Chlamydomonas Reinhardtii
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Noelia Díaz Blanco
Effects of environmental factors on the gonadal transcriptome of European sea bass (Dicentrarchus labrax), juvenile growth and sex ratios Noelia Díaz Blanco Ph.D. thesis 2014 Submitted in partial fulfillment of the requirements for the Ph.D. degree from the Universitat Pompeu Fabra (UPF). This work has been carried out at the Group of Biology of Reproduction (GBR), at the Department of Renewable Marine Resources of the Institute of Marine Sciences (ICM-CSIC). Thesis supervisor: Dr. Francesc Piferrer Professor d’Investigació Institut de Ciències del Mar (ICM-CSIC) i ii A mis padres A Xavi iii iv Acknowledgements This thesis has been made possible by the support of many people who in one way or another, many times unknowingly, gave me the strength to overcome this "long and winding road". First of all, I would like to thank my supervisor, Dr. Francesc Piferrer, for his patience, guidance and wise advice throughout all this Ph.D. experience. But above all, for the trust he placed on me almost seven years ago when he offered me the opportunity to be part of his team. Thanks also for teaching me how to question always everything, for sharing with me your enthusiasm for science and for giving me the opportunity of learning from you by participating in many projects, collaborations and scientific meetings. I am also thankful to my colleagues (former and present Group of Biology of Reproduction members) for your support and encouragement throughout this journey. To the “exGBRs”, thanks for helping me with my first steps into this world. Working as an undergrad with you Dr. -

Identification of Novel Dna Damage Response Genes Using Functional Genomics
IDENTIFICATION OF NOVEL DNA DAMAGE RESPONSE GENES USING FUNCTIONAL GENOMICS by Michael Chang A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Graduate Department of Biochemistry University of Toronto © Copyright by Michael Chang (2005) Identification of novel DNA damage response genes using functional genomics Doctor of Philosophy, 2005; Michael Chang; Department of Biochemistry, University of Toronto ABSTRACT The genetic information required for life is stored within molecules of DNA. This DNA is under constant attack as a result of normal cellular metabolic processes, as well as exposure to genotoxic agents. DNA damage left unrepaired can result in mutations that alter the genetic information encoded within DNA. Cells have consequently evolved complex pathways to combat damage to their DNA. Defects in the cellular response to DNA damage can result in genomic instability, a hallmark of cancer cells. Identifying all the components required for this response remains an important step in fully elucidating the molecular mechanisms involved. I used functional genomic approaches to identify genes required for the DNA damage response in Saccharomyces cerevisiae. I conducted a screen to identify genes required for resistance to a DNA damaging agent, methyl methanesulfonate, and identified several poorly characterized genes that are necessary for proper S phase progression in the presence of DNA damage. Among the genes identified, ESC4/RTT107 has since been shown to be essential for the resumption of DNA replication after DNA damage. Using genome-wide genetic interaction screens to identify genes that are required for viability in the absence of MUS81 and MMS4, two genes required for resistance to DNA damage, I helped identify ELG1, deletion of which causes DNA replication defects, genomic instability, and an inability to properly recover from DNA damage during S phase. -

