FCRL4 Rabbit Pab
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Characteristics of B Cell-Associated Gene Expression in Patients With
MOLECULAR MEDICINE REPORTS 13: 4113-4121, 2016 Characteristics of B cell-associated gene expression in patients with coronary artery disease WENWEN YAN*, HAOMING SONG*, JINFA JIANG, WENJUN XU, ZHU GONG, QIANGLIN DUAN, CHUANGRONG LI, YUAN XIE and LEMIN WANG Department of Internal Medicine, Division of Cardiology, Tongji Hospital, Tongji University School of Medicine, Shanghai 200065, P.R. China Received May 19, 2015; Accepted February 12, 2016 DOI: 10.3892/mmr.2016.5029 Abstract. The current study aimed to identify differentially with the two other groups. Additionally the gene expression expressed B cell-associated genes in peripheral blood mono- levels of B cell regulatory genes were measured. In patients nuclear cells and observe the changes in B cell activation at with AMI, CR1, LILRB2, LILRB3 and VAV1 mRNA expres- different stages of coronary artery disease. Groups of patients sion levels were statistically increased, whereas, CS1 and IL4I1 with acute myocardial infarction (AMI) and stable angina (SA), mRNAs were significantly reduced compared with the SA and as well as healthy volunteers, were recruited into the study control groups. There was no statistically significant difference (n=20 per group). Whole human genome microarray analysis in B cell-associated gene expression levels between patients was performed to examine the expression of B cell-associated with SA and the control group. The present study identified the genes among these three groups. The mRNA expression levels downregulation of genes associated with BCRs, B2 cells and of 60 genes associated with B cell activity and regulation were B cell regulators in patients with AMI, indicating a weakened measured using reverse transcription-quantitative polymerase T cell-B cell interaction and reduced B2 cell activation during chain reaction. -

Usbiological Datasheet
FCRL4, CT (FCRL4, FCRH4, IFGP2, IRTA1, Fc receptor- like protein 4, Fc receptor homolog 4, IFGP family protein 2, Immune receptor translocation-associated protein 1, CD307d) (FITC) Catalog number 035592-FITC Supplier United States Biological This gene encodes a member of the immunoglobulin receptor superfamily and is one of several Fc receptor-like glycoproteins clustered on the long arm of chromosome 1. The encoded protein has four extracellular C2-type immunoglobulin domains, a transmembrane domain and a cytoplasmic domain that contains three immune-receptor tyrosine-based inhibitory motifs. This protein may play a role in the function of memory B-cells in the epithelia. Aberrations in the chromosomal region encoding this gene are associated with non-Hodgkin lymphoma and multiple myeloma. Applications Suitable for use in Western Blot, FLISA and Flow Cytometry (Not Tested). Other applications have not been tested. Recommended Dilution Optimal dilutions to be determined by the researcher. Storage and Stability Store product at 4°C if to be used immediately within two weeks. For long-term storage, aliquot to avoid repeated freezing and thawing and store at -20°C. Aliquots are stable at -20°C for 12 months after receipt. Dilute required amount only prior to immediate use. Further dilutions can be made in assay buffer. Caution: FITC conjugates are sensitive to light. For maximum recovery of product, centrifuge the original vial after thawing and prior to removing the cap. Note Applications are based on unconjugated antibody. Immunogen KLH-conjugated synthetic peptide mapping to a fragment of residues within amino acids 480-507 in the C-terminal region of human FCRL4. -

