protected] (D.R.); [email protected] (V.F.); [email protected] (I.S.); [email protected] (L.A.); [email protected] (L.B.); [email protected] (E.T.) 2 Hematology, Fondazione Cà Granda IRCCS Policlinico, 20122 Milan, Italy; [email protected] 3 Mutagenesis and Cancer Prevention Unit, IRCCS Ospedale Policlinico San Martino, 16132 Genova, Italy; [email protected] (P.M.); [email protected] (P.M.); [email protected] (G.F.) 4 Molecular Pathology Unit, IRCCS Ospedale Policlinico San Martino, 16132 Genova, Italy; [email protected] (G.C.); [email protected] (M.C.); [email protected] (S.M.) 5 Hematology Unit, Department of Onco-Hematology A.O. of Cosenza, 87100 Cosenza, Italy; [email protected] 6 Centre for Genomic Regulation (CRG), The Barcelona Institute of Science and Technology, Dr. Aiguader 88, 08003 Barcelona, -

Tumor Hypoxia Induces Nuclear Paraspeckle Formation Through HIF-2Α Dependent Transcriptional Activation of NEAT1 Leading to Cancer Cell Survival
Oncogene (2015) 34, 4482–4490 OPEN © 2015 Macmillan Publishers Limited All rights reserved 0950-9232/15 www.nature.com/onc ORIGINAL ARTICLE Tumor hypoxia induces nuclear paraspeckle formation through HIF-2α dependent transcriptional activation of NEAT1 leading to cancer cell survival This article has been corrected since Advance Online Publication and a corrigendum is also printed in this issue. H Choudhry1,2, A Albukhari1,3, M Morotti3, S Haider3, D Moralli2, J Smythies4, J Schödel5, CM Green2, C Camps2, F Buffa6, P Ratcliffe4, J Ragoussis2,7,8,9, AL Harris3,9 and DR Mole4,9 Activation of cellular transcriptional responses, mediated by hypoxia-inducible factor (HIF), is common in many types of cancer, and generally confers a poor prognosis. Known to induce many hundreds of protein-coding genes, HIF has also recently been shown to be a key regulator of the non-coding transcriptional response. Here, we show that NEAT1 long non-coding RNA (lncRNA) is a direct transcriptional target of HIF in many breast cancer cell lines and in solid tumors. Unlike previously described lncRNAs, NEAT1 is regulated principally by HIF-2 rather than by HIF-1. NEAT1 is a nuclear lncRNA that is an essential structural component of paraspeckles and the hypoxic induction of NEAT1 induces paraspeckle formation in a manner that is dependent upon both NEAT1 and on HIF-2. Paraspeckles are multifunction nuclear structures that sequester transcriptionally active proteins as well as RNA transcripts that have been subjected to adenosine-to-inosine (A-to-I) editing. We show that the nuclear retention of one such transcript, F11R (also known as junctional adhesion molecule 1, JAM1), in hypoxia is dependent upon the hypoxic increase in NEAT1, thereby conferring a novel mechanism of HIF-dependent gene regulation. -

ID AKI Vs Control Fold Change P Value Symbol Entrez Gene Name *In
ID AKI vs control P value Symbol Entrez Gene Name *In case of multiple probesets per gene, one with the highest fold change was selected. Fold Change 208083_s_at 7.88 0.000932 ITGB6 integrin, beta 6 202376_at 6.12 0.000518 SERPINA3 serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 3 1553575_at 5.62 0.0033 MT-ND6 NADH dehydrogenase, subunit 6 (complex I) 212768_s_at 5.50 0.000896 OLFM4 olfactomedin 4 206157_at 5.26 0.00177 PTX3 pentraxin 3, long 212531_at 4.26 0.00405 LCN2 lipocalin 2 215646_s_at 4.13 0.00408 VCAN versican 202018_s_at 4.12 0.0318 LTF lactotransferrin 203021_at 4.05 0.0129 SLPI secretory leukocyte peptidase inhibitor 222486_s_at 4.03 0.000329 ADAMTS1 ADAM metallopeptidase with thrombospondin type 1 motif, 1 1552439_s_at 3.82 0.000714 MEGF11 multiple EGF-like-domains 11 210602_s_at 3.74 0.000408 CDH6 cadherin 6, type 2, K-cadherin (fetal kidney) 229947_at 3.62 0.00843 PI15 peptidase inhibitor 15 204006_s_at 3.39 0.00241 FCGR3A Fc fragment of IgG, low affinity IIIa, receptor (CD16a) 202238_s_at 3.29 0.00492 NNMT nicotinamide N-methyltransferase 202917_s_at 3.20 0.00369 S100A8 S100 calcium binding protein A8 215223_s_at 3.17 0.000516 SOD2 superoxide dismutase 2, mitochondrial 204627_s_at 3.04 0.00619 ITGB3 integrin, beta 3 (platelet glycoprotein IIIa, antigen CD61) 223217_s_at 2.99 0.00397 NFKBIZ nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta 231067_s_at 2.97 0.00681 AKAP12 A kinase (PRKA) anchor protein 12 224917_at 2.94 0.00256 VMP1/ mir-21likely ortholog -

