Population Genetics of Endangered Catostomid Fishes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
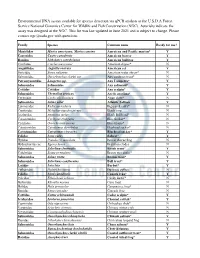
Edna Assay Development
Environmental DNA assays available for species detection via qPCR analysis at the U.S.D.A Forest Service National Genomics Center for Wildlife and Fish Conservation (NGC). Asterisks indicate the assay was designed at the NGC. This list was last updated in June 2021 and is subject to change. Please contact [email protected] with questions. Family Species Common name Ready for use? Mustelidae Martes americana, Martes caurina American and Pacific marten* Y Castoridae Castor canadensis American beaver Y Ranidae Lithobates catesbeianus American bullfrog Y Cinclidae Cinclus mexicanus American dipper* N Anguillidae Anguilla rostrata American eel Y Soricidae Sorex palustris American water shrew* N Salmonidae Oncorhynchus clarkii ssp Any cutthroat trout* N Petromyzontidae Lampetra spp. Any Lampetra* Y Salmonidae Salmonidae Any salmonid* Y Cottidae Cottidae Any sculpin* Y Salmonidae Thymallus arcticus Arctic grayling* Y Cyrenidae Corbicula fluminea Asian clam* N Salmonidae Salmo salar Atlantic Salmon Y Lymnaeidae Radix auricularia Big-eared radix* N Cyprinidae Mylopharyngodon piceus Black carp N Ictaluridae Ameiurus melas Black Bullhead* N Catostomidae Cycleptus elongatus Blue Sucker* N Cichlidae Oreochromis aureus Blue tilapia* N Catostomidae Catostomus discobolus Bluehead sucker* N Catostomidae Catostomus virescens Bluehead sucker* Y Felidae Lynx rufus Bobcat* Y Hylidae Pseudocris maculata Boreal chorus frog N Hydrocharitaceae Egeria densa Brazilian elodea N Salmonidae Salvelinus fontinalis Brook trout* Y Colubridae Boiga irregularis Brown tree snake* -

Endangered Species
FEATURE: ENDANGERED SPECIES Conservation Status of Imperiled North American Freshwater and Diadromous Fishes ABSTRACT: This is the third compilation of imperiled (i.e., endangered, threatened, vulnerable) plus extinct freshwater and diadromous fishes of North America prepared by the American Fisheries Society’s Endangered Species Committee. Since the last revision in 1989, imperilment of inland fishes has increased substantially. This list includes 700 extant taxa representing 133 genera and 36 families, a 92% increase over the 364 listed in 1989. The increase reflects the addition of distinct populations, previously non-imperiled fishes, and recently described or discovered taxa. Approximately 39% of described fish species of the continent are imperiled. There are 230 vulnerable, 190 threatened, and 280 endangered extant taxa, and 61 taxa presumed extinct or extirpated from nature. Of those that were imperiled in 1989, most (89%) are the same or worse in conservation status; only 6% have improved in status, and 5% were delisted for various reasons. Habitat degradation and nonindigenous species are the main threats to at-risk fishes, many of which are restricted to small ranges. Documenting the diversity and status of rare fishes is a critical step in identifying and implementing appropriate actions necessary for their protection and management. Howard L. Jelks, Frank McCormick, Stephen J. Walsh, Joseph S. Nelson, Noel M. Burkhead, Steven P. Platania, Salvador Contreras-Balderas, Brady A. Porter, Edmundo Díaz-Pardo, Claude B. Renaud, Dean A. Hendrickson, Juan Jacobo Schmitter-Soto, John Lyons, Eric B. Taylor, and Nicholas E. Mandrak, Melvin L. Warren, Jr. Jelks, Walsh, and Burkhead are research McCormick is a biologist with the biologists with the U.S. -