Unifying the Genomics-Based Classes of Cancer Fusion Gene Partners: Large Cancer Fusion Genes Are Evolutionarily Conserved
CANCER GENOMICS & PROTEOMICS 9: 389-396 (2012) Unifying the Genomics-based Classes of Cancer Fusion Gene Partners: Large Cancer Fusion Genes Are Evolutionarily Conserved LIBIA M. PAVA, DANIEL T. MORTON, REN CHEN and GEORGE BLANCK Department of Molecular Medicine, Morsani College of Medicine, University of South Florida, Tampa, FL, U.S.A. Abstract. Background: Genes that fuse to cause cancer have fusion of NPM and ALK in anaplastic large-cell lymphoma been studied to determine molecular bases for proliferation, to (3); ABL and BCR in chronic myelogenous leukemia (CML) develop diagnostic tools, and as targets for drugs. To facilitate (4, 5); and C-MYC and IgH in Burkitt’s lymphoma in (6), identification of additional, cancer fusion genes, following among many others. The detection and understanding of the observation of a chromosomal translocation, we have ABL-BCR fusion protein, which stimulates unregulated cell characterized the genomic features of the fusion gene partners. division and leads to leukemia, led to the development of Previous work indicated that cancer fusion gene partners, are Gleevec, a drug able to block the ATP-binding site of the either large or evolutionarily conserved in comparison to the tyrosine kinase domain of ABL-BCR, halting CML (7). This neighboring genes in the region of a chromosomal extraordinary success has led to the hope of designing drugs translocation. These results raised the question of whether targeted against other cancer fusion proteins. large cancer fusion gene partners were also evolutionarily There are about 50,000 unstudied translocations, raising conserved. Methods and Results: We developed two methods the question of whether that information can continue to be for quantifying evolutionary conservation values, allowing the used to facilitate the identification of fusion genes. -

Agricultural University of Athens
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΩΝ ΖΩΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΖΩΙΚΗΣ ΠΑΡΑΓΩΓΗΣ ΕΡΓΑΣΤΗΡΙΟ ΓΕΝΙΚΗΣ ΚΑΙ ΕΙΔΙΚΗΣ ΖΩΟΤΕΧΝΙΑΣ ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ ΑΘΗΝΑ 2020 ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Genome-wide association analysis and gene network analysis for (re)production traits in commercial broilers ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ Τριμελής Επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Επταμελής εξεταστική επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Πηνελόπη Μπεμπέλη (Καθ. ΓΠΑ) Δημήτριος Βλαχάκης (Επ. Καθ. ΓΠΑ) Ευάγγελος Ζωίδης (Επ.Καθ. ΓΠΑ) Γεώργιος Θεοδώρου (Επ.Καθ. ΓΠΑ) 2 Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Περίληψη Σκοπός της παρούσας διδακτορικής διατριβής ήταν ο εντοπισμός γενετικών δεικτών και υποψηφίων γονιδίων που εμπλέκονται στο γενετικό έλεγχο δύο τυπικών πολυγονιδιακών ιδιοτήτων σε κρεοπαραγωγικά ορνίθια. Μία ιδιότητα σχετίζεται με την ανάπτυξη (σωματικό βάρος στις 35 ημέρες, ΣΒ) και η άλλη με την αναπαραγωγική -

Adaptation to Disease Exposure in Indigenous Southern African Populations
Adaptation to disease exposure in indigenous southern African populations Katharine Owers Degree project in biology, Master of science (2 years), 2012 Examensarbete i biologi 30 hp till masterexamen, 2012 Biology Education Centre and Department of Evolutionary Biology, Uppsala University Supervisor: Mattias Jakobsson Abstract Infectious diseases have impacted humans throughout history. While contemporary diseases can be studied with modern methods, allowing rapid collection and dissemination of information about their effects on populations, studies of the effects of diseases in historical times do not have those advantages. Some historical disease events, such as the plague that struck Europe in the 14th century, are relatively well-understood, but in other cases we have little information on the diseases and their impacts. Such is the case for the waves of disease, both known and speculated, caused by migrations into southern Africa of other African groups and European colonists. Southern indigenous societies did not keep written records, so we must reconstruct their history from European reports, oral histories, and information from archeology and linguistics. Recent advances in genetics, however, provide new sources of information on population history, structure, and selection. I analyzed single nucleotide polymorphisms (SNPs) from two indigenous southern African Khoe-San populations with differing levels of contact with the immigrant groups—the ≠Khomani with abundant such contact and the isolated Ju/’hoansi—to search for evidence of adaptation in the genome due to selection pressure from introduced infectious diseases. Two approaches were used. First, I located regions of the genome likely under selection according to a combination of test statistics based on haplotype homozygosity and population differentiation and then examined those regions for enrichment of genes related to the immune system. -