The Network Organization of Cancer-Associated Protein
The Network Organization of Cancer-associated Protein Complexes in SUBJECT AREAS: PROTEOME Human Tissues INFORMATICS COMPUTATIONAL SCIENCE Jing Zhao1, Sang Hoon Lee2,3, Mikael Huss4,5 & Petter Holme2,6 ONCOGENESIS COMPLEX NETWORKS 1Department of Mathematics, Logistical Engineering University, Chongqing, China, 2IceLab, Department of Physics, Umea˚ University, Umea˚, Sweden, 3Oxford Centre for Industrial and Applied Mathematics, Mathematical Institute, University of Oxford, Oxford, United Kingdom, 4Science for Life Laboratory Stockholm, Solna, Sweden, 5Department of Biochemistry and Biophysics, Received Stockholm University, Stockholm, Sweden, 6Department of Energy Science, Sungkyunkwan University, Suwon, Korea. 24 October 2012 Accepted Differential gene expression profiles for detecting disease genes have been studied intensively in systems 7 March 2013 biology. However, it is known that various biological functions achieved by proteins follow from the ability of the protein to form complexes by physically binding to each other. In other words, the functional units are Published often protein complexes rather than individual proteins. Thus, we seek to replace the perspective of 9 April 2013 disease-related genes by disease-related complexes, exemplifying with data on 39 human solid tissue cancers and their original normal tissues. To obtain the differential abundance levels of protein complexes, we apply an optimization algorithm to genome-wide differential expression data. From the differential abundance of Correspondence and complexes, we extract tissue- and cancer-selective complexes, and investigate their relevance to cancer. The method is supported by a clustering tendency of bipartite cancer-complex relationships, as well as a more requests for materials concrete and realistic approach to disease-related proteomics. should be addressed to J.Z. -

The 23 and Me Research Team, Morris, JA, Kemp, JP, Youlten, SE
the 23 and Me Research Team, Morris, J. A., Kemp, J. P., Youlten, S. E., Laurent, L., Logan, J. G., Chai, R. C., Vulpescu, N. A., Forgetta, V., Kleinman, A., Mohanty, S. T., Sergio, C. M., Quinn, J., Nguyen- Yamamoto, L., Luco, A. L., Vijay, J., Simon, M. M., Pramatarova, A., Medina-Gomez, C., ... Evans, D. M. (2019). An atlas of genetic influences on osteoporosis in humans and mice. Nature Genetics, 51, 258–266. https://doi.org/10.1038/s41588-018-0302-x Peer reviewed version Link to published version (if available): 10.1038/s41588-018-0302-x Link to publication record in Explore Bristol Research PDF-document This is the author accepted manuscript (AAM). The final published version (version of record) is available online via Springer Nature at https://www.nature.com/articles/s41588-018-0302-x . Please refer to any applicable terms of use of the publisher. University of Bristol - Explore Bristol Research General rights This document is made available in accordance with publisher policies. Please cite only the published version using the reference above. Full terms of use are available: http://www.bristol.ac.uk/red/research-policy/pure/user-guides/ebr-terms/ An Atlas of Human and Murine Genetic Influences on Osteoporosis John A. Morris1,2§, John P. Kemp3,4§, Scott E. Youlten5, Laetitia Laurent2, John G. Logan6, Ryan Chai5, Nicholas A. Vulpescu7, Vincenzo Forgetta2, Aaron Kleinman8, Sindhu Mohanty5, C. Marcelo Sergio5, Julian Quinn5, Loan Nguyen-Yamamoto9, Aimee Lee Luco9, Jinchu Vijay10, Marie-Michelle Simon10, Albena Pramatarova10, Carolina Medina-Gomez11, Katerina Trajanoska11, Elena J. Ghirardello6, Natalie C. -

