Bacteriophage T4 Infection of Stationary Phase E. Coli: Life After Log from a Phage Perspective
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Concatemerization of Synthetic Oligonucleotides Assembly and Ligand Interactions
THESIS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY CONCATEMERIZATION OF SYNTHETIC OLIGONUCLEOTIDES ASSEMBLY AND LIGAND INTERACTIONS LU SUN Department of Chemical and Biological Engineering CHALMERS UNIVERSITY OF TECHNOLOGY GÖTEBORG, SWEDEN 2014 Concatemerization of Synthetic Oligonucleotides Assembly and Ligand Interactions LU SUN ISBN 978-91-7597-092-9 © LU SUN, 2014 Doktorsavhandlingar vid Chalmers tekniska högskola Ny serie nr 3773 ISSN 0346-718X Department of Chemical and Biological Engineering Chalmers University of Technology SE-412 96 Göteborg Sweden Telephone + 46 (0)31-772 1000 Cover image: Two ways of forming DNA concatemers. (Top right) Self-assembly of oligonucleotides in free solution. (Down) A three-step sequential assembly of DNA concatemer layer on a planar surface and RAD51 induced DNA layer extension. Back cover photo: © Keling Bi Printed by Chalmers Reproservice Göteborg, Sweden 2014 ii Concatemerization of Synthetic Oligonucleotides Assembly and Ligand Interactions LU SUN Department of Chemical and Biological Engineering Chalmers University of Technology ABSTRACT DNA nanotechnology has become an important research field because of its advantages in high predictability and accuracy of base paring recognition. For example, the DNA concatemer, one of the simplest DNA constructs in shape, has been used to enhance signals in biosensing. In this thesis, concatemers were designed and characterized towards sequence-specific target amplifiers for single-molecule mechanical studies. The project firstly focuses on exploring how to obtain concatemers of satisfactory length by self- assembly in bulk. Concatemers formed in solution by mixing of the different components are characterized using gel electrophoresis and AFM. Experimental results demonstrate that the concatemerization yield could be increased primarily by increasing the ionic strength, and linear concatemers of expected length could be separated from mixed sizes and shapes. -

P1 Bacteriophage and Tol System Mutants
P1 BACTERIOPHAGE AND TOL SYSTEM MUTANTS Cari L. Smerk A Thesis Submitted to the Graduate College of Bowling Green State University in partial fulfillment of the requirements for the degree of MASTER OF SCIENCE August 2007 Committee: Ray A. Larsen, Advisor Tami C. Steveson Paul A. Moore Lee A. Meserve ii ABSTRACT Dr. Ray A. Larsen, Advisor The integrity of the outer membrane of Gram negative bacteria is dependent upon proteins of the Tol system, which transduce cytoplasmic-membrane derived energy to as yet unidentified outer membrane targets (Vianney et al., 1996). Mutations affecting the Tol system of Escherichia coli render the cells resistant to a bacteriophage called P1 by blocking the phage maturation process in some way. This does not involve outer membrane interactions, as a mutant in the energy transucer (TolA) retained wild type levels of phage sensitivity. Conversely, mutations affecting the energy harvesting complex component, TolQ, were resistant to lysis by bacteriophage P1. Further characterization of specific Tol system mutants suggested that phage maturation was not coupled to energy transduction, nor to infection of the cells by the phage. Quantification of the number of phage produced by strains lacking this protein also suggests that the maturation of P1 phage requires conditions influenced by TolQ. This study aims to identify the role that the TolQ protein plays in the phage maturation process. Strains of cells were inoculated with bacteriophage P1 and the resulting production by the phage of viable progeny were determined using one step growth curves (Ellis and Delbruck, 1938). Strains that were lacking the TolQ protein rendered P1 unable to produce the characteristic burst of progeny phage after a single generation of phage. -

