Functional Regulation of Foxo1 in Neural Stem Cell Differentiation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Title of the Dissertation
UNIVERSITY OF CALIFORNIA SAN DIEGO Novel network-based integrated analyses of multi-omics data reveal new insights into CD8+ T cell differentiation and mouse embryogenesis A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Bioinformatics and Systems Biology by Kai Zhang Committee in charge: Professor Wei Wang, Chair Professor Pavel Arkadjevich Pevzner, Co-Chair Professor Vineet Bafna Professor Cornelis Murre Professor Bing Ren 2018 Copyright Kai Zhang, 2018 All rights reserved. The dissertation of Kai Zhang is approved, and it is accept- able in quality and form for publication on microfilm and electronically: Co-Chair Chair University of California San Diego 2018 iii EPIGRAPH The only true wisdom is in knowing you know nothing. —Socrates iv TABLE OF CONTENTS Signature Page ....................................... iii Epigraph ........................................... iv Table of Contents ...................................... v List of Figures ........................................ viii List of Tables ........................................ ix Acknowledgements ..................................... x Vita ............................................. xi Abstract of the Dissertation ................................. xii Chapter 1 General introduction ............................ 1 1.1 The applications of graph theory in bioinformatics ......... 1 1.2 Leveraging graphs to conduct integrated analyses .......... 4 1.3 References .............................. 6 Chapter 2 Systematic -

The DNA Methylation of FOXO3 and TP53 As a Blood Biomarker of Late
Yuan et al. J Transl Med (2020) 18:467 https://doi.org/10.1186/s12967-020-02643-y Journal of Translational Medicine RESEARCH Open Access The DNA methylation of FOXO3 and TP53 as a blood biomarker of late-onset asthma Lin Yuan1,2,3, Leyuan Wang2, Xizi Du2, Ling Qin1,3, Ming Yang4, Kai Zhou2, Mengping Wu2, Yu Yang2, Zhiyuan Zheng1,3, Yang Xiang2, Xiangping Qu2, Huijun Liu2, Xiaoqun Qin2 and Chi Liu1,2,5* Abstract Background: Late-onset asthma (LOA) is beginning to account for an increasing proportion of asthma patients, which is often underdiagnosed in the elderly. Studies on the possible relations between aging-related genes and LOA contribute to the diagnosis and treatment of LOA. Forkhead Box O3 (FOXO3) and TP53 are two classic aging-related genes. DNA methylation varies greatly with age which may play an important role in the pathogenesis of LOA. We supposed that the diferentially methylated sites of FOXO3 and TP53 associated with clinical phenotypes of LOA may be useful biomarkers for the early screening of LOA. Methods: The mRNA expression and DNA methylation of FOXO3 and TP53 in peripheral blood of 43 LOA patients (15 mild LOA, 15 moderate LOA and 13 severe LOA) and 60 healthy controls (HCs) were determined. The association of methylated sites with age was assessed by Cox regression to control the potential confounders. Then, the correlation between diferentially methylated sites (DMSs; p-value < 0.05) and clinical lung function in LOA patients was evalu- ated. Next, candidate DMSs combining with age were evaluated to predict LOA by receiver operating characteristic (ROC) analysis and principal components analysis (PCA). -

The E–Id Protein Axis Modulates the Activities of the PI3K–AKT–Mtorc1
