Table S3. List of Predicted Genes Targeted by at Least 5 Mirnas Significantly up Regulated In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Comprehensive Molecular Characterization of Gastric Adenocarcinoma
Comprehensive molecular characterization of gastric adenocarcinoma The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Bass, A. J., V. Thorsson, I. Shmulevich, S. M. Reynolds, M. Miller, B. Bernard, T. Hinoue, et al. 2014. “Comprehensive molecular characterization of gastric adenocarcinoma.” Nature 513 (7517): 202-209. doi:10.1038/nature13480. http://dx.doi.org/10.1038/ nature13480. Published Version doi:10.1038/nature13480 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:12987344 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA NIH Public Access Author Manuscript Nature. Author manuscript; available in PMC 2014 September 22. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Nature. 2014 September 11; 513(7517): 202–209. doi:10.1038/nature13480. Comprehensive molecular characterization of gastric adenocarcinoma A full list of authors and affiliations appears at the end of the article. Abstract Gastric cancer is a leading cause of cancer deaths, but analysis of its molecular and clinical characteristics has been complicated by histological and aetiological heterogeneity. Here we describe a comprehensive molecular evaluation of 295 primary gastric adenocarcinomas as part of The Cancer -

Structure and Function of the Fgd Family of Divergent FYVE Domain Proteins
Biochemistry and Cell Biology Structure and Function of the Fgd Family of Divergent FYVE Domain Proteins Journal: Biochemistry and Cell Biology Manuscript ID bcb-2018-0185.R1 Manuscript Type: Mini Review Date Submitted by the 03-Aug-2018 Author: Complete List of Authors: Eitzen, Gary; University of Alberta Faculty of Medicine and Dentistry Smithers, Cameron C.; University of Alberta, Biochemistry Murray, Allan; University of Alberta Faculty of Medicine and Dentistry Overduin, Michael; University of Alberta Faculty of Medicine and Dentistry Draft Fgd, Pleckstrin Homology domain, FYVE domain, Dbl Homology Domain, Keyword: Rho GEF Is the invited manuscript for consideration in a Special CSMB Special Issue Issue? : https://mc06.manuscriptcentral.com/bcb-pubs Page 1 of 37 Biochemistry and Cell Biology Title: Structure and Function of the Fgd Family of Divergent FYVE Domain Proteins Authors: Gary Eitzen1, Cameron C. Smithers2, Allan G Murray3 and Michael Overduin2* Draft 1Department of Cell Biology, 2Department of Biochemistry, 3Department of Medicine, University of Alberta, Edmonton, Alberta, Canada *Corresponding author. Michael Overduin Telephone: +1 780 492 3518 Fax: +1 780 492-0886 E-mail: [email protected] https://mc06.manuscriptcentral.com/bcb-pubs Biochemistry and Cell Biology Page 2 of 37 Abstract FYVE domains are highly conserved protein modules that typically bind phosphatidylinositol 3-phosphate (PI3P) on the surface of early endosomes. Along with pleckstrin homology (PH) and phox homology (PX) domains, FYVE domains are the principal readers of the phosphoinositide (PI) code that mediate specific recognition of eukaryotic organelles. Of all the human FYVE domain-containing proteins, those within the Faciogenital dysplasia (Fgd) subfamily are particularly divergent, and couple with GTPases to exert unique cellular functions. -

Functional Annotation of Exon Skipping Event in Human Pora Kim1,*,†, Mengyuan Yang1,†,Keyiya2, Weiling Zhao1 and Xiaobo Zhou1,3,4,*