Genomic Analysis of Bacteriophages SP6 and K1-5, an Estranged Subgroup of the T7 Supergroup
doi:10.1016/j.jmb.2003.11.035 J. Mol. Biol. (2004) 335, 1151–1171 Genomic Analysis of Bacteriophages SP6 and K1-5, an Estranged Subgroup of the T7 Supergroup D. Scholl1*, J. Kieleczawa2, P. Kemp3, J. Rush4, C. C. Richardson4 C. Merril1, S. Adhya5 and I. J. Molineux3,6 1Section of Biochemical We have determined the genome sequences of two closely related lytic Genetics, The National bacteriophages, SP6 and K1-5, which infect Salmonella typhimurium LT2 Institute of Mental Health and Escherichia coli serotypes K1 and K5, respectively. The genome organ- NIH, 9000 Rockville Pike ization of these phages is almost identical with the notable exception of Bethesda, MD 20895, USA the tail fiber genes that confer the different host specificities. The two phages have diverged extensively at the nucleotide level but they are still 2Wyeth/Genetics Institute more closely related to each other than either is to any other phage cur- Cambridge, MA 02140, USA rently characterized. The SP6 and K1-5 genomes contain, respectively, 3Microbiology and Molecular 43,769 bp and 44,385 bp, with 174 bp and 234 bp direct terminal repeats. Genetics, University of Texas About half of the 105 putative open reading frames in the two genomes Austin, TX 78712, USA combined show no significant similarity to database proteins with a 4 known or predicted function that is obviously beneficial for growth of a Department of Biological bacteriophage. The overall genome organization of SP6 and K1-5 is com- Chemistry and Molecular parable to that of the T7 group of phages, although the specific order of Pharmacology, Harvard genes coding for DNA metabolism functions has not been conserved. -

Ancestral Gene Acquisition As the Key to Virulence Potential in Environmental Vibrio Populations
The ISME Journal (2018) 12:2954–2966 https://doi.org/10.1038/s41396-018-0245-3 ARTICLE Ancestral gene acquisition as the key to virulence potential in environmental Vibrio populations 1,2 1,2 1,2 1,2 2 1,3 Maxime Bruto ● Yannick Labreuche ● Adèle James ● Damien Piel ● Sabine Chenivesse ● Bruno Petton ● 4 1,2 Martin F. Polz ● Frédérique Le Roux Received: 4 March 2018 / Revised: 30 June 2018 / Accepted: 6 July 2018 / Published online: 2 August 2018 © The Author(s) 2018. This article is published with open access Abstract Diseases of marine animals caused by bacteria of the genus Vibrio are on the rise worldwide. Understanding the eco- evolutionary dynamics of these infectious agents is important for predicting and managing these diseases. Yet, compared to Vibrio infecting humans, knowledge of their role as animal pathogens is scarce. Here we ask how widespread is virulence among ecologically differentiated Vibrio populations, and what is the nature and frequency of virulence genes within these populations? We use a combination of population genomics and molecular genetics to assay hundreds of Vibrio strains for their virulence in the oyster Crassostrea gigas, a unique animal model that allows high-throughput infection assays. We 1234567890();,: 1234567890();,: show that within the diverse Splendidus clade, virulence represents an ancestral trait but has been lost from several populations. Two loci are necessary for virulence, the first being widely distributed across the Splendidus clade and consisting of an exported conserved protein (R5.7). The second is a MARTX toxin cluster, which only occurs within V. splendidus and is for the first time associated with virulence in marine invertebrates. -

Herpesviral Latency—Common Themes
pathogens Review Herpesviral Latency—Common Themes Magdalena Weidner-Glunde * , Ewa Kruminis-Kaszkiel and Mamata Savanagouder Department of Reproductive Immunology and Pathology, Institute of Animal Reproduction and Food Research of Polish Academy of Sciences, Tuwima Str. 10, 10-748 Olsztyn, Poland; [email protected] (E.K.-K.); [email protected] (M.S.) * Correspondence: [email protected] Received: 22 January 2020; Accepted: 14 February 2020; Published: 15 February 2020 Abstract: Latency establishment is the hallmark feature of herpesviruses, a group of viruses, of which nine are known to infect humans. They have co-evolved alongside their hosts, and mastered manipulation of cellular pathways and tweaking various processes to their advantage. As a result, they are very well adapted to persistence. The members of the three subfamilies belonging to the family Herpesviridae differ with regard to cell tropism, target cells for the latent reservoir, and characteristics of the infection. The mechanisms governing the latent state also seem quite different. Our knowledge about latency is most complete for the gammaherpesviruses due to previously missing adequate latency models for the alpha and beta-herpesviruses. Nevertheless, with advances in cell biology and the availability of appropriate cell-culture and animal models, the common features of the latency in the different subfamilies began to emerge. Three criteria have been set forth to define latency and differentiate it from persistent or abortive infection: 1) persistence of the viral genome, 2) limited viral gene expression with no viral particle production, and 3) the ability to reactivate to a lytic cycle. This review discusses these criteria for each of the subfamilies and highlights the common strategies adopted by herpesviruses to establish latency. -