Downloaded from genesdev.cshlp.org on October 6, 2021 - Published by Cold Spring Harbor Laboratory Press The E–Id protein axis modulates the activities of the PI3K–AKT–mTORC1– Hif1a and c-myc/p19Arf pathways to suppress innate variant TFH cell development, thymocyte expansion, and lymphomagenesis Masaki Miyazaki,1,8 Kazuko Miyazaki,1,8 Shuwen Chen,1 Vivek Chandra,1 Keisuke Wagatsuma,2 Yasutoshi Agata,2 Hans-Reimer Rodewald,3 Rintaro Saito,4 Aaron N. Chang,5 Nissi Varki,6 Hiroshi Kawamoto,7 and Cornelis Murre1 1Department of Molecular Biology, University of California at San Diego, La Jolla, California 92093, USA; 2Department of Biochemistry and Molecular Biology, Shiga University of Medical School, Shiga 520-2192, Japan; 3Division of Cellular Immunology, German Cancer Research Center, D-69120 Heidelberg, Germany; 4Department of Medicine, University of California at San Diego, La Jolla, California 92093, USA; 5Center for Computational Biology, Institute for Genomic Medicine, University of California at San Diego, La Jolla, California 92093, USA; 6Department of Pathology, University of California at San Diego, La Jolla, California 92093, USA; 7Department of Immunology, Institute for Frontier Medical Sciences, Kyoto University, Kyoto 606-8507, Japan It is now well established that the E and Id protein axis regulates multiple steps in lymphocyte development. However, it remains unknown how E and Id proteins mechanistically enforce and maintain the naı¨ve T-cell fate. Here we show that Id2 and Id3 suppressed the development and expansion of innate variant follicular helper T (TFH) cells. Innate variant TFH cells required major histocompatibility complex (MHC) class I-like signaling and were associated with germinal center B cells. -

Conserved Functional Motifs of the Nuclear Receptor Superfamily As Potential Pharmacological Targets
INTERNATIONAL JOURNAL OF EPIGenetiCS 1: 3, 2021 Conserved functional motifs of the nuclear receptor superfamily as potential pharmacological targets LOUIS PAPAGEORGIOU1, LIVIA SHALZI1, ASPASIA EFTHIMIADOU2, FLORA BACOPOULOU3, GEORGE P. CHROUSOS3,4, ELIAS ELIOPOULOS1 and DIMITRIOS VLACHAKIS1,3,4 1Laboratory of Genetics, Department of Biotechnology, School of Applied Biology and Biotechnology, Agricultural University of Athens, 11855 Athens; 2Department of Soil Science of Athens, Institute of Soil and Water Resources, Hellenic Agricultural Organization-Demeter, 14123 Lycovrisi; 3University Research Institute of Maternal and Child Health and Precision Medicine, and UNESCO Chair on Adolescent Health Care, National and Kapodistrian University of Athens, ‘Aghia Sophia’ Children's Hospital; 4Division of Endocrinology and Metabolism, Center of Clinical, Experimental Surgery and Translational Research, Biomedical Research Foundation of the Academy of Athens, 11527 Athens, Greece Received March 21, 2021; Accepted May 31, 2021 DOI: 10.3892/ije.2021.3 Abstract. Nuclear receptors (NRs) are one of the most diverse organ physiology, cell differentiation and homeostasis (1,2). In and well-reported family of proteins. They are involved in humans, only 48 members of the superfamily have been found numerous cellular processes as they play pivotal roles in and genetic mutations in these NRs have been proven to cause cell signaling and the cell cycle. The participation of NRs rare diseases, such as cancer, diabetes, rheumatoid arthritis, in various applications in medicine and biology has greatly asthma and hormone resistance syndromes (3). Bearing that attracted the interest of the pharmaceutical industry for the in mind and the fact that nuclear hormone receptors possess discovery of novel and/or improved drugs for the treatment internal pockets, that bind to hydrophobic, drug-like molecules, of several diseases, including cancer, diabetes or infertility. -

A Flexible Microfluidic System for Single-Cell Transcriptome Profiling