D896–D907 Nucleic Acids Research, 2020, Vol. 48, Database issue Published online 23 October 2019 doi: 10.1093/nar/gkz917 ExonSkipDB: functional annotation of exon skipping event in human Pora Kim1,*,†, Mengyuan Yang1,†,KeYiya2, Weiling Zhao1 and Xiaobo Zhou1,3,4,* 1School of Biomedical Informatics, The University of Texas Health Science Center at Houston, Houston, TX 77030, USA, 2College of Electronics and Information Engineering, Tongji University, Shanghai, China, 3McGovern Medical School, The University of Texas Health Science Center at Houston, Houston, TX 77030, USA and 4School of Dentistry, The University of Texas Health Science Center at Houston, Houston, TX 77030, USA Received August 13, 2019; Revised September 21, 2019; Editorial Decision October 03, 2019; Accepted October 03, 2019 ABSTRACT been used as therapeutic targets (3–8). For example, MET has lost the binding site of E3 ubiquitin ligase CBL through Exon skipping (ES) is reported to be the most com- exon 14 skipping event (9), resulting in an enhanced expres- mon alternative splicing event due to loss of func- sion level of MET. MET amplification drives the prolifera- tional domains/sites or shifting of the open read- tion of tumor cells. Multiple tyrosine kinase inhibitors, such ing frame (ORF), leading to a variety of human dis- as crizotinib, cabozantinib and capmatinib, have been used eases and considered therapeutic targets. To date, to treat patients with MET exon 14 skipping (10). Another systematic and intensive annotations of ES events example is the dystrophin gene (DMD) in Duchenne mus- based on the skipped exon units in cancer and cular dystrophy (DMD), a progressive neuromuscular dis- normal tissues are not available. -

Multifactorial Erβ and NOTCH1 Control of Squamous Differentiation and Cancer
Multifactorial ERβ and NOTCH1 control of squamous differentiation and cancer Yang Sui Brooks, … , Karine Lefort, G. Paolo Dotto J Clin Invest. 2014;124(5):2260-2276. https://doi.org/10.1172/JCI72718. Research Article Oncology Downmodulation or loss-of-function mutations of the gene encoding NOTCH1 are associated with dysfunctional squamous cell differentiation and development of squamous cell carcinoma (SCC) in skin and internal organs. While NOTCH1 receptor activation has been well characterized, little is known about how NOTCH1 gene transcription is regulated. Using bioinformatics and functional screening approaches, we identified several regulators of the NOTCH1 gene in keratinocytes, with the transcription factors DLX5 and EGR3 and estrogen receptor β (ERβ) directly controlling its expression in differentiation. DLX5 and ERG3 are required for RNA polymerase II (PolII) recruitment to the NOTCH1 locus, while ERβ controls NOTCH1 transcription through RNA PolII pause release. Expression of several identified NOTCH1 regulators, including ERβ, is frequently compromised in skin, head and neck, and lung SCCs and SCC-derived cell lines. Furthermore, a keratinocyte ERβ–dependent program of gene expression is subverted in SCCs from various body sites, and there are consistent differences in mutation and gene-expression signatures of head and neck and lung SCCs in female versus male patients. Experimentally increased ERβ expression or treatment with ERβ agonists inhibited proliferation of SCC cells and promoted NOTCH1 expression and squamous differentiation both in vitro and in mouse xenotransplants. Our data identify a link between transcriptional control of NOTCH1 expression and the estrogen response in keratinocytes, with implications for differentiation therapy of squamous cancer. Find the latest version: https://jci.me/72718/pdf Research article Multifactorial ERβ and NOTCH1 control of squamous differentiation and cancer Yang Sui Brooks,1,2 Paola Ostano,3 Seung-Hee Jo,1,2 Jun Dai,1,2 Spiro Getsios,4 Piotr Dziunycz,5 Günther F.L. -

Supplementary Materials