The Acquisition of Human B Cell Memory in Response to Plasmodium Falciparum Malaria
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations Summer 2010 The Acquisition of Human B Cell Memory in Response to Plasmodium Falciparum Malaria Gretchen E. Weiss University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Biology Commons, Immunity Commons, Immunology of Infectious Disease Commons, Medical Immunology Commons, and the Parasitology Commons Recommended Citation Weiss, Gretchen E., "The Acquisition of Human B Cell Memory in Response to Plasmodium Falciparum Malaria" (2010). Publicly Accessible Penn Dissertations. 210. https://repository.upenn.edu/edissertations/210 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/210 For more information, please contact [email protected]. The Acquisition of Human B Cell Memory in Response to Plasmodium Falciparum Malaria Abstract Immunity to Plasmodium falciparum (Pf), the most deadly agent of malaria, is only acquired after years of repeated infections and appears to wane rapidly without ongoing exposure. Antibodies (Abs) are central to malaria immunity, yet little is known about the B‐cell biology that underlies Pf‐specific humoral immunity. To address this gap in our knowledge we carried out a year‐long prospective study of the acquisition and maintenance of long‐lived plasma cells (LLPCs) and memory B cells (MBCs) in 225 individuals aged two to twenty‐five years in Mali, in an area of intense seasonal transmission. Using protein microarrays containing approximately 25% of the Pf proteome we determined that Pf‐specific Abs were acquired only gradually, in a stepwise fashion over years of Pf exposure. Pf‐specific Ab levels were significantly boosted each eary during the transmission season but the majority of these Abs were short lived and were lost over the subsequent six month period of no transmission. -

WO 2016/141169 Al 9 September 2016 (09.09.2016) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2016/141169 Al 9 September 2016 (09.09.2016) P O P C T (51) International Patent Classification: (74) Agent: AKHAVAN, Ramin; Caris MPI, Inc., 6655 N. A61K 31/335 (2006.01) G01N 33/50 (2006.01) MacArthur Blvd., Irving, TX 75039 (US). A61K 39/395 (2006.01) G01N 33/53 (2006.01) (81) Designated States (unless otherwise indicated, for every C12Q 1/68 (2006.01) G01N 33/00 (2006.01) kind of national protection available): AE, AG, AL, AM, C40B 30/04 (2006.01) AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, (21) International Application Number: BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, PCT/US2016/020657 DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, (22) International Filing Date: KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, 3 March 2016 (03.03.2016) MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, (25) Filing Language: English PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, (26) Publication Language: English TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (30) Priority Data: (84) Designated States (unless otherwise indicated, for every 62/127,769 3 March 2015 (03.03.2015) kind of regional protection available): ARIPO (BW, GH, 62/167,659 28 May 2015 (28.05.2015) GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, (71) Applicant: CARIS MPI, INC. -
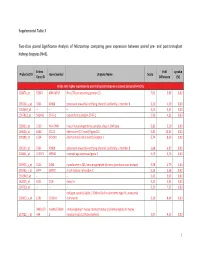
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at -

Characterization of Human FCRL4-Positive B Cells
Characterization of human FCRL4-positive B cells Michel Jourdan, Nicolas Robert, Maïlys Cren, Coraline Thibaut, Christophe Duperray, Alboukadel Kassambara, Michel Cogné, Karin Tarte, Bernard Klein, Jerome Moreaux To cite this version: Michel Jourdan, Nicolas Robert, Maïlys Cren, Coraline Thibaut, Christophe Duperray, et al.. Char- acterization of human FCRL4-positive B cells. PLoS ONE, Public Library of Science, 2017, 12 (6), pp.e0179793. 10.1371/journal.pone.0179793. hal-01560458 HAL Id: hal-01560458 https://hal-univ-rennes1.archives-ouvertes.fr/hal-01560458 Submitted on 28 Jun 2018 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License RESEARCH ARTICLE Characterization of human FCRL4-positive B cells Michel Jourdan1, Nicolas Robert2, Maïlys Cren3, Coraline Thibaut1, Christophe Duperray4, Alboukadel Kassambara1,2, Michel Cogne 5, Karin Tarte6,7, Bernard Klein1,2,8³, JeÂroà me Moreaux1,2,8³* 1 Institute of Human Genetics, UMR 9002 CNRS-UM, Montpellier, France, 2 CHU Montpellier, Laboratory for Monitoring Innovative Therapies, Department of Biological Hematology, Montpellier, France, 3 INSERM, U1040, Montpellier, France, 4 Cytometry IRB, Montpellier Rio Imaging, Montpellier, France, 5 CNRS UMR 7276, Universite de Limoges, Limoges, France, 6 PoÃle Cellules et Tissus, CHU Rennes, Rennes, France, a1111111111 7 INSERM, U917, Rennes, France, 8 Universite Montpellier 1, UFR MeÂdecine, Montpellier, France a1111111111 a1111111111 ³ These authors share last authorship. -