www.nature.com/scientificreports OPEN A fexible microfuidic system for single‑cell transcriptome profling elucidates phased transcriptional regulators of cell cycle Karen Davey1,7, Daniel Wong2,7, Filip Konopacki2, Eugene Kwa1, Tony Ly3, Heike Fiegler2 & Christopher R. Sibley 1,4,5,6* Single cell transcriptome profling has emerged as a breakthrough technology for the high‑resolution understanding of complex cellular systems. Here we report a fexible, cost‑efective and user‑ friendly droplet‑based microfuidics system, called the Nadia Instrument, that can allow 3′ mRNA capture of ~ 50,000 single cells or individual nuclei in a single run. The precise pressure‑based system demonstrates highly reproducible droplet size, low doublet rates and high mRNA capture efciencies that compare favorably in the feld. Moreover, when combined with the Nadia Innovate, the system can be transformed into an adaptable setup that enables use of diferent bufers and barcoded bead confgurations to facilitate diverse applications. Finally, by 3′ mRNA profling asynchronous human and mouse cells at diferent phases of the cell cycle, we demonstrate the system’s ability to readily distinguish distinct cell populations and infer underlying transcriptional regulatory networks. Notably this provided supportive evidence for multiple transcription factors that had little or no known link to the cell cycle (e.g. DRAP1, ZKSCAN1 and CEBPZ). In summary, the Nadia platform represents a promising and fexible technology for future transcriptomic studies, and other related applications, at cell resolution. Single cell transcriptome profling has recently emerged as a breakthrough technology for understanding how cellular heterogeneity contributes to complex biological systems. Indeed, cultured cells, microorganisms, biopsies, blood and other tissues can be rapidly profled for quantifcation of gene expression at cell resolution. -

A Simple Differentiation Protocol for Generation of Induced
1 Article, Special Issue "Induced Pluripotent Stem Cells in Neurodegenerative Diseases: Application for Therapy and 2 Disease Modeling" 3 A simple differentiation protocol for generation of 4 induced pluripotent stem cell-derived basal forebrain 5 cholinergic neurons for Alzheimer’s disease and 6 frontotemporal dementia disease modeling 7 Supplemental information 8 Method 1. Reprogramming and characterisation of MBE2960 healthy control iPSC line 9 The iPSCs were generated using skin fibroblasts obtained from subjects over the age of 18 years 10 by episomal method as described [40]. Briefly, reprogramming was performed on passage 8-10 11 fibroblasts by nucleofection (Lonza Amaxa Nucleofector) with episomal vectors expressing 12 OCT4, SOX2, KLF4, L-MYC, LIN28 and shRNA against p53 [41] in feeder- and serum- free 13 conditions using TeSR-E7 medium (Stemcell Technologies). Subsequently, reprogrammed 14 colonies were manually dissected to establish clonal cell lines [42]. Three clones were assessed 15 for pluripotency markers via immunocytochemistry (Figure S1A). The iPSC line was expanded 16 and characterised. Embryoid bodies were obtained as described [43] and using tri-lineage 17 differentiation kit (Stemcell Technologies). Germ layer differentiation was assessed by 18 immunochemistry (Figure S1B). Copy number variation (CNV) analysis of original fibroblasts 19 and iPSCs from MBE2960 (Figure S1C) was performed using Illumina HumanCore Beadchip 20 arrays as we described [40]. CNV analyses were performed using PennCNV and QuantiSNP with 21 default parameter settings [44,45]. Chromosomal aberrations were deemed to involve at least 10 22 contiguous single nucleotide polymorphisms (SNPs) or a genomic region spanning at least 1MB 23 [44,45]. The B allele frequency (BAF) and the log R ratio (LRR) were extracted from 24 GenomeStudio (Illumina) for representation (Figure S1D). -

