Identi Cation of Drought-Responsive Hub Genes and Their Related
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Loss-Of-Function Mutations in RAB18 Cause Warburg Micro Syndrome
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector REPORT Loss-of-Function Mutations in RAB18 Cause Warburg Micro Syndrome Danai Bem,1 Shin-Ichiro Yoshimura,2,12 Ricardo Nunes-Bastos,2 Frances F. Bond,3 Manju A. Kurian,1,13 Fatima Rahman,1 Mark T.W. Handley,4 Yavor Hadzhiev,1 Imran Masood,5 Ania A. Straatman-Iwanowska,1,13 Andrew R. Cullinane,1,14 Alisdair McNeill,1,3,15 Shanaz S. Pasha,1 Gail A. Kirby,1 Katharine Foster,6 Zubair Ahmed,7 Jenny E. Morton,3 Denise Williams,3 John M. Graham,8 William B. Dobyns,9 Lydie Burglen,10 John R. Ainsworth,11 Paul Gissen,1,13 Ferenc Mu¨ller,1 Eamonn R. Maher,1,3 Francis A. Barr,2 and Irene A. Aligianis1,3,16,* Warburg Micro syndrome and Martsolf syndrome are heterogenous autosomal-recessive developmental disorders characterized by brain, eye, and endocrine abnormalities. Previously, identification of mutations in RAB3GAP1 and RAB3GAP2 in both these syndromes implicated dysregulation of the RAB3 cycle (which controls calcium-mediated exocytosis of neurotransmitters and hormones) in disease pathogenesis. RAB3GAP1 and RAB3GAP2 encode the catalytic and noncatalytic subunits of the hetrodimeric enzyme RAB3GAP (RAB3GTPase-activating protein), a key regulator of the RAB3 cycle. We performed autozygosity mapping in five consanguineous families without RAB3GAP1/2 mutations and identified loss-of-function mutations in RAB18. A c.71T > A (p.Leu24Gln) founder mutation was identified in four Pakistani families, and a homozygous exon 2 deletion (predicted to result in a frameshift) was found in the fifth family. -

The A–Z of Zika Drug Discovery, Drug Discov Today (2018)
Drug Discovery Today Volume 00, Number 00 June 2018 REVIEWS Teaser Zika clinical outcomes might be nefarious impacting newborns for a lifetime. There is still no drug available to cure Zika. We provide guidance to help understand and advance the search for a cure. KEYNOTE REVIEW – The A Z of Zika drug discovery Reviews 1 1 1 Melina Mottin graduated Melina Mottin , Joyce V.V.B. Borba , Rodolpho C. Braga , in pharmacy and earned her 2,3 4 Master’s degree and PhD at Pedro H.M. Torres , Matheus C. Martini , University of Campinas 4 5 (UNICAMP), Brazil. The Jose Luiz Proenca-Modena , Carla C. Judice , main focus of her studies 5 6 7 was to apply molecular Fabio T.M. Costa , Sean Ekins , Alexander L. Perryman and dynamics simulations and 1,5 other computational Carolina Horta Andrade strategies for a nuclear receptor and its association with drugs and coregulatory proteins, implicated in diabetes. 1 LabMol – Laboratory for Molecular Modeling and Drug Design, Faculdade de Farmacia, Universidade Federal de Melina is currently Research Associate of the OpenZika project, at Federal University of Goias, mainly applying Goias – UFG, Goiânia, GO 74605-170, Brazil 2 virtual screening based on docking and QSAR models to Instituto Oswaldo Cruz, Fundação Oswaldo Cruz, Rio de Janeiro, RJ 21040-900, Brazil 3 select drug candidates against Zika virus. Department of Biochemistry, University of Cambridge, 80 Tennis Court Road, Cambridge CB2 1GA, UK 4 Alexander L. Perryman Laboratory of Emerging Viruses (LEVE), Department of Genetics, Evolution, Microbiology and Immunology, earned his PhD in Andy Institute of Biology, UNICAMP, Campinas, SP 13083-864, Brazil 5 McCammon’s lab at UCSD. -