Supplementary Materials: Supplemental Table 1 Abbreviations FMDV Foot and Mouth Disease Virus FMD Foot and Mouth Disease NC Non-treated Control DEGs Differentially Expressed Genes RNA-seq High-throughput Sequencing of Mrna RT-qPCR Quantitative Real-time Reverse Transcriptase PCR TCID50 50% Tissue Culture Infective Doses CPE Cytopathic Effect MOI Multiplicity of Infection DMEM Dulbecco's Modified Eagle Medium FBS Fetal Bovine Serum PBS Phosphate Buffer Saline QC Quality Control FPKM Fragments per Kilo bases per Million fragments method GO Gene Ontology KEGG Kyoto Encyclopedia of Genes and Genomes R Pearson Correlation Coefficient NFKBIA NF-kappa-B Inhibitor alpha IL6 Interleukin 6 CCL4 C-C motif Chemokine 4 CXCL2 C-X-C motif Chemokine 2 TNF Tumor Necrosis Factor VEGFA Vascular Endothelial Growth Gactor A CCL20 C-C motif Chemokine 20 CSF2 Macrophage Colony-Stimulating Factor 2 GADD45B Growth Arrest and DNA Damage Inducible 45 beta MYC Myc proto-oncogene protein FOS Proto-oncogene c-Fos MCL1 Induced myeloid leukemia cell differentiation protein Mcl-1 MAP3K14 Mitogen-activated protein kinase kinase kinase 14 IRF1 Interferon regulatory factor 1 CCL5 C-C motif chemokine 5 ZBTB3 Zinc finger and BTB domain containing 3 OTX1 Orthodenticle homeobox 1 TXNIP Thioredoxin-interacting protein ZNF180 Znc Finger Protein 180 ZNF36 Znc Finger Protein 36 ZNF182 Zinc finger protein 182 GINS3 GINS complex subunit 3 KLF15 Kruppel-like factor 15 Supplemental Table 2 Primers for Verification of RNA-seq-detected DEGs with RT-qPCR TNF F: CGACTCAGTGCCGAGATCAA R: -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Primate Specific Retrotransposons, Svas, in the Evolution of Networks That Alter Brain Function
Title: Primate specific retrotransposons, SVAs, in the evolution of networks that alter brain function. Olga Vasieva1*, Sultan Cetiner1, Abigail Savage2, Gerald G. Schumann3, Vivien J Bubb2, John P Quinn2*, 1 Institute of Integrative Biology, University of Liverpool, Liverpool, L69 7ZB, U.K 2 Department of Molecular and Clinical Pharmacology, Institute of Translational Medicine, The University of Liverpool, Liverpool L69 3BX, UK 3 Division of Medical Biotechnology, Paul-Ehrlich-Institut, Langen, D-63225 Germany *. Corresponding author Olga Vasieva: Institute of Integrative Biology, Department of Comparative genomics, University of Liverpool, Liverpool, L69 7ZB, [email protected] ; Tel: (+44) 151 795 4456; FAX:(+44) 151 795 4406 John Quinn: Department of Molecular and Clinical Pharmacology, Institute of Translational Medicine, The University of Liverpool, Liverpool L69 3BX, UK, [email protected]; Tel: (+44) 151 794 5498. Key words: SVA, trans-mobilisation, behaviour, brain, evolution, psychiatric disorders 1 Abstract The hominid-specific non-LTR retrotransposon termed SINE–VNTR–Alu (SVA) is the youngest of the transposable elements in the human genome. The propagation of the most ancient SVA type A took place about 13.5 Myrs ago, and the youngest SVA types appeared in the human genome after the chimpanzee divergence. Functional enrichment analysis of genes associated with SVA insertions demonstrated their strong link to multiple ontological categories attributed to brain function and the disorders. SVA types that expanded their presence in the human genome at different stages of hominoid life history were also associated with progressively evolving behavioural features that indicated a potential impact of SVA propagation on a cognitive ability of a modern human. -

Supplemental Materials ZNF281 Enhances Cardiac Reprogramming