FCRL4 (N) Antibody, Rabbit Polyclonal
Order: (888)-282-5810 (Phone) (818)-707-0392 (Fax) [email protected] Web: www.Abiocode.com FCRL4 (N) Antibody, Rabbit Polyclonal Cat#: R2921-1 Lot#: Refer to vial Quantity: 100 ul Application: WB Predicted I Observed M.W.: 57 I 60 kDa Uniprot ID: Q96PJ5 Background: Fc receptor-like protein 4 (FCRL4) is a member of the immunoglobulin receptor superfamily and is one of several Fc receptor-like glycoproteins clustered on the long arm of chromosome 1. FCRL4 has four extracellular C2-type immunoglobulin domains, a transmembrane domain and a cytoplasmic domain that contains three immune-receptor tyrosine-based inhibitory motifs. FCRL4 may play a role in the function of memory B-cells in the epithelia. Aberrations in the chromosomal region encoding this gene are associated with non-Hodgkin lymphoma and multiple myeloma. Other Names: Fc receptor-like protein 4, FcR-like protein 4, FcRL4, Fc receptor homolog 4, FcRH4, IFGP family protein 2, hIFGP2, Immune receptor translocation-associated protein 1, CD307d, FCRH4, IFGP2, IRTA1 Source and Purity: Rabbit polyclonal antibodies were produced by immunizing animals with a GST-fusion protein containing the N-terminal region of human FCRL4. Antibodies were purified by affinity purification using immunogen. Storage Buffer and Condition: Supplied in 1 x PBS (pH 7.4), 100 ug/ml BSA, 40% Glycerol, 0.01% NaN3. Store at -20 °C. Stable for 6 months from date of receipt. Species Specificity: Human, Mouse Tested Applications: WB: 1:500-1:2,000 (detect endogenous protein*) *: The apparent protein size on WB may be different from the calculated M.W. due to modifications. -

Increase of MZB1 in B Cells in Systemic Lupus Erythematosus
Miyagawa-Hayashino et al. Arthritis Research & Therapy (2018) 20:13 DOI 10.1186/s13075-018-1511-5 RESEARCHARTICLE Open Access Increase of MZB1 in B cells in systemic lupus erythematosus: proteomic analysis of biopsied lymph nodes Aya Miyagawa-Hayashino1,2,7*, Hajime Yoshifuji3, Koji Kitagori1,3, Shinji Ito4, Takuma Oku1,5, Yoshitaka Hirayama1,5, Adeeb Salah2, Toshiki Nakajima3, Kaori Kiso6, Norishige Yamada6, Hironori Haga2 and Tatsuaki Tsuruyama6 Abstract Background: Systemic lupus erythematosus (SLE) is a prototypical autoimmune disease in which dysregulation of B cells has been recognized. Here, we searched for potential biomarkers of SLE using liquid chromatography-tandem mass spectrometry (LC-MS). Methods: Lymph nodes from SLE patients and controls were analyzed by LC-MS. To validate the identified molecules, immunoblotting and immunohistochemistry were performed and B cells from SLE patients were analyzed by quantitative RT-PCR. B-cell subsets from NZB/W F1 mice, which exhibit autoimmune disease resembling human SLE, were analyzed by flow cytometry. Endoplasmic reticulum (ER) stress was induced by tunicamycin and the serum concentration of anti-dsDNA antibodies was determined by ELISA. TUNEL methods and immunoblotting were used to assess the effect of tunicamycin. Results: MZB1, which comprises part of a B-cell-specific ER chaperone complex and is a key player in antibody secretion, was one of the differentially expressed proteins identified by LC-MS and confirmed by immunoblotting. Immunohistochemically, larger numbers of MZB1+ cells were located mainly in interfollicular areas and scattered in germinal centers in specimens from SLE patients compared with those from controls. MZB1 colocalized with CD138+ plasma cells and IRTA1+ marginal zone B cells.