Rab18 Promotes Lipid Droplet (LD) Growth by Tethering the ER to Lds Through SNARE and NRZ Interactions
Published Online: 24 January, 2018 | Supp Info: http://doi.org/10.1083/jcb.201704184 Article Downloaded from jcb.rupress.org on August 7, 2018 Rab18 promotes lipid droplet (LD) growth by tethering the ER to LDs through SNARE and NRZ interactions Dijin Xu,1* Yuqi Li,1* Lizhen Wu,1* Ying Li,1 Dongyu Zhao,1 Jinhai Yu,1 Tuozhi Huang,1 Charles Ferguson,2 Robert G. Parton,2,3 Hongyuan Yang,4 and Peng Li1 1State Key Laboratory of Membrane Biology, Tsinghua-Peking Center for Life Sciences, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing, China 2Institute for Molecular Bioscience and 3Centre for Microscopy and Microanalysis, University of Queensland, Brisbane, Australia 4School of Biotechnology and Biomolecular Sciences, University of New South Wales, Sydney, Australia Lipid incorporation from endoplasmic reticulum (ER) to lipid droplet (LD) is important in controlling LD growth and intra- cellular lipid homeostasis. However, the molecular link mediating ER and LD cross talk remains elusive. Here, we identi- fied Rab18 as an important Rab guanosine triphosphatase in controlling LD growth and maturation.Rab18 deficiency resulted in a drastically reduced number of mature LDs and decreased lipid storage, and was accompanied by increased ER stress. Rab3GAP1/2, the GEF of Rab18, promoted LD growth by activating and targeting Rab18 to LDs. LD-associated Rab18 bound specifically to the ER-associated NAG-RINT1-ZW10 (NRZ) tethering complex and their associated SNAREs (Syntaxin18, Use1, BNIP1), resulting in the recruitment of ER to LD and the formation of direct ER–LD contact. Cells with defects in the NRZ/SNA RE complex function showed reduced LD growth and lipid storage. -

Convergent Functional Genomics of Schizophrenia: from Comprehensive Understanding to Genetic Risk Prediction
Molecular Psychiatry (2012) 17, 887 -- 905 & 2012 Macmillan Publishers Limited All rights reserved 1359-4184/12 www.nature.com/mp IMMEDIATE COMMUNICATION Convergent functional genomics of schizophrenia: from comprehensive understanding to genetic risk prediction M Ayalew1,2,9, H Le-Niculescu1,9, DF Levey1, N Jain1, B Changala1, SD Patel1, E Winiger1, A Breier1, A Shekhar1, R Amdur3, D Koller4, JI Nurnberger1, A Corvin5, M Geyer6, MT Tsuang6, D Salomon7, NJ Schork7, AH Fanous3, MC O’Donovan8 and AB Niculescu1,2 We have used a translational convergent functional genomics (CFG) approach to identify and prioritize genes involved in schizophrenia, by gene-level integration of genome-wide association study data with other genetic and gene expression studies in humans and animal models. Using this polyevidence scoring and pathway analyses, we identify top genes (DISC1, TCF4, MBP, MOBP, NCAM1, NRCAM, NDUFV2, RAB18, as well as ADCYAP1, BDNF, CNR1, COMT, DRD2, DTNBP1, GAD1, GRIA1, GRIN2B, HTR2A, NRG1, RELN, SNAP-25, TNIK), brain development, myelination, cell adhesion, glutamate receptor signaling, G-protein-- coupled receptor signaling and cAMP-mediated signaling as key to pathophysiology and as targets for therapeutic intervention. Overall, the data are consistent with a model of disrupted connectivity in schizophrenia, resulting from the effects of neurodevelopmental environmental stress on a background of genetic vulnerability. In addition, we show how the top candidate genes identified by CFG can be used to generate a genetic risk prediction score (GRPS) to aid schizophrenia diagnostics, with predictive ability in independent cohorts. The GRPS also differentiates classic age of onset schizophrenia from early onset and late-onset disease. -

Avian Binocularity and Adaptation to Nocturnal Environments: Genomic Insights Froma Highly Derived Visual Phenotype Rui Borges Universidade Do Porto - Portugal
Nova Southeastern University NSUWorks Biology Faculty Articles Department of Biological Sciences 8-22-2019 Avian Binocularity and Adaptation to Nocturnal Environments: Genomic Insights froma Highly Derived Visual Phenotype Rui Borges Universidade do Porto - Portugal Joao Fonseca Universidade do Porto - Portugal Cidalia Gomes Universidade do Porto - Portugal Warren E. Johnson Smithsonian Institution Stephen James O'Brien St. Petersburg State University - Russia; Nova Southeastern University, [email protected] See next page for additional authors Follow this and additional works at: https://nsuworks.nova.edu/cnso_bio_facarticles Part of the Biology Commons NSUWorks Citation Borges, Rui; Joao Fonseca; Cidalia Gomes; Warren E. Johnson; Stephen James O'Brien; Guojie Zhang; M. Thomas P. Gilbert; Erich D. Jarvis; and Agostinho Antunes. 2019. "Avian Binocularity and Adaptation to Nocturnal Environments: Genomic Insights froma Highly Derived Visual Phenotype." Genome Biology and Evolution 11, (8): 2244-2255. doi:10.1093/gbe/evz111. This Article is brought to you for free and open access by the Department of Biological Sciences at NSUWorks. It has been accepted for inclusion in Biology Faculty Articles by an authorized administrator of NSUWorks. For more information, please contact [email protected]. Authors Rui Borges, Joao Fonseca, Cidalia Gomes, Warren E. Johnson, Stephen James O'Brien, Guojie Zhang, M. Thomas P. Gilbert, Erich D. Jarvis, and Agostinho Antunes This article is available at NSUWorks: https://nsuworks.nova.edu/cnso_bio_facarticles/982 GBE Avian Binocularity and Adaptation to Nocturnal Environments: Genomic Insights from a Highly Derived Visual Downloaded from https://academic.oup.com/gbe/article-abstract/11/8/2244/5544263 by Nova Southeastern University/HPD Library user on 16 September 2019 Phenotype Rui Borges1,2,Joao~ Fonseca1,Cidalia Gomes1, Warren E. -