DNA Replication and Sister Chromatid Cohesion 1 (DSCC1)
Journal of Cancer 2019, Vol. 10 6142 Ivyspring International Publisher Journal of Cancer 2019; 10(24): 6142-6153. doi: 10.7150/jca.32339 Research Paper DNA Replication and Sister Chromatid Cohesion 1 (DSCC1) of the Replication Factor Complex CTF18- RFC is Critical for Colon Cancer Cell Growth Jong-Tae Kim1*, Hee Jun Cho1*, Sang Yoon Park1, Byung Moo Oh1,2, Yo Sep Hwang1,2, Kyoung Eun Baek1, Young-Ha Lee3, Hee Cheol Kim4 and Hee Gu Lee1,2 1. Immunotherapy Research Center, Korea Research Institute of Bioscience and Biotechnology (KRIBB), Daejeon, Republic of Korea. 2. Department of Biomolecular Science, University of Science and Technology (UST), Daejeon, Republic of Korea. 3. Department of Infection Biology, Chungnam National University School of Medicine, Daejeon, Republic of Korea. 4. Department of Surgery, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Republic of Korea. *These authors contributed equally to this work. Corresponding authors: Hee Cheol Kim, M.D., Ph.D., Department of Surgery, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Republic of Korea. E-mail: [email protected] or Hee Gu Lee, Ph.D., Immunotherapy Research Center, Korea Research Institute of Bioscience and Biotechnology, Daejeon 34141, Republic of Korea. Tel: +82-42-860-4182; Fax: +82-42-860-4593; E-mail: [email protected] © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2018.12.17; Accepted: 2019.08.26; Published: 2019.10.15 Abstract DNA replication and sister chromatid cohesion 1 (DSCC1) combines with chromosome transmission-fidelity protein 18 (CTF18) to form a CTF18-DSCC1-CTF8 (CTF18-1-8) module, which in combination with CTF18-replication factor C (RFC) acts as a proliferating cell nuclear antigen (PCNA) loader during DNA replication-associated processes. -
CHTF18 Monoclonal Antibody (M01), Clone 1F5
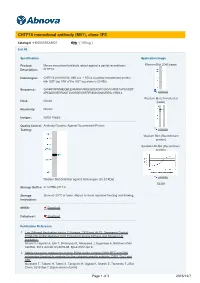
CHTF18 monoclonal antibody (M01), clone 1F5 Catalog # : H00063922-M01 規格 : [ 100 ug ] List All Specification Application Image Product Mouse monoclonal antibody raised against a partial recombinant Western Blot (Cell lysate) Description: CHTF18. Immunogen: CHTF18 (AAH18184, 886 a.a. ~ 975 a.a) partial recombinant protein with GST tag. MW of the GST tag alone is 26 KDa. Sequence: GVHRPAPRNHEQRLEHIMRRAAREEQPEKDFFGRVVVRSTAVPSAGDT APEQDSVERRMGTAVGRSEVWFRFNEGVSNAVRRSLYIRDLL enlarge Western Blot (Transfected Host: Mouse lysate) Reactivity: Human Isotype: IgG2a Kappa Quality Control Antibody Reactive Against Recombinant Protein. Testing: enlarge Western Blot (Recombinant protein) Sandwich ELISA (Recombinant protein) enlarge Western Blot detection against Immunogen (35.53 KDa) . ELISA Storage Buffer: In 1x PBS, pH 7.4 Storage Store at -20°C or lower. Aliquot to avoid repeated freezing and thawing. Instruction: MSDS: Download Datasheet: Download Publication Reference 1. Two Different Replication Factor C Proteins, Ctf18 and RFC1, Separately Control PCNA-CRL4Cdt2-Mediated Cdt1 Proteolysis during S Phase and following UV Irradiation. Shiomi Y, Hayashi A, Ishii T, Shinmyozu K, Nakayama J, Sugasawa K, Nishitani H.Mol Cell Biol. 2012 Jun;32(12):2279-88. Epub 2012 Apr 9. 2. Stable interaction between the human PCNA loader complex Ctf18-RFC and DNA polymerase {epsilon} is mediated by the cohesion specific subunits, Ctf18, Dcc1 and Ctf8. Murakami T, Takano R, Takeo S, Taniguchi R, Ogawa K, Ohashi E, Tsurimoto T.J Biol Chem. 2010 Sep 7. [Epub ahead of print] Page 1 of 3 2016/12/7 Applications Western Blot (Cell lysate) CHTF18 monoclonal antibody (M01), clone 1F5 Western Blot analysis of CHTF18 expression in HeLa ( Cat # L013V1 ). Protocol Download Western Blot (Transfected lysate) Western Blot analysis of CHTF18 expression in transfected 293T cell line by CHTF18 monoclonal antibody (M01), clone 1F5. -