Diagnosis of FOXG1 Syndrome Caused by Recurrent Balanced Chromosomal Rearrangements: Case Study and Literature Review Connor P
Craig et al. Mol Cytogenet (2020) 13:40 https://doi.org/10.1186/s13039-020-00506-1 CASE REPORT Open Access Diagnosis of FOXG1 syndrome caused by recurrent balanced chromosomal rearrangements: case study and literature review Connor P. Craig1,2, Emily Calamaro3, Chin‑To Fong3, Anwar M. Iqbal1, Alexander R. Paciorkowski3,4,5,6 and Bin Zhang1,3* Abstract Background: The FOXG1 gene plays a vital role in mammalian brain diferentiation and development. Intra‑ and intergenic mutations resulting in loss of function or altered expression of the FOXG1 gene cause FOXG1 syndrome. The hallmarks of this syndrome are severe developmental delay with absent verbal language, post‑natal growth restric‑ tion, post‑natal microcephaly, and a recognizable movement disorder characterized by chorea and dystonia. Case presentation: Here we describe a case of a 7‑year‑old male patient found to have a de novo balanced translo‑ cation between chromosome 3 at band 3q14.1 and chromosome 14 at band 14q12 via G‑banding chromosome and Fluorescence In Situ Hybridization (FISH) analyses. This rearrangement disrupts the proximity of FOXG1 to a previously described smallest region of deletion overlap (SRO), likely resulting in haploinsufciency. Conclusions: This case adds to the growing body of literature implicating chromosomal structural variants in the manifestation of this disorder and highlights the vital role of cis‑acting regulatory elements in the normal expression of this gene. Finally, we propose a protocol for refex FISH analysis to improve diagnostic efciency for patients with suspected FOXG1 syndrome. Keywords: FOXG1, Haploinsufciency, Postnatal microcephaly, FISH, Enhancer, Chromosomal rearrangement, Diagnosis Introduction brain development, with high levels of expression in the Te Forkhead Box G1 (FOXG1) gene [OMIM: 164874], developing fetal telencephalon [1–4]. -
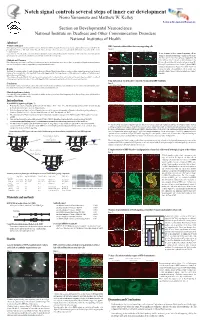
Notch Signal Controls Several Steps of Inner Ear Development Norio Yamamoto and Matthew W
NIDCD Notch signal controls several steps of inner ear development Norio Yamamoto and Matthew W. Kelley Section on Developmental Neuroscience Section on Developmental Neuroscience National Institute on Deafness and Other Communication Disorders National Institutes of Health Abstract Problem addressed RBP-J mutant cochleae did not have any supporting cells Notch signaling has been reported to contribute to inner ear development, however, its specific functions remain unclear, partly because of discrepancies between the phenotypes of mutant mice with single deletion of specific Notch related genes. These discrepancies are probably due to functional compensation by other Notch Figure 4 receptors or ligands. Foxg1 Cre;RBP-J floxed/+ Foxg1 Cre;RBP-J floxed/floxed A B C D To determine the effects of complete elimination of Notch signaling, we used a conditional knockout of the Rbpsuh gene. RBP-J protein is a critical transcriptional p27 Phalloidin p27 Phalloidin To test if mutant cochleae contained supporting cells we co-activator for all Notch molecules and thus deletion of this protein inhibits all Notch signaling. examined expression of supporting cell markers such as p27 and Prox1. On E17.5 p27 was expressed in supporting cells Methods and Measures ***** ***** under or between hair cells such as inner pharyngeal cells, Floxed Rbpsuh mice were crossed with Foxg1-Cre knock-in mice to delete the Rbpsuh gene in the inner ear. Inner ear phenotypes in Rbpsuh conditional knockout Deiter's cells and pillar cells (asterisks in figure 4 A and B). mice were determined at various developmental stages using immunohistochemistry. But no p27 expression was detected in those regions of RBP-J E Prox1 F Phalloidin G Prox1 H Phalloidin conditional knockout mice (Figure 4 C and D). -