Supplemental Materials ZNF281 enhances cardiac reprogramming by modulating cardiac and inflammatory gene expression Huanyu Zhou, Maria Gabriela Morales, Hisayuki Hashimoto, Matthew E. Dickson, Kunhua Song, Wenduo Ye, Min S. Kim, Hanspeter Niederstrasser, Zhaoning Wang, Beibei Chen, Bruce A. Posner, Rhonda Bassel-Duby and Eric N. Olson Supplemental Table 1; related to Figure 1. Supplemental Table 2; related to Figure 1. Supplemental Table 3; related to the “quantitative mRNA measurement” in Materials and Methods section. Supplemental Table 4; related to the “ChIP-seq, gene ontology and pathway analysis” and “RNA-seq” and gene ontology analysis” in Materials and Methods section. Supplemental Figure S1; related to Figure 1. Supplemental Figure S2; related to Figure 2. Supplemental Figure S3; related to Figure 3. Supplemental Figure S4; related to Figure 4. Supplemental Figure S5; related to Figure 6. Supplemental Table S1. Genes included in human retroviral ORF cDNA library. Gene Gene Gene Gene Gene Gene Gene Gene Symbol Symbol Symbol Symbol Symbol Symbol Symbol Symbol AATF BMP8A CEBPE CTNNB1 ESR2 GDF3 HOXA5 IL17D ADIPOQ BRPF1 CEBPG CUX1 ESRRA GDF6 HOXA6 IL17F ADNP BRPF3 CERS1 CX3CL1 ETS1 GIN1 HOXA7 IL18 AEBP1 BUD31 CERS2 CXCL10 ETS2 GLIS3 HOXB1 IL19 AFF4 C17ORF77 CERS4 CXCL11 ETV3 GMEB1 HOXB13 IL1A AHR C1QTNF4 CFL2 CXCL12 ETV7 GPBP1 HOXB5 IL1B AIMP1 C21ORF66 CHIA CXCL13 FAM3B GPER HOXB6 IL1F3 ALS2CR8 CBFA2T2 CIR1 CXCL14 FAM3D GPI HOXB7 IL1F5 ALX1 CBFA2T3 CITED1 CXCL16 FASLG GREM1 HOXB9 IL1F6 ARGFX CBFB CITED2 CXCL3 FBLN1 GREM2 HOXC4 IL1F7 -

Towards a Molecular Understanding of Microrna-Mediated Gene Silencing
REVIEWS NON-CODING RNA Towards a molecular understanding of microRNA-mediated gene silencing Stefanie Jonas and Elisa Izaurralde Abstract | MicroRNAs (miRNAs) are a conserved class of small non-coding RNAs that assemble with Argonaute proteins into miRNA-induced silencing complexes (miRISCs) to direct post-transcriptional silencing of complementary mRNA targets. Silencing is accomplished through a combination of translational repression and mRNA destabilization, with the latter contributing to most of the steady-state repression in animal cell cultures. Degradation of the mRNA target is initiated by deadenylation, which is followed by decapping and 5ʹ‑to‑3ʹ exonucleolytic decay. Recent work has enhanced our understanding of the mechanisms of silencing, making it possible to describe in molecular terms a continuum of direct interactions from miRNA target recognition to mRNA deadenylation, decapping and 5ʹ‑to‑3ʹ degradation. Furthermore, an intricate interplay between translational repression and mRNA degradation is emerging. Deadenylation MicroRNAs (miRNAs) are conserved post-transcriptional recruit additional protein partners to mediate silenc- 5,6 Shortening of mRNA poly(A) regulators of gene expression that are integral to ing . Silencing occurs through a combination of tails. In eukaryotes, this almost all known biological processes, including translational repression, deadenylation, decapping and process is catalysed by the cell growth, proliferation and differentiation, as well 5ʹ‑to‑3ʹ mRNA degradation5,6 (FIG. 1). The GW182 pro- consecutive but partially as organismal metabolism and development1. The teins play a central part in this process and are among redundant action of two 5,6 cytoplasmic deadenylase number of miRNAs encoded within the genomes of the most extensively studied AGO partners . -

Profiling Data
Compound Name DiscoveRx Gene Symbol Entrez Gene Percent Compound Symbol Control Concentration (nM) JNK-IN-8 AAK1 AAK1 69 1000 JNK-IN-8 ABL1(E255K)-phosphorylated ABL1 100 1000 JNK-IN-8 ABL1(F317I)-nonphosphorylated ABL1 87 1000 JNK-IN-8 ABL1(F317I)-phosphorylated ABL1 100 1000 JNK-IN-8 ABL1(F317L)-nonphosphorylated ABL1 65 1000 JNK-IN-8 ABL1(F317L)-phosphorylated ABL1 61 1000 JNK-IN-8 ABL1(H396P)-nonphosphorylated ABL1 42 1000 JNK-IN-8 ABL1(H396P)-phosphorylated ABL1 60 1000 JNK-IN-8 ABL1(M351T)-phosphorylated ABL1 81 1000 JNK-IN-8 ABL1(Q252H)-nonphosphorylated ABL1 100 1000 JNK-IN-8 ABL1(Q252H)-phosphorylated ABL1 56 1000 JNK-IN-8 ABL1(T315I)-nonphosphorylated ABL1 100 1000 JNK-IN-8 ABL1(T315I)-phosphorylated ABL1 92 1000 JNK-IN-8 ABL1(Y253F)-phosphorylated ABL1 71 1000 JNK-IN-8 ABL1-nonphosphorylated ABL1 97 1000 JNK-IN-8 ABL1-phosphorylated ABL1 100 1000 JNK-IN-8 ABL2 ABL2 97 1000 