S-Phase Checkpoint Genes Safeguard High-Fidelity Sister Chromatid Cohesion□D Cheryl D
Molecular Biology of the Cell Vol. 15, 1724–1735, April 2004 S-Phase Checkpoint Genes Safeguard High-Fidelity Sister Chromatid Cohesion□D Cheryl D. Warren,* D. Mark Eckley,* Marina S. Lee,* Joseph S. Hanna,* Adam Hughes,* Brian Peyser,* Chunfa Jie,* Rafael Irizarry,† and Forrest A. Spencer*‡ *McKusick-Nathans Institute of Genetic Medicine, Ross 850, Johns Hopkins University School of Medicine, Baltimore, Maryland 21205; and †Department of Biostatistics, Johns Hopkins University Bloomberg School of Public Health, Baltimore, Maryland 21205 Submitted September 2, 2003; Revised December 10, 2003; Accepted December 23, 2003 Monitoring Editor: Douglas Koshland Cohesion establishment and maintenance are carried out by proteins that modify the activity of Cohesin, an essential complex that holds sister chromatids together. Constituents of the replication fork, such as the DNA polymerase ␣-binding protein Ctf4, contribute to cohesion in ways that are poorly understood. To identify additional cohesion components, we analyzed a ctf4⌬ synthetic lethal screen performed on microarrays. We focused on a subset of ctf4⌬- interacting genes with genetic instability of their own. Our analyses revealed that 17 previously studied genes are also necessary for the maintenance of robust association of sisters in metaphase. Among these were subunits of the MRX complex, which forms a molecular structure similar to Cohesin. Further investigation indicated that the MRX complex did not contribute to metaphase cohesion independent of Cohesin, although an additional role may be contributed by XRS2. In general, results from the screen indicated a sister chromatid cohesion role for a specific subset of genes that function in DNA replication and repair. This subset is particularly enriched for genes that support the S-phase checkpoint. -

Cshperspect-REP-A015727 Table3 1..10
Table 3. Nomenclature for proteins and protein complexes in different organisms Mammals Budding yeast Fission yeast Flies Plants Archaea Bacteria Prereplication complex assembly H. sapiens S. cerevisiae S. pombe D. melanogaster A. thaliana S. solfataricus E. coli Hs Sc Sp Dm At Sso Eco ORC ORC ORC ORC ORC [Orc1/Cdc6]-1, 2, 3 DnaA Orc1/p97 Orc1/p104 Orc1/Orp1/p81 Orc1/p103 Orc1a, Orc1b Orc2/p82 Orc2/p71 Orc2/Orp2/p61 Orc2/p69 Orc2 Orc3/p66 Orc3/p72 Orc3/Orp3/p80 Orc3/Lat/p82 Orc3 Orc4/p50 Orc4/p61 Orc4/Orp4/p109 Orc4/p52 Orc4 Orc5L/p50 Orc5/p55 Orc5/Orp5/p52 Orc5/p52 Orc5 Orc6/p28 Orc6/p50 Orc6/Orp6/p31 Orc6/p29 Orc6 Cdc6 Cdc6 Cdc18 Cdc6 Cdc6a, Cdc6b [Orc1/Cdc6]-1, 2, 3 DnaC Cdt1/Rlf-B Tah11/Sid2/Cdt1 Cdt1 Dup/Cdt1 Cdt1a, Cdt1b Whip g MCM helicase MCM helicase MCM helicase MCM helicase MCM helicase Mcm DnaB Mcm2 Mcm2 Mcm2/Nda1/Cdc19 Mcm2 Mcm2 Mcm3 Mcm3 Mcm3 Mcm3 Mcm3 Mcm4 Mcm4/Cdc54 Mcm4/Cdc21 Mcm4/Dpa Mcm4 Mcm5 Mcm5/Cdc46/Bob1 Mcm5/Nda4 Mcm5 Mcm5 Mcm6 Mcm6 Mcm6/Mis5 Mcm6 Mcm6 Mcm7 Mcm7/Cdc47 Mcm7 Mcm7 Mcm7/Prolifera Gmnn/Geminin Geminin Mcm9 Mcm9 Hbo1 Chm/Hat1 Ham1 Ham2 DiaA Ihfa Ihfb Fis SeqA Replication fork assembly Hs Sc Sp Dm At Sso Eco Mcm8 Rec/Mcm8 Mcm8 Mcm10 Mcm10/Dna43 Mcm10/Cdc23 Mcm10 Mcm10 DDK complex DDK complex DDK complex DDK complex Cdc7 Cdc7 Hsk1 l(1)G0148 Hsk1-like 1 Dbf4/Ask Dbf4 Dfp1/Him1/Rad35 Chif/chiffon Drf1 Continued 2 Replication fork assembly (Continued ) Hs Sc Sp Dm At Sso Eco CDK complex CDK complex CDK complex CDK complex CDK complex Cdk1 Cdc28/Cdk1 Cdc2/Cdk1 Cdc2 CdkA Cdk2 Cdc2c CcnA1, A2 CycA CycA1, A2, -