Single-Cell Transcriptomics Characterizes Cell Types in the Subventricular Zone and Uncovers
bioRxiv preprint doi: https://doi.org/10.1101/365619; this version posted July 9, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Single-cell transcriptomics characterizes cell types in the subventricular zone and uncovers molecular defects underlying impaired adult neurogenesis Vera Zywitza1,+, Aristotelis Misios1,+, Lena Bunatyan2, Thomas E. Willnow2,*, and Nikolaus Rajewsky1,3,* 1 Laboratory for Systems Biology of Gene Regulatory Elements, Berlin Institute for Medical Systems Biology, Max Delbrück Center for Molecular Medicine, Robert-Rössle-Str. 10, Berlin-Buch, Germany 2 Molecular Cardiovascular Research, Max Delbrück Center for Molecular Medicine, Robert-Rössle- Str. 10, Berlin-Buch, Germany 3 Lead Contact + These authors contributed equally *Correspondence: [email protected]; [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/365619; this version posted July 9, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. SUMMARY Neural stem cells (NSCs) contribute to plasticity and repair of the adult brain. Niches harboring NSCs are crucial for regulating stem cell self-renewal and differentiation. We used single-cell RNA profiling to generate an unbiased molecular atlas of all cell types in the largest neurogenic niche of the adult mouse brain, the subventricular zone (SVZ). We characterized > 20 neural and non-neural cell types and gained insights into the dynamics of neurogenesis by predicting future cell states based on computational analysis of RNA kinetics. Furthermore, we apply our single-cell approach to mice lacking LRP2, an endocytic receptor required for SVZ maintenance. -

Regulation of PERK Expression by FOXO3: a Vulnerability of Drug- Resistant Cancer Cells
Oncogene (2019) 38:6382–6398 https://doi.org/10.1038/s41388-019-0890-7 ARTICLE Regulation of PERK expression by FOXO3: a vulnerability of drug- resistant cancer cells 1 1 1 1 1 Glowi Alasiri ● Yannasittha Jiramongkol ● Stefania Zona ● Lavender Y. -N. Fan ● Zimam Mahmud ● 2 2 1 Gyungyub Gong ● Hee Jin Lee ● Eric W. -F. Lam Received: 15 February 2019 / Revised: 28 June 2019 / Accepted: 2 July 2019 / Published online: 16 July 2019 © The Author(s) 2019. This article is published with open access Abstract The major impediment to effective cancer therapy has been the development of drug resistance. The tumour suppressive transcription factor FOXO3 promotes cell cycle arrest, senescence and cell death, and mediates the cytotoxic and cytostatic functions of cancer therapeutics. In consequence, FOXO3 is often downregulated as an adaptive response in cancer and particularly in chemotherapeutic drug-resistant cells. Consistently, we find that FOXO3 expression is attenuated in the drug- resistant MCF-7-EpiR and MCF-7-TaxR compared to the parental MCF-7 breast cancer cells. Using ChIP, short-interfering RNA (siRNA) knockdown, and overexpression assays as well as Foxo1/3/4−/− MEFs, we establish the endoplasmic reticulum 1234567890();,: 1234567890();,: (ER)-stress defence modulator PERK (eIF2AK3) as a direct downstream transcriptional target of FOXO3. In agreement, there is also a positive correlation between FOXO3 and PERK expression at the protein and RNA levels in breast cancer patient samples. We uncover that PERK expression is downregulated but its activity constitutively elevated in the drug- resistant cells. With this in mind, we exploit this adaptive response of low FOXO3 and PERK expression, and high PERK activity in drug-resistant breast cancer cells and show that these drug-resistant cells are specifically sensitive to PERK inhibition. -