Conservation Status of Threatened Fishes in Warner Basin, Oregon
Great Basin Naturalist Volume 50 Number 3 Article 5 10-31-1990 Conservation status of threatened fishes in arnerW Basin, Oregon Jack E. Williams Division of Wildlife and Fisheries, Bureau of Land Management, Washington, D.C. Mark A. Stern The Nature Conservancy, Portland, Oregon Alan V. Munhall Bureau of Land Management, Lakeview, Oregon Gary A. Anderson Oregon Department of Fish and Wildlife, Lakeview, Oregon Follow this and additional works at: https://scholarsarchive.byu.edu/gbn Recommended Citation Williams, Jack E.; Stern, Mark A.; Munhall, Alan V.; and Anderson, Gary A. (1990) "Conservation status of threatened fishes in arnerW Basin, Oregon," Great Basin Naturalist: Vol. 50 : No. 3 , Article 5. Available at: https://scholarsarchive.byu.edu/gbn/vol50/iss3/5 This Article is brought to you for free and open access by the Western North American Naturalist Publications at BYU ScholarsArchive. It has been accepted for inclusion in Great Basin Naturalist by an authorized editor of BYU ScholarsArchive. For more information, please contact [email protected], [email protected]. Creat &Isio N:l.luraUst 50(3), 1900, pp. 243-248 CONSERVATION STATUS OF THREATENED FISHES IN WARNER BASIN, OREGON 1 l 3 Jack E. Williams , MarkA. Stern \ Alan V. Munhall , and Cary A. Anderson"" A8S'TRACT.-Two fedemlJy listed fisbes, the Foskett speckled daceand Warnersucker, are endemic to Warner Basin in south central Oregon. The Foskett speckled dace is native only to a single spring in Coleman Valley. Anearby'spring was stocked with dace in 1979 and 1980, and now provides a second population. The present numbers ofdace probably are at their Wgbest levels since settlement ofthe region. -

Threats, Conservation Strategies, and Prognosis for Suckers (Catostomidae) in North America: Insights from Regional Case Studies of a Diverse Family of Non-Game fishes
BIOLOGICAL CONSERVATION Biological Conservation 121 (2005) 317–331 www.elsevier.com/locate/biocon Review Threats, conservation strategies, and prognosis for suckers (Catostomidae) in North America: insights from regional case studies of a diverse family of non-game fishes Steven J. Cooke a,b,*,1, Christopher M. Bunt c, Steven J. Hamilton d, Cecil A. Jennings e, Michael P. Pearson f, Michael S. Cooperman g, Douglas F. Markle g a Department of Forest Sciences, Centre for Applied Conservation Research, University of British Columbia, 2424 Main Mall, Vancouver, BC, Canada V6T 1Z4 b Centre for Aquatic Ecology, Illinois Natural History Survey, 607 E. Peabody Dr., Champaign, IL 61820, USA c Biotactic Inc., 691 Hidden Valley Rd., Kitchener, Ont., Canada N2C 2S4 d Yankton Field Research Station, Columbia Environmental Research Center, United States Geological Survey, Yankton, SD 57078, USA e United States Geological Survey, Georgia Cooperative Fish and Wildlife Research Unit, School of Forest Resources, University of Georgia, Athens, GA 30602, USA f Fisheries Centre and Institute for Resources, Environment, and Sustainability, University of British Columbia, Vancouver, BC, Canada V6T 1Z4 g Department of Fisheries and Wildlife, Oregon State University, Corvallis, OR 97331, USA Received 10 December 2003; received in revised form 6 May 2004; accepted 18 May 2004 Abstract Catostomid fishes are a diverse family of 76+ freshwater species that are distributed across North America in many different habitats. This group of fish is facing a variety of impacts and conservation issues that are somewhat unique relative to more economically valuable and heavily managed fish species. Here, we present a brief series of case studies to highlight the threats such as migration barriers, flow regulation, environmental contamination, habitat degradation, exploitation and impacts from introduced (non-native) species that are facing catostomids in different regions. -