High-Resolution Acgh and Expression Profiling Identifies a Novel Genomic
Open Access Research2007ChinetVolume al. 8, Issue 10, Article R215 High-resolution aCGH and expression profiling identifies a novel genomic subtype of ER negative breast cancer Suet F Chin¤*, Andrew E Teschendorff¤*†, John C Marioni¤†§, Yanzhong Wang*, Nuno L Barbosa-Morais†, Natalie P Thorne†§, Jose L Costa#, Sarah E Pinder¥, Mark A van de Wiel**††, Andrew R Green¶, Ian O Ellis¶, Peggy L Porter‡‡, Simon Tavar醧, James D Brenton‡, Bauke Ylstra# and Carlos Caldas*¥ Addresses: *Breast Cancer Functional Genomics, Cancer Research UK Cambridge Research Institute and Department of Oncology University of Cambridge, Li Ka-Shing Centre, Robinson Way, Cambridge CB2 0RE, UK. †Computational Biology Group, Cancer Research UK Cambridge Research Institute and Department of Oncology University of Cambridge, Li Ka-Shing Centre, Robinson Way, Cambridge CB2 0RE, UK. ‡Functional Genomics of Drug Resistance, Cancer Research UK Cambridge Research Institute and Department of Oncology University of Cambridge, Li Ka-Shing Centre, Robinson Way, Cambridge CB2 0RE, UK. §Computational Biology Group, Department of Applied Mathematics and Theoretical Physics, University of Cambridge, Centre for Mathematical Sciences, Wilberforce Road, Cambridge CB3 0WA, UK. ¶Histopathology, Nottingham City Hospital NHS Trust and University of Nottingham, Nottingham NG5 1PB, UK. ¥Cambridge Breast Unit, Addenbrookes Hospital, Cambridge University Hospitals NHS Foundation Trust, Hills Road, Cambridge, UK. #Department of Pathology, VU University Medical Center, PO Box 7057, 1007MB Amsterdam, The Netherlands. **Department of Biostatistics, VU University Medical Center, PO Box 7057, 1007MB Amsterdam, The Netherlands. ††Department of Mathematics, Vrije Universiteit, Amsterdam, Netherlands. ‡‡Division of Human Biology, Division of Public Health Sciences, Fred Hutchinson Cancer Research Center, Seattle, WA 98109, USA. -

Program and Abstracts Book
16th International Conference on the Cell and Molecular Biology of Chlamydomonas June 8-13, 2014 Asilomar Conference Center, Pacific Grove, CA, USA Program and Abstracts 16th International Conference on the Cell and Molecular Biology of Chlamydomonas June 8-13, 2014 Asilomar Conference Grounds Pacific Grove, California Program and Abstracts Organizers: Kris Niyogi, University of California, Berkeley Winfield Sale, Emory University Marilyn Kobayashi, University of California, Berkeley Advisory Committee: José Luis Crespo, CSIC - Universidad de Sevilla Susan Dutcher, Washington University School of Medicine Arthur Grossman, Carnegie Institution for Science Sabeeha Merchant, University of California, Los Angeles Jun Minagawa, National Institute for Basic Biology David Mitchell, SUNY Upstate Medical University Rachael Morgan-Kiss, Miami University Michael Schroda, University of Kaiserslautern Carolyn Silflow, University of Minnesota James Umen, Donald Danforth Plant Science Center Chia-Lin Wei, DOE Joint Genome Institute William Zerges, Concordia University 1 2 TABLE OF CONTENTS General Information ............................................................................................................................ 4 Exhibitors ............................................................................................................................................. 4 Schedule of Events ............................................................................................................................... 5 Plenary Session Listings -