Large Homozygous RAB3GAP1 Gene Microdeletion Causes Warburg
Picker-Minh et al. Orphanet Journal of Rare Diseases 2014, 9:113 http://www.ojrd.com/content/9/1/113 LETTER TO THE EDITOR Open Access Large homozygous RAB3GAP1 gene microdeletion causes Warburg Micro Syndrome 1 Sylvie Picker-Minh1,2,3, Andreas Busche4, Britta Hartmann4, Birgit Spors5, Eva Klopocki6,7, Christoph Hübner1, Denise Horn6 and Angela M Kaindl1,2,3* Abstract Warburg micro syndrome (WARBM) is a genetic heterogeneous disease characterized by microcephaly, intellectual disability, brain, ocular, and endocrine anomalies. WARBM1-4 can be caused by biallelic mutations of the RAB3GAP1 (RAB3 GTPase-activating protein 1), RAB3GAP2, RAB18 (RAS-associated protein RAB18), or TBC1D20 (TBC1 domain protein, member 20) gene, respectively. Here, we delineate the so far largest intragenic homozygous RAB3GAP1 microdeletion. Despite the size of the RAB3GAP1 gene deletion, the patient phenotype is mainly consistent with that of other WARBM1 patients, supporting strongly the theory that WARBM1 is caused by a loss of RAB3GAP1 function. We further highlight osteopenia as a feature of WARBM1. Keywords: RAB3GAP1, WARBM, Warburg micro syndrome, Microcephaly, Intellectual disability, Congenital cataract, Array CGH Letter to the editor protein-function [1,2,4-7], putatively explaining the lack of Warburg micro syndrome (WARBM) is a rare autosomal a genotype-phenotype correlation. We here report the lar- recessive disorder characterized by neurodevelopmental gest RAB3GAP1 gene microdeletion to date in patients abnormalities such as congenital or postnatal microcephaly, with WARBM1 and compare their phenotype with that of severe intellectual disability, pachy- or polymicrogyria, other WARBM1 patients. The two index patients were and hypoplasia/agenesis of the corpus callosum as well as born at term without complications as the first and second ocular manifestations including congenital cataract, micro- child of healthy, consanguineous parents of Kurdish- cornea, microphthalmia, and optic atrophy [1-3]. -

