Human Transcription Factor Protein Interaction Networks
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Table S1. List of Proteins in the BAHD1 Interactome
Table S1. List of proteins in the BAHD1 interactome BAHD1 nuclear partners found in this work yeast two-hybrid screen Name Description Function Reference (a) Chromatin adapters HP1α (CBX5) chromobox homolog 5 (HP1 alpha) Binds histone H3 methylated on lysine 9 and chromatin-associated proteins (20-23) HP1β (CBX1) chromobox homolog 1 (HP1 beta) Binds histone H3 methylated on lysine 9 and chromatin-associated proteins HP1γ (CBX3) chromobox homolog 3 (HP1 gamma) Binds histone H3 methylated on lysine 9 and chromatin-associated proteins MBD1 methyl-CpG binding domain protein 1 Binds methylated CpG dinucleotide and chromatin-associated proteins (22, 24-26) Chromatin modification enzymes CHD1 chromodomain helicase DNA binding protein 1 ATP-dependent chromatin remodeling activity (27-28) HDAC5 histone deacetylase 5 Histone deacetylase activity (23,29,30) SETDB1 (ESET;KMT1E) SET domain, bifurcated 1 Histone-lysine N-methyltransferase activity (31-34) Transcription factors GTF3C2 general transcription factor IIIC, polypeptide 2, beta 110kDa Required for RNA polymerase III-mediated transcription HEYL (Hey3) hairy/enhancer-of-split related with YRPW motif-like DNA-binding transcription factor with basic helix-loop-helix domain (35) KLF10 (TIEG1) Kruppel-like factor 10 DNA-binding transcription factor with C2H2 zinc finger domain (36) NR2F1 (COUP-TFI) nuclear receptor subfamily 2, group F, member 1 DNA-binding transcription factor with C4 type zinc finger domain (ligand-regulated) (36) PEG3 paternally expressed 3 DNA-binding transcription factor with -
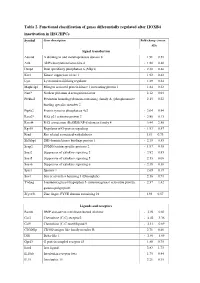
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Activated Peripheral-Blood-Derived Mononuclear Cells
Transcription factor expression in lipopolysaccharide- activated peripheral-blood-derived mononuclear cells Jared C. Roach*†, Kelly D. Smith*‡, Katie L. Strobe*, Stephanie M. Nissen*, Christian D. Haudenschild§, Daixing Zhou§, Thomas J. Vasicek¶, G. A. Heldʈ, Gustavo A. Stolovitzkyʈ, Leroy E. Hood*†, and Alan Aderem* *Institute for Systems Biology, 1441 North 34th Street, Seattle, WA 98103; ‡Department of Pathology, University of Washington, Seattle, WA 98195; §Illumina, 25861 Industrial Boulevard, Hayward, CA 94545; ¶Medtronic, 710 Medtronic Parkway, Minneapolis, MN 55432; and ʈIBM Computational Biology Center, P.O. Box 218, Yorktown Heights, NY 10598 Contributed by Leroy E. Hood, August 21, 2007 (sent for review January 7, 2007) Transcription factors play a key role in integrating and modulating system. In this model system, we activated peripheral-blood-derived biological information. In this study, we comprehensively measured mononuclear cells, which can be loosely termed ‘‘macrophages,’’ the changing abundances of mRNAs over a time course of activation with lipopolysaccharide (LPS). We focused on the precise mea- of human peripheral-blood-derived mononuclear cells (‘‘macro- surement of mRNA concentrations. There is currently no high- phages’’) with lipopolysaccharide. Global and dynamic analysis of throughput technology that can precisely and sensitively measure all transcription factors in response to a physiological stimulus has yet to mRNAs in a system, although such technologies are likely to be be achieved in a human system, and our efforts significantly available in the near future. To demonstrate the potential utility of advanced this goal. We used multiple global high-throughput tech- such technologies, and to motivate their development and encour- nologies for measuring mRNA levels, including massively parallel age their use, we produced data from a combination of two distinct signature sequencing and GeneChip microarrays. -

Expression of Oncogenes ELK1 and ELK3 in Cancer