Bacterial Virus Ontology; Coordinating Across Databases
viruses Article Bacterial Virus Ontology; Coordinating across Databases Chantal Hulo 1, Patrick Masson 1, Ariane Toussaint 2, David Osumi-Sutherland 3, Edouard de Castro 1, Andrea H. Auchincloss 1, Sylvain Poux 1, Lydie Bougueleret 1, Ioannis Xenarios 1 and Philippe Le Mercier 1,* 1 Swiss-Prot group, SIB Swiss Institute of Bioinformatics, CMU, University of Geneva Medical School, 1211 Geneva, Switzerland; [email protected] (C.H.); [email protected] (P.M.); [email protected] (E.d.C.); [email protected] (A.H.A.); [email protected] (S.P.); [email protected] (L.B.); [email protected] (I.X.) 2 University Libre de Bruxelles, Génétique et Physiologie Bactérienne (LGPB), 12 rue des Professeurs Jeener et Brachet, 6041 Charleroi, Belgium; [email protected] 3 European Molecular Biology Laboratory, European Bioinformatics Institute (EMBL-EBI), Wellcome Trust Genome Campus, Hinxton CB10 1SD, UK; [email protected] * Correspondence: [email protected]; Tel.: +41-22379-5870 Academic Editors: Tessa E. F. Quax, Matthias G. Fischer and Laurent Debarbieux Received: 13 April 2017; Accepted: 17 May 2017; Published: 23 May 2017 Abstract: Bacterial viruses, also called bacteriophages, display a great genetic diversity and utilize unique processes for infecting and reproducing within a host cell. All these processes were investigated and indexed in the ViralZone knowledge base. To facilitate standardizing data, a simple ontology of viral life-cycle terms was developed to provide a common vocabulary for annotating data sets. New terminology was developed to address unique viral replication cycle processes, and existing terminology was modified and adapted. -
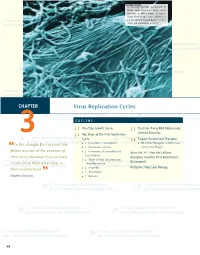
Virus Replication Cycles
© Jones and Bartlett Publishers. NOT FOR SALE OR DISTRIBUTION A scanning electron micrograph of Ebola virus particles. Ebola virus contains an RNA genome. It causes Ebola hemorrhagic fever, which is a severe and often fatal disease in hu- mans and nonhuman primates. CHAPTER Virus Replication Cycles OUTLINE 3.1 One-Step Growth Curves 3.3 The Error-Prone RNA Polymerases: 3 3.2 Key Steps of the Viral Replication Genetic Diversity Cycle 3.4 Targets for Antiviral Therapies In the struggle for survival, the ■ 1. Attachment (Adsorption) ■ RNA Virus Mutagens: A New Class “ ■ 2. Penetration (Entry) of Antiviral Drugs? fi ttest win out at the expense of ■ 3. Uncoating (Disassembly and Virus File 3-1: How Are Cellular Localization) their rivals because they succeed Receptors Used for Viral Attachment ■ 4. Types of Viral Genomes and Discovered? in adapting themselves best to Their Replication their environment. ■ 5. Assembly Refresher: Molecular Biology ” ■ 6. Maturation Charles Darwin ■ 7. Release 46 229329_CH03_046_069.indd9329_CH03_046_069.indd 4466 11/18/08/18/08 33:19:08:19:08 PPMM © Jones and Bartlett Publishers. NOT FOR SALE OR DISTRIBUTION CASE STUDY The campus day care was recently closed during the peak of the winter fl u season because many of the young children were sick with a lower respiratory tract infection. An email an- nouncement was sent to all students, faculty, and staff at the college that stated the closure was due to a metapneumovirus outbreak. The announcement briefed the campus com- munity with information about human metapneumonoviruses (hMPVs). The announcement stated that hMPV was a newly identifi ed respiratory tract pathogen discovered in the Netherlands in 2001. -