Fishes As a Template for Reticulate Evolution
University of Arkansas, Fayetteville ScholarWorks@UARK Theses and Dissertations 12-2016 Fishes as a Template for Reticulate Evolution: A Case Study Involving Catostomus in the Colorado River Basin of Western North America Max Russell Bangs University of Arkansas, Fayetteville Follow this and additional works at: http://scholarworks.uark.edu/etd Part of the Evolution Commons, Molecular Biology Commons, and the Terrestrial and Aquatic Ecology Commons Recommended Citation Bangs, Max Russell, "Fishes as a Template for Reticulate Evolution: A Case Study Involving Catostomus in the Colorado River Basin of Western North America" (2016). Theses and Dissertations. 1847. http://scholarworks.uark.edu/etd/1847 This Dissertation is brought to you for free and open access by ScholarWorks@UARK. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of ScholarWorks@UARK. For more information, please contact [email protected], [email protected]. Fishes as a Template for Reticulate Evolution: A Case Study Involving Catostomus in the Colorado River Basin of Western North America A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biology by Max Russell Bangs University of South Carolina Bachelor of Science in Biological Sciences, 2009 University of South Carolina Master of Science in Integrative Biology, 2011 December 2016 University of Arkansas This dissertation is approved for recommendation to the Graduate Council. _____________________________________ Dr. Michael E. Douglas Dissertation Director _____________________________________ ____________________________________ Dr. Marlis R. Douglas Dr. Andrew J. Alverson Dissertation Co-Director Committee Member _____________________________________ Dr. Thomas F. Turner Ex-Officio Member Abstract Hybridization is neither simplistic nor phylogenetically constrained, and post hoc introgression can have profound evolutionary effects. -

Distribution and Biology of Suckers in Lower Klamath Reservoirs
DISTRIBUTION AND BIOLOGY OF SUCKERS IN LOWER KLAMATH RESERVOIRS 1999 FINAL REPORT SUBMITTED TO PacifiCorp 825 NE Multnomah Blvd., Ste. 1500 Portland, OR 97232 By Marc Desjardins & Douglas F. Markle 104 Nash Hall Department of Fisheries and Wildlife Oregon State University Corvallis, Oregon 97331-3803 March 28, 2000 Abstract The objectives of this two-year study (1998-1999) were to document distribution, abundance, age class structure, recruitment success, and habitat use by all life history stages of shortnose and Lost River suckers in three lower Klamath River hydroelectric reservoirs (J. C. Boyle, Copco, and Iron Gate). Lost River sucker catches were sporadic (only 3 adult individuals total) and the focus of our analyses, therefore, shifted to shortnose suckers. Adult and larval suckers were found in all reservoirs both years. All life history stages (larvae, juveniles and adults) were found in J. C. Boyle during both years and in Copco in 1999. Juvenile suckers were not found in Copco in 1998. The number of adult shortnose suckers was highest in Copco reservoir (n=165), followed by J.C. Boyle (n=50) and Iron Gate (n=22). Larger and older individuals dominated Copco and Iron Gate reservoirs and little size structure was detected. J. C. Boyle tended to have smaller adult shortnose suckers and many size classes were present. Unidentifiable larval suckers were most abundant in Copco reservoir where historic spawning of shortnose suckers has been documented. Larval suckers in Copco and Iron Gate reservoirs were most abundant in mid to late June before quickly disappearing from catches. J. C. Boyle larval suckers peaked in mid July, attained larger sizes, and were caught later in the season. -

Amphibian Cascades Frog SOC X X Mountain Meadows, Bogs, Ponds Or
Taxa Species Species SMU/ESU/D Federal State BM CP CR EC KM NR WC WV NS Special needs Limiting factors Data gaps Conservation actions Key reference or plan, if available Common Scientific PS/Group Listing listing Name Name Status status Amphibian Cascades SOC X X Mountain meadows, bogs, ponds or potholes Montane species vulnerable to genetic Habitat requirements and how they may vary by Maintain connectivity of habitat. Monitor effects of fish http://www.fs.fed.us/psw/publications/documents/ frog above 2,400 feet elevation. Requires access isolation. Experiencing substantial elevation within the species' range. Habitat stocking and water quality on populations. Carefully manage psw_gtr244/psw_gtr244.pdf to permanent water. Lays eggs in shallow reductions in southern parts of range characteristics that could enhance migration livestock grazing in occupied wet meadows. Use prescribed sunny edges of ponds, or on low vegetation (e.g., CA). Potentially sensitive to and gene flow. Feeding habits. Effects of burning or hand-felling of trees periodically to set plant near ponds where warm sunlight speeds egg waterborne pathogens. pathogens airborne environmental pollution. succession. If reintroductions are warranted, use individuals development. Larvae may “school” in large Feasibility studies on reintroduction at historic from nearby populations and consult results of feasibility masses. sites. studies. Conservation actions in Oregon are particularly valuable given reductions in other parts of range. Amphibian Cascade X X Cold, fast-flowing, clear, permanent headwater Larvae take several years to reach sexual Basic inventory, abundance and population Maintain stream buffers to maintain cool water Howell, B.L. and N. M. Maggiulli. 2011. -