Review Article Annals of Colorectal Cancer Research Published: 11 Nov, 2019 Expression of Oncogenes ELK1 and ELK3 in Cancer Akhlaq Ahmad and Asif Hayat* College of Chemistry, Fuzhou University, China Abstract Cancer is the uncontrolled growth of abnormal cells anywhere in a body, ELK1 and ELK3 is a member of the Ets-domain transcription factor family and the TCF (Ternary Complex Factor) subfamily. Proteins in this subfamily regulate transcription when recruited by SRF (Serum Response Factor) to bind to serum response elements. ELK1 and ELK3 transcription factors are known as oncogenes. Both transcription factors are proliferated in a different of type of cancer. Herein, we summarized the expression of transcription factor ELK1 and ELK3 in cancer cells. Keywords: ETS; ELK1; ELK3; Transcription factor; Cancer Introduction The ETS, a transcription factor of E twenty-six family based on a dominant ETS amino acids that integrated with a ~10-basepair element arrange in highly mid core sequence 5′-GGA(A/T)-3′ [1-2]. The secular family alter enormous 28/29 members which has been assigned in human and mouse and similarly the family description are further sub-divided into nine sub-families according to their homology and domain factor [3]. More importantly, one of the subfamily members such as ELK (ETS-like) adequate an N-terminal ETS DNA-binding domain along with a B-box domain that transmit the response of serum factor upon the formation of ternary complex and therefore manifested as ternary complex factors [4]. Further the ELK sub-divided into Elk1, Elk3 (Net, Erp or Sap2) and Elk4 (Sap1) proteins [3,4], which simulated varied proportional of potential protein- protein interactions [4,5]. -

TRANSCRIPTIONAL REGULATION of Hur in RENAL STRESS
TRANSCRIPTIONAL REGULATION OF HuR IN RENAL STRESS DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Sudha Suman Govindaraju Graduate Program in Biochemistry The Ohio State University 2014 Dissertation Committee: Dr. Beth S. Lee, Ph.D., Advisor Dr. Kathleen Boris-Lawrie, Ph.D. Dr. Sissy M. Jhiang, Ph.D. Dr. Arthur R. Strauch, Ph.D Abstract HuR is a ubiquitously expressed RNA-binding protein that affects the post- transcriptional life of thousands of cellular mRNAs by regulating transcript stability and translation. HuR can post-transcriptionally regulate gene expression and modulate cellular responses to stress, differentiation, proliferation, apoptosis, senescence, inflammation, and the immune response. It is an important mediator of survival during cellular stress, but when inappropriately expressed, can promote oncogenic transformation. Not surprisingly, the expression of HuR itself is tightly regulated at multiple transcriptional and post-transcriptional levels. Previous studies demonstrated the existence of two alternate HuR transcripts that differ in their 5’ untranslated regions and have markedly different translatabilities. These forms were also found to be reciprocally expressed following cellular stress in kidney proximal tubule cell lines, and the shorter, more readily translatable variant was shown to be regulated by Smad 1/5/8 pathway and bone morphogenetic protein-7 (BMP-7) signaling. In this study, the factors that promote transcription of the longer alternate form were identified. NF-κB was shown to be important for expression of the long HuR mRNA, as was a newly identified region with potential for binding the Sp/KLF families of transcription factors. -

View Is Portrayed Schematically in Figure 7B
BASIC RESEARCH www.jasn.org Recombination Signal Binding Protein for Ig-kJ Region Regulates Juxtaglomerular Cell Phenotype by Activating the Myo-Endocrine Program and Suppressing Ectopic Gene Expression † † ‡ Ruth M. Castellanos-Rivera,* Ellen S. Pentz,* Eugene Lin,* Kenneth W. Gross, † Silvia Medrano,* Jing Yu,§ Maria Luisa S. Sequeira-Lopez,* and R. Ariel Gomez* *Department of Pediatrics, School of Medicine, †Department of Biology, Graduate School of Arts and Sciences, and §Department of Cell Biology, University of Virginia, Charlottesville, Virginia; and ‡Department of Molecular and Cellular Biology, Roswell Park Cancer Institute, Buffalo, New York ABSTRACT Recombination signal binding protein for Ig-kJ region (RBP-J), the major downstream effector of Notch signaling, is necessary to maintain the number of renin-positive juxtaglomerular cells