Viruses Chapter 19
Viruses Chapter 19 What you must know: The components of a virus. The differences between lytic and lysogenic cycles. How viruses can introduce genetic variation into host organisms. Mechanisms that introduce genetic variation into viral populations. Bacteria vs. Viruses Bacteria Virus Prokaryotic cell Not a living cell (genes Most are free-living (some packaged in protein shell) parasitic) Intracellular parasite Relatively large size 1/1000 size of bacteria Antibiotics used to kill Vaccines used to prevent bacteria viral infection Antiviral treatment Viruses Very small (<ribosomes) Components = nucleic acid + capsid ◦ Nucleic acid: DNA or RNA (double or single-stranded) ◦ Capsid: protein shell ◦ Some viruses also have viral envelopes that surround capsid Derived from host cell membranes and viral proteins Viruses Limited host range ◦ Entry = attach to host cell membrane receptors through capsid proteins or glycoproteins on viral envelope (animal) ◦ Eg. human cold virus (rhinovirus) upper respiratory tract (mouth & nose) Reproduce quickly within host cells Can mutate easily ◦ RNA viruses: no error-checking mechanisms Simplified viral replicative cycle Entry: Injection endocytosis, fusion VIDEO: T4 PHAGE INFECTION Bacteriophage Virus that infects bacterial cells Viral Reproduction - phages Lytic Cycle: ◦ Use host machinery to replicate, assemble, and release copies of virus ◦ Virulent phages: Cells die through lysis or apoptosis Lysogenic (Latent) Cycle: ◦ DNA incorporated into host DNA and replicated along with it -

Ab Komplet 6.07.2018
CONTENTS 1. Welcome addresses 2 2. Introduction 3 3. Acknowledgements 10 4. General information 11 5. Scientific program 16 6. Abstracts – oral presentations 27 7. Abstracts – poster sessions 99 8. Participants 419 1 EMBO Workshop Viruses of Microbes 2018 09 – 13 July 2018 | Wrocław, Poland 1. WELCOME ADDRESSES Welcome to the Viruses of Microbes 2018 EMBO Workshop! We are happy to welcome you to Wrocław for the 5th meeting of the Viruses of Microbes series. This series was launched in the year 2010 in Paris, and was continued in Brussels (2012), Zurich (2014), and Liverpool (2016). This year our meeting is co-organized by two partner institutions: the University of Wrocław and the Hirszfeld Institute of Immunology and Experimental Therapy, Polish Academy of Sciences. The conference venue (University of Wrocław, Uniwersytecka 7-10, Building D) is located in the heart of Wrocław, within the old, historic part of the city. This creates an opportunity to experience the over 1000-year history of the city, combined with its current positive energy. The Viruses of Microbes community is constantly growing. More and more researchers are joining it, and they represent more and more countries worldwide. Our goal for this meeting was to create a true global platform for networking and exchanging ideas. We are most happy to welcome representatives of so many countries and continents. To accommodate the diversity and expertise of the scientists and practitioners gathered by VoM2018, the leading theme of this conference is “Biodiversity and Future Application”. With the help of your contribution, this theme was developed into a program covering a wide range of topics with the strongest practical aspect. -