D82b54407ece9642da45e12eee
Oxford Medical Case Reports, 2020;4,1–3 doi: 10.1093/omcr/omaa031 Case Report CASE REPORT Novel mutation in the RAB3GAP1 gene, the first diagnosed Warburg Micro syndrome case in Syria Soubhi Tenawi1,†, Rawan Al Khudari1,†,* and Diana Alasmar2 1Faculty of Medicine, Damascus University, Damascus, Syria, 2Professor of Inborn Errors of Metabolism, Pediatrics Department, Damascus University, Damascus, Syria *Correspondence address. Faculty of Medicine, Damascus University, Damascus, Syria. Tel: +963992336336; E-mail: [email protected] Abstract Warburg Micro syndrome is a rare autosomal recessive disease due to mutation in the RAB3GAP1, RAB3GAP2, RAB18 and TBC1D20 genes. It is commonly seen in consanguineous marriages, characterized by optic (microcornea, microphthalmia, congenital cataracts), neurologic )microcephaly, corpus callosum hypoplasia, severe mental retardation( and hypogonadism; some non-typical findings could be present (cardiomyopathy, peripheral neuropathy). We report a novel homozygous mutation in the RAB3GAP1 gene in a 7-month-old boy from healthy nonconsanguineous parents from the same village in Syria, with bilateral congenital cataracts, hypogonadism, muscular hypotonia and severe developmental delay. Whole exome sequencing (WES) showed a homozygous mutation in the c.2195del p.(Pro732Glnfs∗6) in exon 19 of the RAB3GAP1 gene, which is likely pathogenic and correlates with Warburg Micro syndrome type 1. INTRODUCTION eye and brain [4]. Here, we report a novel RAB3GAP1 homozygous mutation in a child with WARBM. Warburg Micro syndrome (WARBM) is a rare autosomal reces- sive disorder characterized by postnatal growth retardation, hypoplasia of the corpus callosum, microcephalus, delayed CASE REPORT motor development, severe intellectual disability, microph- thalmia, microcornea, congenital cataracts, optic atrophy and A 7-month-old boy presented to Children’s University Hospi- hypogonadism [1]. -
Primepcr™Assay Validation Report
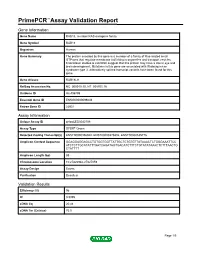
PrimePCR™Assay Validation Report Gene Information Gene Name RAB18, member RAS oncogene family Gene Symbol RAB18 Organism Human Gene Summary The protein encoded by this gene is a member of a family of Ras-related small GTPases that regulate membrane trafficking in organelles and transport vesicles. Knockdown studies is zebrafish suggest that this protein may have a role in eye and brain development. Mutations in this gene are associated with Warburg micro syndrome type 3. Alternatively spliced transcript variants have been found for this gene. Gene Aliases RAB18LI1 RefSeq Accession No. NC_000010.10, NT_008705.16 UniGene ID Hs.406799 Ensembl Gene ID ENSG00000099246 Entrez Gene ID 22931 Assay Information Unique Assay ID qHsaCED0042708 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000356940, ENST00000375802, ENST00000535776 Amplicon Context Sequence GGAGGAGGAGCCTGTGGTGGTTATTGCTCTGTGTTATAAACTCTGGGAAATTCC ATCTCTTGCATATTTGATCAGATAGTGACATCTTTCTGTATATAAACTCTTTAACTG CTATTTT Amplicon Length (bp) 88 Chromosome Location 10:27826942-27827059 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 96 R2 0.9995 cDNA Cq 20.44 cDNA Tm (Celsius) 76.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq 24.45 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report RAB18, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard -

A SARS-Cov-2 Protein Interaction Map Reveals Targets for Drug Repurposing
Article A SARS-CoV-2 protein interaction map reveals targets for drug repurposing https://doi.org/10.1038/s41586-020-2286-9 A list of authors and affiliations appears at the end of the paper Received: 23 March 2020 Accepted: 22 April 2020 A newly described coronavirus named severe acute respiratory syndrome Published online: 30 April 2020 coronavirus 2 (SARS-CoV-2), which is the causative agent of coronavirus disease 2019 (COVID-19), has infected over 2.3 million people, led to the death of more than Check for updates 160,000 individuals and caused worldwide social and economic disruption1,2. There are no antiviral drugs with proven clinical efcacy for the treatment of COVID-19, nor are there any vaccines that prevent infection with SARS-CoV-2, and eforts to develop drugs and vaccines are hampered by the limited knowledge of the molecular details of how SARS-CoV-2 infects cells. Here we cloned, tagged and expressed 26 of the 29 SARS-CoV-2 proteins in human cells and identifed the human proteins that physically associated with each of the SARS-CoV-2 proteins using afnity-purifcation mass spectrometry, identifying 332 high-confdence protein–protein interactions between SARS-CoV-2 and human proteins. Among these, we identify 66 druggable human proteins or host factors targeted by 69 compounds (of which, 29 drugs are approved by the US Food and Drug Administration, 12 are in clinical trials and 28 are preclinical compounds). We screened a subset of these in multiple viral assays and found two sets of pharmacological agents that displayed antiviral activity: inhibitors of mRNA translation and predicted regulators of the sigma-1 and sigma-2 receptors. -