Long Noncoding RNA NEAT1, Regulated by the EGFR Pathway
Published OnlineFirst November 14, 2017; DOI: 10.1158/1078-0432.CCR-17-0605 Biology of Human Tumors Clinical Cancer Research Long Noncoding RNA NEAT1, Regulated by the EGFR Pathway, Contributes to Glioblastoma Progression Through the WNT/b-Catenin Pathway by Scaffolding EZH2 Qun Chen1,2,3, Jinquan Cai1,2,3, Qixue Wang3,4, Yunfei Wang3,4, Mingyang Liu5, Jingxuan Yang5, Junhu Zhou3,4, Chunsheng Kang3,4, Min Li5, and Chuanlu Jiang1,2,3 Abstract Purpose: Long noncoding RNAs have been implicated in NFkB (p65) downstream of the EGFR pathway. Moreover, we gliomagenesis, but their mechanisms of action are mainly undoc- found that NEAT1 was critical for glioma cell growth and umented. Through public glioma mRNA expression data sets, we invasion by increasing b-catenin nuclear transport and down- found that NEAT1 was a potential oncogene. We systematically regulating ICAT, GSK3B, and Axin2. Taken together, we found analyzed the clinical significance and mechanism of NEAT1 in that NEAT1 could bind to EZH2 and mediate the trimethyla- glioblastoma. tion of H3K27 in their promoters. NEAT1 depletion also Experimental Design: Initially, we evaluated whether NEAT1 inhibited GBM cell growth and invasion in the intracranial expression levels could be regulated by EGFR pathway activity. We animal model. subsequently evaluated the effect of NEAT1 on the WNT/ Conclusions: The EGFR/NEAT1/EZH2/b-catenin axis serves as b-catenin pathway and its target binding gene. The animal model a critical effector of tumorigenesis and progression, suggesting supported the experimental findings. new therapeutic directions in glioblastoma. Clin Cancer Res; 1–12. Results: We found that NEAT1 levels were regulated by Ó2017 AACR. -
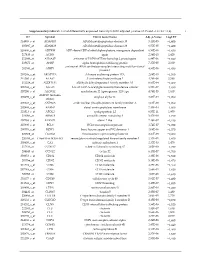
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

Long Non-Coding RNA Neat1 Regulates Adaptive Behavioural Response To
bioRxiv preprint doi: https://doi.org/10.1101/773333; this version posted September 23, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Long non-coding RNA Neat1 regulates adaptive behavioural response to stress in mice Abbreviated title: LncRNA Neat1 regulates adaptive behaviour in mice Michail S. Kukharsky1, Natalia N. Ninkina1, Haiyan An1,2, Vsevolod Telezhkin3, Wenbin Wei4,5, Camille Rabesahala de Meritens2, Johnathan Cooper-Knock4, Shinichi Nakagawa6, Tetsuro Hirose6, Vladimir L. Buchman1 and Tatyana A. Shelkovnikova1,2* 1School of Biosciences, Cardiff University, Cardiff, UK, CF10 3AX 2Medicines Discovery Institute, Cardiff University, Cardiff, UK, CF10 3AT 3School of Dental Sciences, Newcastle University, Newcastle upon Tyne, UK, NE2 4BW 4Sheffield Institute for Translational Neuroscience, University of Sheffield, Sheffield, UK, S10 2HQ 5Department of Biosciences, Durham University, Durham, UK, DH1 3LE 6Faculty of Pharmaceutical Sciences, Hokkaido University, Sapporo, Japan, 060-0812 *To whom correspondence should be addressed at Medicines Discovery Institute, Cardiff University, Main Building, Cardiff, CF10 3AT, United Kingdom Tel.: +44 (0)29 225 11072 E-mail: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/773333; this version posted September 23, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. -

Long Non-Coding Rnas Mechanisms of Action in HIV-1 Modulation and the Identification of Novel Therapeutic Targets
non-coding RNA Review Long Non-coding RNAs Mechanisms of Action in HIV-1 Modulation and the Identification of Novel Therapeutic Targets Roslyn M. Ray and Kevin V. Morris * The Center for Gene Therapy, The Beckman Research Institute, City of Hope Medical Center, Duarte, CA 91010, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-6262182839 Received: 8 October 2019; Accepted: 10 March 2020; Published: 13 March 2020 Abstract: This review aims to highlight the role of long non-coding RNAs in mediating human immunodeficiency virus (HIV-1) viral replication, latency, disease susceptibility and progression. In particular, we focus on identifying possible lncRNA targets and their purported mechanisms of action for future drug design or gene therapeutics. Keywords: HIV; lncRNA; transcription; antisense; epigenetic 1. Introduction The interplay between host gene expression and exposure to pathogens is a vital process that occurs in vivo. Cellular fate is determined through the differential expression of key pathways that are regulated and fine-tuned, in part, by long non-coding RNAs (lncRNAs). Manipulation of these lncRNAs by pathogens, genomic rearrangements or single nucleotide polymorphisms (SNPs) have been shown to exert great effects on disease outcomes [1–3]. Long non-coding RNAs are an important class of RNAs involved in orchestrating and manipulating transcriptional and post-transcriptional effects in vivo. Further, lncRNAs have been found to undergo purifying selection, suggesting their functional importance in biological processes [4]. LncRNAs are RNAs that are over 200 nucleotides in length and usually have no functional coding potential [5]. LncRNAs work in cis or trans, or both, to exert their gene modulatory effects.