Bacteriophage T4 Lysis and Lysis Inhibition
BACTERIOPHAGE T4 LYSIS AND LYSIS INHIBITION: MOLECULAR BASIS OF AN ANCIENT STORY A Dissertation by TRAM ANH THI TRAN Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY May 2007 Major subject: Biochemistry BACTERIOPHAGE T4 LYSIS AND LYSIS INHIBITION: MOLECULAR BASIS OF AN ANCIENT STORY A Dissertation by TRAM ANH THI TRAN Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY Approved by: Chair of Committee, Ryland F. Young Committee Members, Deborah Siegele Paul Fitzpatrick Michael Polymenis Head of Department, Gregory D. Reinhart May 2007 Major subject: Biochemistry iii ABSTRACT Bacteriophage T4 Lysis and Lysis Inhibition: Molecular Basis of an Ancient Story. (May 2007) Tram Anh Thi Tran, B.S., Stephen F. Austin State University; M.S., Stephen F. Austin State University Chair of Advisory Committee: Dr. Ryland F. Young T4 requires two proteins: holin, T (lesion formation and lysis timing) and endolysin, E (cell wall degradation) to lyse the host at the end of its life cycle. E is a cytoplasmic protein that sequestered away from its substrate, but the inner membrane lesion formed by T allows E to gain access to the cell wall. T4 exhibits lysis inhibition (LIN), a phenomenon in which a second T4 infection occurs ≤ 3 min after primary infection results a delay in lysis. Mutations that abolish LIN mapped to several genes but only rV encoding the holin, T, and rI, encoding the antiholin, RI, are required for LIN in all hosts which support T4 replication. -

The Lytic Cycle and the Lysogenic Cycle
Chapter 19 Viruses PowerPoint® Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright © 2008 Pearson Education, Inc., publishing as Pearson Benjamin Cummings Overview: A Borrowed Life • Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli • Viruses lead “a kind of borrowed life” between life-forms and chemicals • The origins of molecular biology lie in early studies of viruses that infect bacteria Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Fig. 19-1 0.5 µm Concept 19.1: A virus consists of a nucleic acid surrounded by a protein coat • Viruses were detected indirectly long before they were actually seen Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings The Discovery of Viruses: Scientific Inquiry • Tobacco mosaic disease stunts growth of tobacco plants and gives their leaves a mosaic coloration • In the late 1800s, researchers hypothesized that a particle smaller than bacteria caused the disease • In 1935, Wendell Stanley confirmed this hypothesis by crystallizing the infectious particle, now known as tobacco mosaic virus (TMV) Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Fig. 19-2 RESULTS 1 Extracted sap 2 Passed sap 3 Rubbed filtered from tobacco through a sap on healthy plant with porcelain tobacco plants tobacco filter known mosaic to trap disease bacteria -

DNA Concatemers MICHAEL D
Proc. Natl. Acad. Sci. USA Vol. 91, pp. 153-157, January 1994 Biochemistry Deletion of the E4 region of the genome produces adenovirus DNA concatemers MICHAEL D. WEIDEN AND HAROLD S. GINSBERG* Departments of Medicine and Microbiology, College of Physicians and Surgeons, Columbia University, 630 West 168th Street, New York, NY 10032 Contributed by Harold S. Ginsberg, September 20, 1993 ABSTRACT Two mutants containing large deletions in the by replication-induced recombination (11). The last 45 bp of E4 region of the adenovirus genome H5dl366 (91.9-98.3 map the genome contain the necessary sequence information for units) and H2dl808 (93.0-97.1 map units) were used to inves- replication, and any deletion of the terminal cytosine abol- tigate the role of E4 genes in adenovirus DNA synthesis. ishes DNA synthesis (12, 13). Infection of KB human epidermoid carcinoma cells with either Other viral gene products may be involved in viral DNA mutant resulted in production of large concatemers of viral synthesis, and other mechanisms not yet elucidated may be DNA. Only monomer viral genome forms were produced, required for completion of viral DNA replication. Some however, when mutants infected W162 cells, a monkey kidney phenomena not accounted for in the current model include cell line transformed with and expressing the E4 genes. Dif- the following: circular, supercoiled, and multimeric viral fusible E4 gene products, therefore, complement the E4 mutant DNA molecules, which were identified in electron micro- phenotype. The viral DNA concatemers produced in d1366- and scopic studies (14); the pL 5'-to-3' exonuclease required in d1808-infected KB cells did not have any specific orientation of the in vitro model (9) produces defective strands that need monomer joining: the junctions consisted of head-to-head, additional processing for maturation; a large proportion of head-to-tail, and tall-to-tail joints.