and the plasticity of arteriolar smooth muscle cells to re-express renin when homeostasis is threatened. We hypothesized that RBP-J controls a repertoire of genes that defines the phenotype of the renin cell. Mice bearing a bacterial artificial chromosome reporter with a mutated RBP-J binding site in the renin promoter had markedly reduced reporter expression at the basal state and in response to a homeostatic challenge. Mice with conditional deletion of RBP-J in renin cells had decreased expression of endocrine (renin and Akr1b7)and smooth muscle (Acta2, Myh11, Cnn1,andSmtn) genes and regulators of smooth muscle expression (miR- 145, SRF, Nfatc4, and Crip1). To determine whether RBP-J deletion decreased the endowment of renin cells, we traced the fate of these cells in RBP-J conditional deletion mice. Notably, the lineage staining patterns in mutant and control kidneys were identical, although mutant kidneys had fewer or no renin- expressing cells in the juxtaglomerular apparatus. -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

Olig2 and Ngn2 Function in Opposition to Modulate Gene Expression in Motor Neuron Progenitor Cells
Downloaded from genesdev.cshlp.org on September 29, 2021 - Published by Cold Spring Harbor Laboratory Press Olig2 and Ngn2 function in opposition to modulate gene expression in motor neuron progenitor cells Soo-Kyung Lee,1 Bora Lee,1 Esmeralda C. Ruiz, and Samuel L. Pfaff2 Gene Expression Laboratory, The Salk Institute for Biological Studies, La Jolla, California 92037, USA Spinal motor neurons and oligodendrocytes are generated sequentially from a common pool of progenitors termed pMN cells. Olig2 is a bHLH-class transcription factor in pMN cells, but it has remained unclear how its transcriptional activity is modulated to first produce motor neurons and then oligodendrocytes. Previous studies have shown that Olig2 primes pMN cells to become motor neurons by triggering the expression of Ngn2 and Lhx3. Here we show that Olig2 also antagonizes the premature expression of post-mitotic motor neuron genes in pMN cells. This blockade is counteracted by Ngn2, which accumulates heterogeneously in pMN cells, thereby releasing a subset of the progenitors to differentiate and activate expression of post-mitotic motor neuron genes. The antagonistic relationship between Ngn2 and Olig2 is mediated by protein interactions that squelch activity as well as competition for shared DNA-binding sites. Our data support a model in which the Olig2/Ngn2 ratio in progenitor cells serves as a gate for timing proper gene expression during the development of pMN cells: Olig2high maintains the pMN state, thereby holding cells in reserve for oligodendrocyte generation, whereas Ngn2high favors the conversion of pMN cells into post-mitotic motor neurons. [Keywords: Motor neuron; oligodendrocyte; development; basic helix–loop–helix (bHLH); neurogenin (Ngn); Olig] Supplemental material is available at http://www.genesdev.org. -

Human Transcription Factor Protein-Protein Interactions in Health and Disease
HELKA GÖÖS GÖÖS HELKA Recent Publications in this Series 45/2019 Mgbeahuruike Eunice Ego Evaluation of the Medicinal Uses and Antimicrobial Activity of Piper guineense (Schumach & Thonn) 46/2019 Suvi Koskinen AND DISEASE IN HEALTH INTERACTIONS PROTEIN-PROTEIN FACTOR HUMAN TRANSCRIPTION Near-Occlusive Atherosclerotic Carotid Artery Disease: Study with Computed Tomography Angiography 47/2019 Flavia Fontana DISSERTATIONES SCHOLAE DOCTORALIS AD SANITATEM INVESTIGANDAM Biohybrid Cloaked Nanovaccines for Cancer Immunotherapy UNIVERSITATIS HELSINKIENSIS 48/2019 Marie Mennesson Kainate Receptor Auxiliary Subunits Neto1 and Neto2 in Anxiety and Fear-Related Behaviors 49/2019 Zehua Liu Porous Silicon-Based On-Demand Nanohybrids for Biomedical Applications 50/2019 Veer Singh Marwah Strategies to Improve Standardization and Robustness of Toxicogenomics Data Analysis HELKA GÖÖS 51/2019 Iryna Hlushchenko Actin Regulation in Dendritic Spines: From Synaptic Plasticity to Animal Behavior and Human HUMAN TRANSCRIPTION FACTOR PROTEIN-PROTEIN Neurodevelopmental Disorders 52/2019 Heini Liimatta INTERACTIONS IN HEALTH AND DISEASE Efectiveness of Preventive Home Visits among Community-Dwelling Older People 53/2019 Helena Karppinen Older People´s Views Related to Their End of Life: Will-to-Live, Wellbeing and Functioning 54/2019 Jenni Laitila Elucidating Nebulin Expression and Function in Health and Disease 55/2019 Katarzyna Ciuba Regulation of Contractile Actin Structures in Non-Muscle Cells 56/2019 Sami Blom Spatial Characterisation of Prostate Cancer by Multiplex -