CHTF18 Polyclonal Antibody Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP55350
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 CHTF18 Polyclonal Antibody Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP55350 Specification CHTF18 Polyclonal Antibody - Product Information Application WB, IHC-P, IHC-F, IF, ICC Primary Accession Q8WVB6 Reactivity Rat Host Rabbit Clonality Polyclonal Calculated MW 107383 CHTF18 Polyclonal Antibody - Additional Information Gene ID 63922 Other Names Chromosome transmission fidelity protein 18 homolog, hCTF18, CHL12, CHTF18, C16orf41, CTF18 Format 0.01M TBS(pH7.4) with 1% BSA, 0.09% (W/V) sodium azide and 50% Glyce Storage Store at -20 ℃ for one year. Avoid repeated freeze/thaw cycles. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 ℃. CHTF18 Polyclonal Antibody - Protein Information Name CHTF18 Synonyms C16orf41, CTF18 Function Chromosome cohesion factor involved in sister chromatid cohesion and fidelity of chromosome transmission. Component of one of the cell nuclear antigen loader complexes, CTF18-replication factor C (CTF18-RFC), which consists of CTF18, CTF8, DCC1, RFC2, RFC3, RFC4 and RFC5. Page 1/2 10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 The CTF18-RFC complex binds to single-stranded and primed DNAs and has weak ATPase activity that is stimulated by the presence of primed DNA, replication protein A (RPA) and by proliferating cell nuclear antigen (PCNA). The CTF18-RFC complex catalyzes the ATP- dependent loading of PCNA onto primed and gapped DNA. Interacts with and stimulates DNA polymerase POLH. -

Unequal Evolutionary Conservation of Human Protein Interactions In
Open Access Research2007BrownVolume and 8, Issue Jurisica 5, Article R95 Unequal evolutionary conservation of human protein interactions comment in interologous networks Kevin R Brown*† and Igor Jurisica*†‡ Addresses: *Department of Medical Biophysics, University of Toronto, Toronto, Canada M5G 1L7. †Ontario Cancer Institute, Toronto Medical Discovery Tower, Toronto, Canada M5G 1L7. ‡Department of Computer Science, University of Toronto, Toronto, Canada M5G 1L71. Correspondence: Igor Jurisica. Email: [email protected] reviews Published: 29 May 2007 Received: 16 November 2006 Revised: 2 March 2007 Genome Biology 2007, 8:R95 (doi:10.1186/gb-2007-8-5-r95) Accepted: 29 May 2007 The electronic version of this article is the complete one and can be found online at http://genomebiology.com/2007/8/5/R95 © 2007 Brown and Jurisica; licensee BioMed Central Ltd. This is an open access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which reports permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Conservation<p>Theorganisms, conservation revealing of protein that of interactionsprotein- proteinprotein complexes interaction are preferentially networks can conserved, be examined and that by mapping such conservation human proteins can yield to yeastbiological and otherinsights.</p> model Abstract deposited research Background: Protein-protein interaction (PPI) networks have been transferred between organisms using interologs, allowing model organisms to supplement the interactomes of higher eukaryotes. However, the conservation of various network components has not been fully explored. Unequal conservation of certain network components may limit the ability to fully expand the target interactomes using interologs.