Lost River Sucker 5-Year Status Review
Lost River Sucker (Deltistes luxatus) 5-Year Review Summary and Evaluation U.S. Fish and Wildlife Service Klamath Falls Fish and Wildlife Office Klamath Falls, Oregon July 2007 5-YEAR REVIEW Lost River Sucker (Deltistes luxatus) TABLE OF CONTENTS 1.0 GENERAL INFORMATION.......................................................................................... 1 1.1. Reviewers............................................................................................................................ 1 1.2. Methodology used to complete the review....................................................................... 1 1.3. Background ........................................................................................................................ 1 2.0 REVIEW ANALYSIS....................................................................................................... 2 2.1. Application of the 1996 Distinct Populations Segment (DPS) policy ............................ 2 2.2. Biology and Habitat ........................................................................................................... 3 2.3. Recovery Criteria............................................................................................................. 12 2.4. Five-Factor Analysis ........................................................................................................ 15 2.5. Synthesis............................................................................................................................ 29 3.0 RESULTS ....................................................................................................................... -

Juvenile Lost River and Shortnose Sucker Year Class Strength
Juvenile Lost River and Shortnose Sucker Year Class Strength, Survival, and Growth in Upper Klamath Lake, Oregon, and Clear Lake Reservoir, California: 2016 Monitoring Report Open-File Report 2018–1066 U.S. Department of the Interior U.S. Geological Survey Cover: Photograph showing Upper Klamath Lake, Oregon in July with a cyanobacterial bloom. Photograph taken from Modoc Rim on the eastern shore looking to the northwest. Photograph from U.S. Geological Survey. Juvenile Lost River and Shortnose Sucker Year Class Strength, Survival, and Growth in Upper Klamath Lake, Oregon, and Clear Lake Reservoir, California—2016 Monitoring Report By Summer M. Burdick, Carl O. Ostberg, and Marshal S. Hoy Open-File Report 2018–1066 U.S. Department of the Interior U.S. Geological Survey U.S. Department of the Interior RYAN K. ZINKE, Secretary U.S. Geological Survey William H. Werkheiser, Deputy Director exercising the authority of the Director U.S. Geological Survey, Reston, Virginia: 2018 For more information on the USGS—the Federal source for science about the Earth, its natural and living resources, natural hazards, and the environment—visit https://www.usgs.gov/ or call 1–888–ASK–USGS (1–888–275–8747). For an overview of USGS information products, including maps, imagery, and publications, visit https://store.usgs.gov. Any use of trade, firm, or product names is for descriptive purposes only and does not imply endorsement by the U.S. Government. Although this information product, for the most part, is in the public domain, it also may contain copyrighted materials as noted in the text. Permission to reproduce copyrighted items must be secured from the copyright owner. -