CD56+ T-Cells in Relation to Cytomegalovirus in Healthy Subjects and Kidney Transplant Patients
CD56+ T-cells in Relation to Cytomegalovirus in Healthy Subjects and Kidney Transplant Patients Institute of Infection and Global Health Department of Clinical Infection, Microbiology and Immunology Thesis submitted in accordance with the requirements of the University of Liverpool for the degree of Doctor in Philosophy by Mazen Mohammed Almehmadi December 2014 - 1 - Abstract Human T cells expressing CD56 are capable of tumour cell lysis following activation with interleukin-2 but their role in viral immunity has been less well studied. The work described in this thesis aimed to investigate CD56+ T-cells in relation to cytomegalovirus infection in healthy subjects and kidney transplant patients (KTPs). Proportions of CD56+ T cells were found to be highly significantly increased in healthy cytomegalovirus-seropositive (CMV+) compared to cytomegalovirus-seronegative (CMV-) subjects (8.38% ± 0.33 versus 3.29%± 0.33; P < 0.0001). In donor CMV-/recipient CMV- (D-/R-)- KTPs levels of CD56+ T cells were 1.9% ±0.35 versus 5.42% ±1.01 in D+/R- patients and 5.11% ±0.69 in R+ patients (P 0.0247 and < 0.0001 respectively). CD56+ T cells in both healthy CMV+ subjects and KTPs expressed markers of effector memory- RA T-cells (TEMRA) while in healthy CMV- subjects and D-/R- KTPs the phenotype was predominantly that of naïve T-cells. Other surface markers, CD8, CD4, CD58, CD57, CD94 and NKG2C were expressed by a significantly higher proportion of CD56+ T-cells in healthy CMV+ than CMV- subjects. Functional studies showed levels of pro-inflammatory cytokines IFN-γ and TNF-α, as well as granzyme B and CD107a were significantly higher in CD56+ T-cells from CMV+ than CMV- subjects following stimulation with CMV antigens. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Structural Basis of UGUA Recognition by the Nudix Protein CFI 25
Structural basis of UGUA recognition by the Nudix protein CFIm25 and implications for a regulatory role in mRNA 3′ processing Qin Yang, Gregory M. Gilmartin1, and Sylvie Doublié1 Department of Microbiology and Molecular Genetics, Stafford Hall, University of Vermont, Burlington, VT 05405 Edited by Joan A. Steitz, Howard Hughes Medical Institute, New Haven, CT, and approved April 20, 2010 (received for review January 21, 2010) — Human Cleavage Factor Im (CFIm) is an essential component of the C-terminal RS/RD alternating charge domain a structure simi- pre-mRNA 3′ processing complex that functions in the regulation of lar to that of the SR-protein family of splicing regulators. The 25 poly(A) site selection through the recognition of UGUA sequences small subunit (CFIm ) contains a Nudix domain, a protein do- upstream of the poly(A) site. Although the highly conserved 25 kDa main that most often participates in the hydrolysis of substrates subunit (CFIm25) of the CFIm complex possesses a characteristic containing a nucleotide diphosphate linked to a variable moiety α β α / / Nudix fold, CFIm25 has no detectable hydrolase activity. Here X (15). Found throughout all three kingdoms, Nudix proteins we report the crystal structures of the human CFIm25 homodimer in participate in a wide range of crucial housekeeping functions, in- complex with UGUAAA and UUGUAU RNA sequences. CFIm25 is the cluding the hydrolysis of mutagenic nucleotides, the modulation first Nudix protein to be reported to bind RNA in a sequence- of the levels of signaling molecules, and the monitoring of meta- 25 α β specific manner. The UGUA sequence contributes to binding speci- bolic intermediates (15).