JNK-IN-8 ACVR1 ACVR1 100 1000 JNK-IN-8 ACVR1B ACVR1B 88 1000 JNK-IN-8 ACVR2A ACVR2A 100 1000 JNK-IN-8 ACVR2B ACVR2B 100 1000 JNK-IN-8 ACVRL1 ACVRL1 96 1000 JNK-IN-8 ADCK3 CABC1 100 1000 JNK-IN-8 ADCK4 ADCK4 93 1000 JNK-IN-8 AKT1 AKT1 100 1000 JNK-IN-8 AKT2 AKT2 100 1000 JNK-IN-8 AKT3 AKT3 100 1000 JNK-IN-8 ALK ALK 85 1000 JNK-IN-8 AMPK-alpha1 PRKAA1 100 1000 JNK-IN-8 AMPK-alpha2 PRKAA2 84 1000 JNK-IN-8 ANKK1 ANKK1 75 1000 JNK-IN-8 ARK5 NUAK1 100 1000 JNK-IN-8 ASK1 MAP3K5 100 1000 JNK-IN-8 ASK2 MAP3K6 93 1000 JNK-IN-8 AURKA AURKA 100 1000 JNK-IN-8 AURKA AURKA 84 1000 JNK-IN-8 AURKB AURKB 83 1000 JNK-IN-8 AURKB AURKB 96 1000 JNK-IN-8 AURKC AURKC 95 1000 JNK-IN-8 -

Redefining the Specificity of Phosphoinositide-Binding by Human
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.20.163253; this version posted June 21, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Redefining the specificity of phosphoinositide-binding by human PH domain-containing proteins Nilmani Singh1†, Adriana Reyes-Ordoñez1†, Michael A. Compagnone1, Jesus F. Moreno Castillo1, Benjamin J. Leslie2, Taekjip Ha2,3,4,5, Jie Chen1* 1Department of Cell & Developmental Biology, University of Illinois at Urbana-Champaign, Urbana, IL 61801; 2Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD 21205; 3Department of Biophysics, Johns Hopkins University, Baltimore, MD 21218; 4Department of Biomedical Engineering, Johns Hopkins University, Baltimore, MD 21205; 5Howard Hughes Medical Institute, Baltimore, MD 21205, USA †These authors contributed equally to this work. *Correspondence: [email protected]. bioRxiv preprint doi: https://doi.org/10.1101/2020.06.20.163253; this version posted June 21, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. ABSTRACT Pleckstrin homology (PH) domains are presumed to bind phosphoinositides (PIPs), but specific interaction with and regulation by PIPs for most PH domain-containing proteins are unclear. Here we employed a single-molecule pulldown assay to study interactions of lipid vesicles with full-length proteins in mammalian whole cell lysates. -
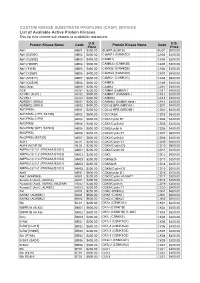
CUSTOM KINASE SUBSTRATE PROFILING (CKSP) SERVICE List of Available Active Protein Kinases This List May Change with Respect to Availability and Pricing
CUSTOM KINASE SUBSTRATE PROFILING (CKSP) SERVICE List of Available Active Protein Kinases This list may change with respect to availability and pricing. Protein Kinase Name Code U.S. Protein Kinase Name Code U.S. Price Price Abl1 AB01 $200.00 BUBR1(BUB1B) BU01 $600.00 Abl1 [E255K] AB02 $400.00 CaMK1δ (CAMK1D) CA03 $200.00 Abl1 [G250E] AB03 $400.00 CAMK1γ CA04 $200.00 Abl1 [H369P] AB04 $600.00 CAMK2α (CAMK2B) CA05 $200.00 Abl1 [T315I] AB05 $400.00 CaMK2β (CAMK2B) CA06 $200.00 Abl1 [Y253F] AB06 $400.00 CaMK2δ (CAMK2D) CA07 $400.00 Abl1 [M351T] AB07 $600.00 CaMK2γ (CAMK2G) CA08 $600.00 Abl1 [Q252H] AB08 $600.00 CAMK3γ CA09 $200.00 Abl2 (Arg) AB09 $200.00 CAMK4 CA10 $200.00 ACK AC01 $200.00 CAMK4 (CaMKIV ) CA11 $400.00 ACVR1 (ALK2 ) AC02 $400.00 CAMKK1 (CAMKKA ) CA12 $200.00 ACVRL1 AC03 $400.00 CAMKK2 CA13 $200.00 ADRBK1 (GRK2) AD01 $200.00 CAMKK2 (CaMKK beta ) CA14 $400.00 ADRBK2 (GRK3) AD02 $400.00 CDC42 BPA (MRCKA ) CD01 $400.00 Akt1/PKBα AK01 $200.00 CDC42 BPB (MRCKB ) CD02 $400.00 Akt1/PKBα [δPH, S473D] AK02 $600.00 CDC7/ASK CD03 $600.00 Akt1/PKBα [δPH] AK03 $600.00 CDK1/cyclin B1 CD04 $400.00 Akt2/PKBβ AK04 $200.00 CDK1/CyclinA2 CD05 $200.00 Akt2/PKBβ [δPH, S474D] AK05 $600.00 CDK2/cyclin A CD06 $400.00 Akt3/PKBγ AK06 $200.00 CDK2/Cyclin E1 CD07 $600.00 Akt3/PKBγ [S472D] AK07 $600.00 CDK2/CyclinA2 CD08 $200.00 ALK1 AL01 $200.00 CDK3/Cyclin E1 CD09 $600.00 ALK4 (ACVR1B) AL02 $200.00 CDK4//Cyclin D3 CD10 $600.00 AMPKα1/β1/γ1 (PRKAA1/B1/G1) AM01 $200.00 CDK4/Cyclin D1 CD11 $200.00 AMPKα1/β1/γ2 (PRKAA1/B1/G2) AM02 $200.00 CDK5 CD12 $600.00