Microsoft Outlook
Joey Steil From: Leslie Jordan <[email protected]> Sent: Tuesday, September 25, 2018 1:13 PM To: Angela Ruberto Subject: Potential Environmental Beneficial Users of Surface Water in Your GSA Attachments: Paso Basin - County of San Luis Obispo Groundwater Sustainabilit_detail.xls; Field_Descriptions.xlsx; Freshwater_Species_Data_Sources.xls; FW_Paper_PLOSONE.pdf; FW_Paper_PLOSONE_S1.pdf; FW_Paper_PLOSONE_S2.pdf; FW_Paper_PLOSONE_S3.pdf; FW_Paper_PLOSONE_S4.pdf CALIFORNIA WATER | GROUNDWATER To: GSAs We write to provide a starting point for addressing environmental beneficial users of surface water, as required under the Sustainable Groundwater Management Act (SGMA). SGMA seeks to achieve sustainability, which is defined as the absence of several undesirable results, including “depletions of interconnected surface water that have significant and unreasonable adverse impacts on beneficial users of surface water” (Water Code §10721). The Nature Conservancy (TNC) is a science-based, nonprofit organization with a mission to conserve the lands and waters on which all life depends. Like humans, plants and animals often rely on groundwater for survival, which is why TNC helped develop, and is now helping to implement, SGMA. Earlier this year, we launched the Groundwater Resource Hub, which is an online resource intended to help make it easier and cheaper to address environmental requirements under SGMA. As a first step in addressing when depletions might have an adverse impact, The Nature Conservancy recommends identifying the beneficial users of surface water, which include environmental users. This is a critical step, as it is impossible to define “significant and unreasonable adverse impacts” without knowing what is being impacted. To make this easy, we are providing this letter and the accompanying documents as the best available science on the freshwater species within the boundary of your groundwater sustainability agency (GSA). -

United States Department of the Interior
United States Department of the Interior FISH AND WILDLIFE SERVICE Oregon Fish and Wildlife Office 2600 SE 98th Avenue, Suite 100 Portland, Oregon 97266 Phone: (503) 231-6179 FAX: (503) 231-6195 Reply To: 8330.F0047(09) File Name: CREP BO 2009_final.doc TS Number: 09-314 TAILS: 13420-2009-F-0047 Doc Type: Final Don Howard, Acting State Executive Director U.S. Department of Agriculture Farm Service Agency, Oregon State Office 7620 SW Mohawk St. Tualatin, OR 97062-8121 Dear Mr. Howard, This letter transmits the U.S. Fish and Wildlife Service’s (Service) Biological and Conference Opinion (BO) and includes our written concurrence based on our review of the proposed Oregon Conservation Reserve Enhancement Program (CREP) to be administered by the Farm Service Agency (FSA) throughout the State of Oregon, and its effects on Federally-listed species in accordance with section 7 of the Endangered Species Act (Act) of 1973, as amended (16 U.S.C. 1531 et seq.). Your November 24, 2008 request for informal and formal consultation with the Service, and associated Program Biological Assessment for the Oregon Conservation Reserve Enhancement Program (BA), were received on November 24, 2008. We received your letter providing a 90-day extension on March 26, 2009 based on the scope and complexity of the program and the related species that are covered, which we appreciated. This Concurrence and BO covers a period of approximately 10 years, from the date of issuance through December 31, 2019. The BA also includes species that fall within the jurisdiction of the National Oceanic and Atmospheric Administration’s Fisheries Service (NOAA Fisheries Service). -

1 CWU Comparative Osteology Collection, List of Specimens
CWU Comparative Osteology Collection, List of Specimens List updated November 2019 0-CWU-Collection-List.docx Specimens collected primarily from North American mid-continent and coastal Alaska for zooarchaeological research and teaching purposes. Curated at the Zooarchaeology Laboratory, Department of Anthropology, Central Washington University, under the direction of Dr. Pat Lubinski, [email protected]. Facility is located in Dean Hall Room 222 at CWU’s campus in Ellensburg, Washington. Numbers on right margin provide a count of complete or near-complete specimens in the collection. Specimens on loan from other institutions are not listed. There may also be a listing of mount (commercially mounted articulated skeletons), part (partial skeletons), skull (skulls), or * (in freezer but not yet processed). Vertebrate specimens in taxonomic order, then invertebrates. Taxonomy follows the Integrated Taxonomic Information System online (www.itis.gov) as of June 2016 unless otherwise noted. VERTEBRATES: Phylum Chordata, Class Petromyzontida (lampreys) Order Petromyzontiformes Family Petromyzontidae: Pacific lamprey ............................................................. Entosphenus tridentatus.................................... 1 Phylum Chordata, Class Chondrichthyes (cartilaginous fishes) unidentified shark teeth ........................................................ ........................................................................... 3 Order Squaliformes Family Squalidae Spiny dogfish ........................................................