Downloaded from Bioscientifica.Com at 09/26/2021 08:38:14PM Via Free Access
1 185 L C Gregory, P Gergics, OTX2 mutations in congenital 185:1 121–135 Clinical Study M Nakaguma and others hypopituitarism The phenotypic spectrum associated with OTX2 mutations in humans Louise C Gregory 1,*, Peter Gergics2,* , Marilena Nakaguma3,* , Hironori Bando2 , Giuseppa Patti1,4,5, Mark J McCabe1, Qing Fang 2, Qianyi Ma2 , Ayse Bilge Ozel2 , Jun Z Li2 , Michele Moreira Poina3, Alexander A L Jorge 3, Anna F Figueredo Benedetti3, Antonio M Lerario3, Ivo J P Arnhold3 , Berenice B Mendonca3 , Mohamad Maghnie4,5, Sally A Camper2 , Luciani R S Carvalho3 and Mehul T Dattani1 1Section of Molecular Basis of Rare Disease, Genetics and Genomic Medicine Research & Teaching Department, UCL Great Ormond Street Institute of Child Health, London, UK, 2Department of Human Genetics, University of Michigan, Correspondence Ann Arbor, Michigan, USA, 3Developmental Endocrinology Unit, Hospital das Clinicas da Faculdade de Medicina da should be addressed Universidade de São Paulo, São Paulo, Brazil, 4Department of Pediatrics, IRCCS Istituto Giannina Gaslini and to S A Camper or M T 5Department of Neuroscience, Rehabilitation, Ophthalmology, Genetics, Maternal and Child Health, University of Dattani Genova, Genova, Italy Email *(L C Gregory, P Gergics and M Nakaguma contributed equally to this work) [email protected] or [email protected] Abstract Objective: The transcription factor OTX2 is implicated in ocular, craniofacial, and pituitary development. Design: We aimed to establish the contribution of OTX2 mutations in congenital hypopituitarism patients with/without eye abnormalities, study functional consequences, and establish OTX2 expression in the human brain, with a view to investigate the mechanism of action. Methods: We screened patients from the UK (n = 103), international centres (n = 24), and Brazil (n = 282); 145 were within the septo-optic dysplasia spectrum, and 264 had no eye phenotype. -
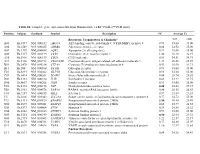
Table S1. Complete Gene Expression Data from Human Diabetes RT² Profiler™ PCR Array Receptors, Transporters & Channels* A
Table S1. Complete gene expression data from Human Diabetes RT² Profiler™ PCR Array Position Unigene GenBank Symbol Description FC Average Ct Receptors, Transporters & Channels* NGT GDM A01 Hs,5447 NM_000352 ABCC8 ATP-binding cassette, sub-family C (CFTR/MRP), member 8 0.93 35.00 35.00 A04 0Hs,2549 NM_000025 ADRB3 Adrenergic, beta-3-, receptor 0.88 34.92 35.00 A07 Hs,1307 NM_000486 AQP2 Aquaporin 2 (collecting duct) 0.93 35.00 35.00 A09 30Hs,5117 NM_001123 CCR2 Chemokine (C-C motif) receptor 2 1.00 26.28 26.17 A10 94Hs,5916 396NM_006139 CD28 CD28 molecule 0.81 34.51 34.71 A11 29Hs,5126 NM_001712 CEACAM1 Carcinoembryonic antigen-related cell adhesion molecule 1 1.31 26.08 25.59 B01 82Hs,2478 NM_005214 CTLA4 (biliaryCytotoxic glycoprotein) T-lymphocyte -associated protein 4 0.53 30.90 31.71 B11 24Hs,208 NM_000160 GCGR Glucagon receptor 0.93 35.00 35.00 C01 Hs,3891 NM_002062 GLP1R Glucagon-like peptide 1 receptor 0.93 35.00 35.00 C07 03Hs,6434 NM_000201 ICAM1 Intercellular adhesion molecule 1 0.84 28.74 28.89 D02 47Hs,5134 NM_000418 IL4R Interleukin 4 receptor 0.64 34.22 34.75 D06 57Hs,4657 NM_000208 INSR Insulin receptor 0.93 35.00 35.00 E05 44Hs,4312 NM_006178 NSF N-ethylmaleimide-sensitive factor 0.48 28.42 29.37 F08 79Hs,2961 NM_004578 RAB4A RAB4A, member RAS oncogene family 0.88 20.55 20.63 F10 69Hs,7287 NM_000655 SELL Selectin L 0.97 23.89 23.83 F11 56Hs,3806 NM_001042 SLC2A4 Solute carrier family 2 (facilitated glucose transporter), member 4 0.77 34.72 35.00 F12 91Hs,5111 NM_003825 SNAP23 Synaptosomal-associated protein, 23kDa 3.90