RAB18 Gene RAB18, Member RAS Oncogene Family
RAB18 gene RAB18, member RAS oncogene family Normal Function The RAB18 gene provides instructions for producing the RAB18 protein, which functions as a GTPase. Often referred to as molecular switches, GTPases can be turned on and off. They are turned on (active) when they are attached (bound) to a molecule called GTP and are turned off (inactive) when they are bound to another molecule called GDP. When active, RAB18 is involved in a process called vesicle trafficking, which moves proteins and other molecules within cells in sac-like structures called vesicles. RAB18 regulates the movement of substances between compartments in cells and the storage and release of fats (lipids) by structures called lipid droplets. The protein also appears to play a role in a process called autophagy, which helps clear unneeded materials from cells. RAB18 is important for the organization of a cell structure called the endoplasmic reticulum, which is involved in protein processing and transport. Health Conditions Related to Genetic Changes RAB18 deficiency At least five mutations in the RAB18 gene have been found to cause Warburg micro syndrome, which is the most severe of the disorders caused by RAB18 deficiency. Warburg micro syndrome is characterized by multiple eye abnormalities, vision impairment, severe intellectual disability, and a reduction of the hormones that direct sexual development (hypogonadotropic hypogonadism). The RAB18 gene mutations that cause Warburg micro syndrome eliminate the function of the RAB18 protein. It is unclear how a shortage -

Analysis of Mammalian Gene Function Through Broad-Based Phenotypic
ARTICLES Analysis of mammalian gene function through broad-based phenotypic screens across a consortium of mouse clinics The function of the majority of genes in the mouse and human genomes remains unknown. The mouse embryonic stem cell knockout resource provides a basis for the characterization of relationships between genes and phenotypes. The EUMODIC consortium developed and validated robust methodologies for the broad-based phenotyping of knockouts through a pipeline comprising 20 disease-oriented platforms. We developed new statistical methods for pipeline design and data analysis aimed at detecting reproducible phenotypes with high power. We acquired phenotype data from 449 mutant alleles, representing 320 unique genes, of which half had no previous functional annotation. We captured data from over 27,000 mice, finding that 83% of the mutant lines are phenodeviant, with 65% demonstrating pleiotropy. Surprisingly, we found significant differences in phenotype annotation according to zygosity. New phenotypes were uncovered for many genes with previously unknown function, providing a powerful basis for hypothesis generation and further investigation in diverse systems. Phenotypic annotations of knockout mutants have been generated for (EMPReSS) database10 catalogs the standard operating procedures about a third of the genes in the mouse genome1. However, the way in (SOPs) that were developed, including operational details and the which the phenotype is screened is often dependent on the expertise and parameters measured. More recently, a major single-center effort interests of the investigator, and in only a few cases has a broad-based to analyze several hundred knockout lines through a phenotyping assessment of phenotype been undertaken that encompassed the analy- pipeline has illuminated the pleiotropy that can be found and the sis of developmental, biochemical, physiological and organ systems2–4. -

A Novel Mouse Model of Warburg Micro Syndrome Reveals Roles for RAB18 in Eye Development and Organisation of the Neuronal Cytoskeleton Sarah M
© 2014. Published by The Company of Biologists Ltd | Disease Models & Mechanisms (2014) 7, 711-722 doi:10.1242/dmm.015222 RESEARCH ARTICLE A novel mouse model of Warburg Micro syndrome reveals roles for RAB18 in eye development and organisation of the neuronal cytoskeleton Sarah M. Carpanini1,2, Lisa McKie1, Derek Thomson3, Ann K. Wright3, Sarah L. Gordon4, Sarah L. Roche3, Mark T. Handley1, Harris Morrison1, David Brownstein5, Thomas M. Wishart2,3, Michael A. Cousin4, Thomas H. Gillingwater3,*,‡, Irene A. Aligianis1,* and Ian J. Jackson1,2,*,‡ ABSTRACT INTRODUCTION Mutations in RAB18 have been shown to cause the heterogeneous Warburg Micro syndrome (WARBM) is a heterogenous autosomal autosomal recessive disorder Warburg Micro syndrome (WARBM). recessive disorder (Warburg et al., 1993). Loss-of-function Individuals with WARBM present with a range of clinical symptoms, mutations have been identified in RAB3GAP1 (Aligianis et al., including ocular and neurological abnormalities. However, the 2005), RAB3GAP2 (Borck et al., 2011), RAB18 (Bem et al., 2011) underlying cellular and molecular pathogenesis of the disorder and TBC1D20 (Liegel et al., 2013), and these mutations result in remains unclear, largely owing to the lack of any robust animal models clinically indistinguishable phenotypes. The clinical features of that phenocopy both the ocular and neurological features of the WARBM are primarily ocular and neurological (Abdel-Salam et al., disease. We report here the generation and characterisation of a novel 2007; Derbent et al., 2004; Handley et al., 2013): affected children Rab18-mutant mouse model of WARBM. Rab18-mutant mice are have visual impairment and eye abnormalities – including viable and fertile. They present with congenital nuclear cataracts and congenital bilateral cataracts, microphthalmia, microcornea (<10 atonic pupils, recapitulating the characteristic ocular features that are mm diameter) and small atonic pupils that do not react to dark or associated with WARBM.